Abstract
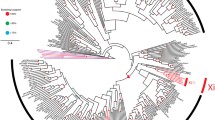
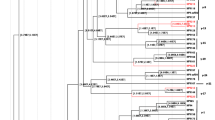
Papillomaviruses are responsible for a variety of diseases in humans and animals, ranging from harmless skin warts to lethal cancers. They also make up one of the most genetically diversified families of viruses known, and could represent a model system of DNA-virus evolution. A specialized genetic sequences database, The Human Papillomavirus Database and Analysis Project, was recently established in an effort to provide database services that are specific to papillomaviruses to the research community and to perform a variety of sequence-based analyses. This review is intended to present the scope of the information currently contained in the database and to outline some of the analyses that have been performed on the genetic sequences. These analyses will address issues including phylogenetic relationships, recombination events, selective pressures on different genes and the possibility of cross-species transmission in the case of the papillomaviruses.
Article PDF
Similar content being viewed by others
References
Bernard HU, Chan SY, Delius H. Evolution of papillomaviruses. In: zur Hausen H, ed. Human Pathogenic Papillomaviruses. Berlin, Springer, 34–54;1994.
Bernard HU, Chan SY, Ong CK, Villa LL, Delius H, Peyton CL, Bauer HM, Manos MM, Wheeler CM. Identification and assessment of known and novel human papillomaviruses by polymerase chain reaction, restriction digestion fingerprinting, nucleotide sequence, and phylogenetic algorithms. J Infect Dis 170:1077–1085;1994.
Chan SY, Delius H, Bernard HU. Phylogenetic and taxonomic evaluation of 96 papillomavirus types. (submitted).
Chan SY, Ho L, Ong CK, Chow V, Drescher B, Durst M, ter Meulen J, Villa L, Luande J, Mgaya HN, Bernard HU. Molecular variants of human papillomavirus type 16 from four continents suggest ancient pandemic spread of the virus and its coevolution with humankind. J Virol 66:2057–2066;1992.
Chong T, Chan WK, Bernard HU. Transcriptional activation of human papillomavirus 16 by nuclear factor I, AP1, steroid receptors and a possibly novel transcription factor, PVF: A model for the composition of genital papillomavirus enhancers. Nucleic Acids Res 18:465–470;1990.
Coggins JR, zur Hausen H. Workshop on papillomaviruses and cancer. Cancer Res 39:545–546;1979.
Deau MC, Favre M, Orth G. Genetic heterogeneity among human papillomaviruses (HPV) associated with epidermodysplasia verruciformis: Evidence for multiple allelic forms of HPV5 and HPV8 E6 genes. Virology 184:492–503;1991.
de Villiers EM. Human pathogenic papillomavirus types: An update. In: zu Hausen H, ed. Human Pathogenic Papillomaviruses. Berlin, Springer, 1–12;1994.
Hillis DM, Allard MW, Miyamoto MM. Analysis of DNA sequence data: Phylogenetic inference. Methods Enzymol 224:456–487;1993.
Hillis DM, Huelsenbeck JP, Cunningham CW. Application and accuracy of molecular phylogenies. Science 264:671–677;1994.
Jukes TH, Cantor CR. Evolution of protein molecules. In: Munroe H, ed. Mammalian Protein Metabolism. New York, Academic Press, 21–132;1969.
Kimura M. A simple method for estimating evolutionary rate of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120;1980.
Korber BTM, Farber RM, Wolpert DH, Lapedes AS. Covariation of mutations in the V3 loop of human immunodeficiency virus type 1 envelope protein: An information theoretic analysis. Proc Natl Acad Sci USA 90:9061–9065;1993.
Korber BTM, MacInnes K, Smith RF, Myers G. Mutational trends in V3 loop protein sequences observed in different genetic lineages of human immunodeficiency virus type 1. J Virol 68:6730–6744;1994.
Korber B, Myers G. Signature pattern analysis: A method for assessing viral sequence relatedness. AIDS Res Hum Retroviruses 8:1549–1560;1992.
Laga M, Diallo MO, Buve A. Inter-relationship of sexually transmitted diseases and HIV: Where are we now? AIDS 8 (suppl 1):S119-S124;1994.
Maddison WP, Maddison DR. MacClade: Analysis of Phylogeny and Character Evolution. Sunderland, Sinauer, 1992.
Myers G. Assimilating HIV sequences. AIDS Res Hum Retroviruses 9:697–702;1993.
Myers G, Bernard HU, Delius H, Favre M, Icenogel J, van Ranst M, Wheeler C. Human Papillomaviruses: A Compilation and Analysis of Nucleic Acid and Amino Acid Sequences. Los Alamos, Theoretical Biology and Biophysics, Los Alamos National Laboratory, 1994.
Nei M, Gojobori T. Simple methods for estimating the numbers of synonymous and nonsynonymous substitutions. Mol Biol Evol 2:418–426;1986.
Ong CK, Chan SY, Campo MS, Fujinaga K, Mavromara-Nagos P, Labropoulou V, Pfister H, Tay SK, ter Meulen J, Villa LL, Bernard HU. Evolution of human papillomavirus type 18: An ancient phylogenetic root in Africa and intratype diversity reflect coevolution with human ethnic groups. J Virol 67:6424–6431;1993.
Rowson KEK, Mahy BWJ. Human papova (wart) virus. Bacteriol Rev 31:110–131;1967.
Smith RF, Smith TF. Automatic generation of primary sequence pattern from sets of related sequences. Proc Natl Acad Sci USA 87:118–121;1990.
Sundberg JP. Papillomavirus infections in animals. In: Syrjaenen K, Gissmann L, Koss LG, eds. Papillomaviruses and Human Disease. Berlin, Springer, 40–103;1987.
Swofford DL. Paup: Phylogenetic Analysis Using Parsimony (Version 3.1). Computer program distributed by the Illinois Natural History Survey, Champaign, Illinois, 1991.
Van Ranst MA, Kaplan JB, Burk RD. Phylogenetic classification of papillomaviruses: Correlation with clinical manifestations. J Gen Virol 73:2653–2660;1992.
Van Ranst MA, Tachezy R, Burk RD. Human papillomavirus nucleotide sequences: What's in stock? Papillomavirus Rep 6:65–75;1994.
Zur Hausen H. Preface. In: zur Hausen H, ed. Human Pathogenic Papillomaviruses. Berlin, Springer, v-ix;1994.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Farmer, A.D., Calef, C.E., Millman, K. et al. The Human Papillomavirus Database. J Biomed Sci 2, 90–104 (1995). https://doi.org/10.1007/BF02253061
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF02253061