Abstract
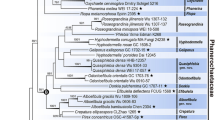
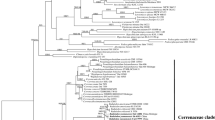
Nucleotide sequences of the small-subunit (SSU) ribosomal DNA were determined forPelvetia babingtonii, P. canaliculate, Pelvetiopsis limitata, andAscophyllum nodosum in the family Fucaceae. A total of 1755 positions were aligned for the whole sequence. The positional differences in the primary structure among the taxa ranged from 16 to 30 nucleotide changes in pairwise comparisons. There was a minimum divergence betweenPs. limitata andP. babingtonii while a maximum betweenPs. limitata andP. canaliculata. The SSU rDNA trees showed that the genusPelvetia was not monophyletic and the genusPelvetiopsis was not closely related toPelvetia. Our results suggest that the taxonomic revision of the genusPelvetia as well as the family Fucaceae is needed based on detailed morphological observations.
Similar content being viewed by others
Literature Cited
Bhattacharya, D., L. Medlin, P.O. Wainright, E.V. Ariztia, C. Biheau, S.K. Stickel and M.L. Sogin. 1992. Algae containing chlorophlls a+c arc para-phylctic: molecular evolutionary analysis of the chloro-phyta.Evolution46: 1801–1817.
Capesius, I. and M. Stech. 1997. Molecular relationships within mosses based on 1SS rRNA gene sequences.Nova Hedw.64: 525–533.
Clayton, M.N. 1984. Evolution of the Phaeophyceae with particular reference to the Fucales.Prog. Phycol. Res.3: 11–46.
De Toni, G.B. 1895. Sylloge algarum vol 3. Patavii. p. 216.
Felsenstein, J. 1985. Confidence limits on phylogeneties: an approach using the bootstrap.Evolution39: 783–791.
Felsenstein, J. 1995. Phylogeny Inference Packages, version 3.57. The University of Washington.
Gardner, N.L. 1913. New Fucaceae.Univ. Calif. Publ. Hot.4: 317–374.
Hillis, D.M. and J.J. Bull. 1993. An empirical test of boolstrapping as a method for assessing confidence in phylogenetic analysis.Syst. Biol.42: 182–192.
Kawai, H., H. Muto, T. Fujii and A. Kato. 1995. A linked 5S rRNA gene inScytosiphon lomentaria (Scytosiphonaecae, Phaeophyceae).J. Phycol.31: 306–311.
Kimura, M. 1980. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences.J. Mol. Evol.16: 111–120.
Lee, R.E. 1989. Phycology. 2nd ed. Cambridge Univ. Press. Cambridge. 645 pp.
Lee, W.J. and R.J. King. 1996. The molecular characteristics of five genera of Dictyotaceae (Phaeophyta) from Australia: based on DNA sequence of Nuclear rDNA internal transcribed spacer (ITS) and 5.8s.Algae11: 381–388.
Lim, B.L., H. Kawai, H. Hori and S. Osawa. 1986. Molecular evolution of 5S ribosomal RNA from red and brown algae.Jpn. J. Genet.61: 169–176.
Nakayama, T., S. Watanabe, K. Mitsui, H. Uchida and I. Inouye. 1996. The Phylogenetic relationship between the Chlamydomonadales and Chlorococcales infered from 18s rDNA sequence data.Phycol. Res.44: 47–55.
Powell, H.T. 1963. Speciation in the genusFucus L. and related genera. Speciation in the Sea. pp. 66–77.
Rousseau, F., M.-C. Leclerg and B. de Reviers. 1997. Molecular phylogeny of European Fucales (Phaeophyceae) based on partial large-subunit rDNA sequence comparisons.Phycologia36: 438–446.
Saunders, G.W. and L.D. Druehl. 1992. Nucleotide sequences of the small-subunit ribosomal RNA genes from selected Laminariales (Phaeophyta); implications for kelp evolution.J. Phycol.28: 544–549.
Saunders, G.W. and G.T. Kraft. 1995. The phylogenetic affinities ofNotheia anomala (Fucales, Phaeophyceae) as determinded from partial small-subunit rRNA gene sequences.Phycologia34: 383–389.
Shchapova, T. 1945. On the amphipacific distribution of certain species of Phaeophyceacc.Comp. Ren. Acad. Sci. URSS.52: 171–173.
Serrao, E.A. and S.H. Brawley. 1997. Phylogeny of the Fucaceae based on DNA sequences of rRNA-ITS.Phycologia36 (supplement): 101.
Setchell, W.A. and N.L. Gardner. 1925. The marine algae of the Pacific coast of North America. Melano-phyceae.Univ. Calif. Publ. Bot.8: 590–649.
Song, H.S., K.S. Seo and S.M. Boo. 1996. Field studies of the brown algaPelvetia siliquosa with implication for taxonomy and distribution.Algae11: 65–71.
Swofford, D.L. 1993. PAUP: Phylogenetic analysis using parsimony. Version 3.1.1. Illinois Natural History Survey. Champaign.
Ian, I.H. and L.D. Druehl. 1993. Phylogeny of the Northeast Pacific brown algal (Phaeophyceae) orders as inferred from 18S rDNA gene sequences.Hy-drobiologia.260/261: 699–704.
Tan, I.H. and L.D. Druehl. 1996. A ribosomal DNA phylogeny supports the close evolutionary relationships among the Sporochanles, Desmarestiales and Laminariales (Phaeophyceae).J. Phycol.32: 112–118.
Thompson, J.D., D.G. Higgins and T.J. Gibson. 1994. CLUSTAL W. improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice.Nucl. Acids Res.22: 4673–4680.
Tseng, C.K. and C.F. Chang. 1953. On a new species ofPelvetia and its distribution.Acta Biol. Sinica2: 280–297.
van den Hoek, C., D.G. Mann and H.M. Jahns. 1995. Algae: an introduction to phycology. Cambridge Univ. Press. Cambridge. 623 pp.
Wray, C.G., N.H. Landman, W.B. Saunders and J. Bonacum. 1995. Generic divergence and geographic diversification inNautilus.Paleobiology21: 220–228.
Wynne, M.J. and S. Loiseaux. 1978. Phycological reviews 5: recent advances in life history studies of the Phaeophyta.Phycologia15: 425–452.
Yoshida, T. and P.C. Silva. 1992. On the identity ofFucus babingtonit Harvery Fucales. Phaeophyta).Jpn. J. Phvcol.40: 121–124.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Lee, W.J., Yoon, H.S. & Boo, S.M. Phylogenetic relationships ofPelvetia andPelvetiopsis (Phaeophyceae) based on small subunit ribosomal DNA sequences. J. Plant Biol. 41, 103–109 (1998). https://doi.org/10.1007/BF03030396
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF03030396