Summary
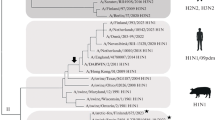
The nucleoprotein (NP) gene of influenza A viruses is decisive for separating two large individually evolving reservoirs in birds and humans. A phylogenetic analysis of the NP gene revealed that all mammalian influenza viruses originated — directly or indirectly — from an avian ancestor. The stable introduction of an avian influenza A virus into a mammalian species seems to be a relatively rare event, the latest one occurred in 1979 when such an avian virus was introduced into pigs in Northern Europe which gave rise to a new lineage. At least two concomitant events are required for such a new and stable introduction: (1) The new species has to become infected, and (2) a mutation in the polymerase complex has to establish a labile variant, which is prone to provide a large number of different variants, from which some can adapt rapidly to the new host (or to any unusual environments). Since such mutator mutations might be advantageous only during stress periods, variants with a less error prone polymerase might emerge again after adaptation. Examples for such fluctuations in terms of mutational and evolutionary rates are discussed in this brief review.
Similar content being viewed by others
References
Altmüller A, Fitch WM, Scholtissek C (1989) Biological and genetic evolution of the nucleoprotein (NP) gene of human influenza A viruses. J Gen Virol 70: 2111–2119
Altmüller A, Kunerl M, Müller K, Hinshaw VS, Fitch WM, Scholtissek C (1991) Genetic relatedness of the nucleoprotein (NP) of recent swine, turkey, and human influenza A virus (H1 N1) isolates. Virus Res 82: 79–87
Bean WJ, Kawaoka Y, Wood JM, Pearson JE, Webster RG (1985) Characterization of virulent and avirulent A/chicken/Pennsylvania/83 influenza A viruses: potential role of defective interfering RNAs in nature. J Virol 54: 151–160
Donis RO, Bean WJ, Kawaoka Y, Webster RG (1989) Distinct lineages of influenza virus H 4 hemagglutinin genes in different regions of the world. Virology 169: 408–417
Fitch WM (1971) Toward defining the course of evolution: minimum change for a specific tree topology. Syst Zool 20: 406–416
Gammelin M, Mandler J, Scholtissek C (1989) Two subtypes of nucleoproteins (NP) of influenza A viruses. Virology 170: 71–80
Gammelin M, Altmüller A, Reinhardt U, Mandler J, Harley VR, Hudson PJ, Fitch WM, Scholtissek C (1990) Phylogenetic analysis of nucleoproteins suggests that human influenza A viruses emerged from a 19th century avian ancestor. Mol Biol Evol 7: 194–200
Gorman OT, Bean WJ, Kawaoka Y, Webster RG (1990) Evolution of the nucleoprotein gene of influenza A virus. J Virol 64: 1487–1497
Gorman OT, Bean WJ, Kawaoka Y, Donatelli I, Guo Y, Webster RG (1991) Evolution of influenza A virus nucleoprotein genes: implications for the origin of H1 N1 human and classical swine viruses. J Virol 65: 3704–3714
Gorman OT, Bean WJ, Webster RG (1992) Evolutionary processes in influenza viruses: Divergence, rapid evolution, and stasis. Curr Top Microbiol Immunol 176: 75–97
Hinshaw VS, Alexander DJ, Aymard M, Bachmann PA, Easterday BC, Hannoun C, Kida H, Lipkind M, MacKenzie JS, Nerome K, Schild GC, Scholtissek C, Senne DA, Shortridge KF, Skehel JJ, Webster RG (1984) Antigenic comparisons of swine influenza-like H1 N1 isolates from pigs, birds, and humans: an international collaborative study. Bull World Health Organ 62: 871–878
Kaplan MM, Webster RG (1977) The epidemiology of influenza. Sci Am 12: 88–106
Kistner O, Müller K, Scholtissek C (1989) Differential phosphorylation of the nucleoprotein of influenza A viruses. J Gen Virol 70: 2421–2431
Lamb RA (1983) The influenza virus RNA segments and their encoded proteins. In: Palese P, Kingsbury DW (eds) Genetics of influenza viruses. Springer, Wien New York, pp 21–69
Ludwig S, Schultz U, Mandler J, Fitch WM, Scholtissek C (1991) Phylogenetic relationship of the nonstructural (NS) genes of influenza A viruses. Virology 183: 568–577
