Abstract
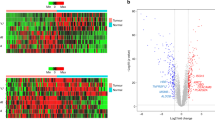
Various genomic technologies have been applied to address crucial problems in cancer biology, because cancer develops through the accumulation of various genetic alterations. Of these, gene expression profiling analysis using microarray technology has been widely applied not only to classify cancers at molecular levels, but also to identify novel molecular targets for therapeutics and/or diagnostics. To gain molecular understanding of gastric carcinogenesis, progression, and diversity, we analyzed primary advanced gastric cancer and noncancerous gastric tissues by high-density oligonucleotide microarray. Genes differentially expressed between cancer and noncancerous tissues were identified. In cancer tissues, genes related to cell cycle, growth factor, cell motility, cell adhesion, and matrix remodeling were highly expressed, whereas those related to gastrointestinal-specific function and immune response were rather downregulated. These results provide not only a new molecular basis for understanding biological properties of gastric cancer but also useful resources for future development of therapeutic and diagnostic biomarkers for gastric cancer. Several microarray studies have been published since and have been compared for validation in meta-analysis. As integration of transcriptome information with other biological data is crucial to interpret gene expression data, we have applied oligonucleotide microarray technology to assess allelic gene dosage at 10000 polymorphic loci, namely with an average interval of 200 kb. Using a newly developed algorithm, genome imbalance map, loss of heterozygosity (LOH) status can be determined simultaneously. Besides several loci with genomic amplification, we also identified a homozygously deleted chromosomal region in 7q, where frequent chromosomal instability was observed. Finally, we are currently developing novel biomarkers for gastroenterological cancers. Glypican 3 is detected at high levels in serum of hepatocellular carcinoma patients and could be a potential target for antibody therapy.
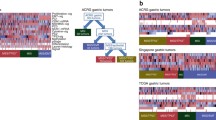
Similar content being viewed by others
References
Stears RL, Martinsky T, Schena M. Trends in microarray analysis. Nat Med 2003;9: 140–5.
Hippo Y, Taniguchi H, Tsutsumi S, Machida N, Chong JM, Fukayama M, et al. Global gene expression analysis of gastric cancer by oligonucleotide microarrays. Cancer Res 2002;62: 233–40.
Leung SY, Chen X, Chu KM, Yuen ST, Mathy J, Ji J, et al. Phospholipase A2 group IIA expression in gastric adenocarcinoma is associated with prolonged survival and less frequent metastasis. Proc Natl Acad Sci U S A 2002; 99: 16203–8.
Boussioutas A, Li H, Liu J, Waring P, Lade S, Holloway AJ, et al. Distinctive patterns of gene expression in premalignant gastric mucosa and gastric cancer. Cancer Res 2003;63: 2569–77.
Tay ST, Leong SH, Yu K, Aggarwal A, Tan SY, Lee CH, et al. A combined comparative genomic hybridization and expression microarray analysis of gastric cancer reveals novel molecular subtypes. Cancer Res 2003; 63: 3309–16.
Jinawath N, Furukawa Y, Hasegawa S, Li M, Tsunoda T, Satoh S, et al. Comparison of gene-expression profiles between diffuse-and intestinal-type gastric cancers using a genome-wide cDNA microarray. Oncogene 2004; 23: 6830–44.
Hippo Y, Yashiro M, Ishii M, Taniguchi H, Tsutsumi S, Hirakawa K, et al. Differential gene expression profiles of scirrhous gastric cancer cells with high metastatic potential to peritoneum or lymph nodes. Cancer Res 2001; 61: 889–95.
Leung SY, Yuen ST, Chu KM, Mathy JA, Li R, Chan AS, et al. Expression profiling identifies chemokine (C-C motif) ligand 18 as an independent prognostic indicator in gastric cancer. Gastroenterology 2004;127: 459–69.
Kano M, Nishimura K, Ishikawa S, Tsutsumi S, Hirota K, Hirose M, et al. Expression imbalance map: a new visualization method for detection of mRNA expression imbalance regions. Physiol Genomics 2003; 13: 31–46.
Midorikawa Y, Tsutsumi S, Nishimura K, Kamimura N, Kano M, Sakamoto H, et al. Distinct chromosomal bias of gene expression signatures in the progression of hepatocellular carcinoma. Cancer Res 2004; 64: 7263–70.
Mantripragada KK, Buckley PG, de Stahl TD, Dumanski JP. Genomic microarrays in the spotlight. Trends Genet 2004; 20: 87–94.
Pinkel D, Segraves R, Sudar D, Clark S, Poole I, Kowbel D, et al. High resolution analysis of DNA copy number variation using comparative genomic hybridization to microarrays. Nat Genet 1998; 20: 207–11.
Pollack JR, Perou CM, Alizadeh AA, Eisen MB, Pergamenschikov A, Williams CF, et al. Genome-wide analysis of DNA copy-number changes using cDNA microarrays. Nat Genet 1999; 23: 41–6.
Lucito R, Healy J, Alexander J, Reiner A, Esposito D, Chi M, et al. Representational oligonucleotide microarray analysis: a high-resolution method to detect genome copy number variation. Genome Res 2003;13: 2291–305.
Bignell GR, Huang J, Greshock J, Watt S, Butler A, West S, et al. High-resolution analysis of DNA copy number using oligonucleotide microarrays. Genome Res 2004; 14: 287–95.
Strausberg RL, Simpson AJ, Old LJ, Riggins GJ. Oncogenomics and the development of new cancer therapies. Nature (Lond) 2004; 429: 469–74.
Midorikawa Y, Ishikawa S, Iwanari H, Imamura T, Sakamoto H, Miyazono K, et al. Glypican-3, overexpressed in hepatocellular carcinoma, modulates FGF2 and BMP-7 signaling. Int J Cancer 2003; 103: 455–65.
Hippo Y, Watanabe K, Watanabe A, Midorikawa Y, Yamamoto S, Ihara S, et al. Identification of soluble NH2-terminal fragment of glypican-3 as a serological marker for early-stage hepatocellular carcinoma. Cancer Res 2004; 64: 2418–23.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Aburatani, H. Discovery of a new biomarker for gastroenterological cancers. J Gastroenterol 40 (Suppl 16), 1–6 (2005). https://doi.org/10.1007/BF02990571
Issue Date:
DOI: https://doi.org/10.1007/BF02990571