Ludwig S, Haustein A, Kaleta EF, Scholtissek C (1993) Recent influenza infections in turkeys and pigs in Northern Europe. In: Hinshaw VS, Easterday BC (eds) Proceedings of the third international symposium on avian influenza. Extension Duplicating Services, University of Wisconsin, Madison, pp 131–135
Mandler J, Gorman OT, Ludwig S, Schroeder E, Fitch WM, Webster RG, Scholtissek C (1990) Derivation of the nucleoproteins (NP) of influenza A viruses isolated from marine mammals. Virology 176: 255–261
Pinto LH, Holsinger LJ, Lamb RA (1992) Influenza virus M 2 protein has ion channel activity. Cell 69: 517–528
Scholtissek C, Harms E, Rohde W, Orlich M, Rott R (1976) Correlation between RNA fragments of fowl plague virus and their corresponding gene functions. Virology 74: 332–344
Scholtissek C, Rohde W, von Hoyningen V, Rott R (1978) On the origin of the human influenza virus subtypes H2 N2 and H3 N2. Virology 87: 13–20
Scholtissek C, Könnecke I, Rott R (1978) Host range recombinants of fowl plague (influenza A) virus. Virology 91: 79–85
Scholtissek C, Vallbracht A, Flehmig B, Rott R (1979) Correlation of pathogenicity and gene constellation of influenza A viruses. II. Highly neurovirulent recombinants derived from non-virulent or weekly neurovirulent parent virus strains. Virology 95: 492–500
Scholtissek C, Bürger H, Bachmann PA, Hannoun C (1983) Genetic relatedness of hemagglutinins of the H 1 subtype of influenza A viruses isolated from swine and birds. Virology 129: 521–523
Scholtissek C, Bürger H, Kistner O, Shortridge KF (1985) The nucleoprotein as a possible major factor in determining host specificity of influenza H3 N2 viruses. Virology 147: 287–294
Scholtissek C, Naylor E (1988) Fish farming and influenza pandemics. Nature 331: 215
Schultz U, Fitch WM, Ludwig S, Mandler J, Scholtissek C (1991) Evolution of pig influenza viruses. Virology 183: 61–73
Snyder MH, Buckler-White AJ, London WT, Tierney EL, Murphy BR (1987) The avian influenza virus nucleoprotein gene and a specific constellation of avian and human virus polymerase genes each specify attenuation of avian-human influenza A/pintail/79 reassortant viruses for monkeys. J Virol 61: 2857–2863
Suárez P, Valcácell J, Ortín J (1992) Heterogeneity of the mutation rates of influenza A viruses: isolation of mutator mutants. J Virol 66: 2491–2494
Tian S-F, Buckler-White AJ, London WT, Recle LJ, Chanock RM, Murphy BR (1985) Nucleoprotein and membrane protein genes are associated with restriction of replication of influenza A/mallard/NY/78 virus and its reassortants in squirrel monkey respiratory tract. J Virol 53: 771–775
Vogel U, Kunerl M, Scholtissek C (1993) Influenza A virus late mRNAs are specifically retained in the nucleus in the presence of a methyltransferase or a protein kinase inhibitor. Virology (in press)
von Hoyningen-Huene V, Scholtissek C (1983) Low genetic mixing between influenza viruses of different geographic regions. Arch Virol 76: 63–67
Webster RG, Bean WJ, Gorman OT, Chambers TM, Kawaoka Y (1992) Evolution and ecology of influenza A viruses. Microbiol Rev 56: 152–179
Witte KH, Nienhoff E, Ernst H, Schmidt U, Prager D (1981) Erstmaliges Auftreten einer durch das Schweineinfluenzavirus verursachten Epizootie in Schweinebeständen der Bundesrepublik Deutschland. Tierärztl Umschau 36: 591–606
Zebedee SJ, Lamb RA (1988) Influenza A virus M 2 protein monoclonal antibody restriction of virus growth and detection of M 2 in virions. J Virol 62: 2762–2772
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Scholtissek, C., Ludwig, S. & Fitch, W.M. Analysis of influenza A virus nucleoproteins for the assessment of molecular genetic mechanisms leading to new phylogenetic virus lineages. Archives of Virology 131, 237–250 (1993). https://doi.org/10.1007/BF01378629
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF01378629