Summary
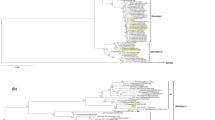
The nucleotide sequences of the NS 1 genes from five Thai and three Sri Lankan dengue-2 viruses were determined by sequencing the viral RNA using synthetic oligonucleotide primers. The results were shown to be similar to four published dengue-2 NS 1 sequences and the classification of these genes was compared with the one obtained for the envelope genes of the same viruses. The classification was similar and showed that the Thai isolates could be divided into two separate groups and that the Sri Lankan isolates were distinct. We found no correlation between disease severity, serological response (1° or 2°), or year of isolation and various aspects of NS 1 protein sequence variation; and no particular amino acid changes were correlated with virulence. The sequences were combined with those published and classified elsewhere to provide a comprehensive E/NS 1 gene taxonomy of dengue-2 virus isolates.
Article PDF
Similar content being viewed by others
References
Air GM, Gibbs AJ, Laver WG, Webster RG (1990) Evolutionary changes in influenza B are not primarily governed by antibody selection. Proc Natl Acad Sci USA 87: 3884–3888
Blok J, Air GM (1980) Comparative nucleotide sequences at the 3′ end of the neuraminidase gene from eleven influenza type A viruses. Virology 107: 50–60
Blok J, Henchal EA, Gorman BM (1984) Comparison of dengue viruses and some other flaviviruses by cDNA-RNA hybridization analysis and detection of a close relationship between dengue virus serotype 2 and Edge Hill virus. J Gen Virol 65: 2173–2181
Blok J, Kay BH, Hall RA, Gorman BM (1988) Isolation and characterization of dengue viruses serotype 1 from an epidemic in northern Queensland, Australia. Arch Virol 100: 213–220
Blok J, Samuel S, Gibbs AJ, Vitarana UT (1989) Variation of the nucleotide and encoded amino acid sequences of the envelope gene from eight dengue-2 viruses. Arch Virol 105: 39–53
Brandt WE, Cardiff RD, Russell PK (1970) Dengue virions and antigens in brain and serum of infected mice. J Virol 6: 500–506
Brandt WE, Chiewslip D, Harris DL, Russell PK (1970) Partial purification and characterization of a dengue virus soluble complement fixing antigen. J Immunol 105: 1565–1568
Cardiff RD, Lund JL (1976) Distribution of dengue-2 antigens by electron immunocytochemistry. Infect Immun 13: 1699–1709
Deubel V, Kinney RM, Trent DW (1988) Nucleotide sequence and deduced amino acid sequence of the nonstructural proteins of dengue type 2 virus, Jamaica genotype: comparative analysis of the full-length genome. Virology 165: 234–244
Fong MY, Koh CL, Samuel S, Pang T, Lam SK (1990) Nucleotide sequences of the nonstructural protein NS 1 gene of three dengue-2 viruses, M 1, M 2, and M 3, isolated in Malaysia from patients with dengue haemorrhagic fever, dengue shock syndrome and dengue fever, respectively. Nucleic Acids Res 18: 1642
Gibbs AJ, Air GM, Laver WG (1982) Analysis of variation among haemagglutinin genes of influenza A viruses. In: Mackenzie JS (ed) Viral diseases in South-East Asia and the Western Pacific. Academic Press, New York, pp 546–549
Hahn YS, Galler R, Hunkapiller T, Dalrymple JM, Strauss JH, Strauss EG (1988) Nucleotide sequence of dengue 2 RNA and comparison of the encoded proteins with those of other flaviviruses. Virology 162: 167–180
Halstead SB, Simasthien P (1970) Observations related to the pathogeneis of dengue hemorrhagic fever II. Antigenic and biologic properties of dengue viruses and their association with disease response in the host. Yale J Biol Med 42: 276–292
Henchal EA, Henchal LS, Schlesinger JJ (1988) Synergistic interactions of anti-NS 1 monoclonal antibodies protect passively immunized mice from lethal challenge with dengue 2 virus. J Gen Virol 69: 2101–2107
Igarashi A (1978) Isolation of a Singh'sAedes albopictus cell clone sensitive to dengue and Chikungunya viruses. J Gen Virol 40: 531–544
Irie K, Mohan PM, Sasaguri Y, Putnak R, Padmanabhan R (1989) Sequence analysis of cloned dengue virus type 2 genome (New Guinea-C strain). Gene 75: 197–211
Jin L, Nei M (1990) Limitations of the evolutionary parsimony method of phylogenetic analysis. Mol Biol Evol 7: 82–102
Kerschner JAH, Vorndam AV, Monath TP, Trent DW (1986) Genetic and epidemiological studies of dengue type 2 viruses by hybridization using synthetic deoxyoligonucleotides as probes. J Gen Virol 67: 2645–2661
Li W-H, Wu C-I, Luo C-C (1984) Non randomness of point mutations is reflected by nucleotide substitutions in pseudogenes and its evolutionary implications. J Mol Evol 21: 58–71
Mackow E, Makino Y, Zhao B, Zhang Y-M, Markoff L, Buckler-White A, Guiler M, Chanock R, Lai C-J (1987) The nucleotide sequence of dengue type 4 virus: analysis of genes coding for nonstructural proteins. Virology 159: 217–228
Mason PW, McAda PC, Mason TL, Fournier MJ (1987) Sequence of the dengue-1 genome in the region encoding the three structural proteins and the major nonstructural protein NS 1. Virology 161: 262–267
Monath TP, Wands JR, Hill LJ, Brown NV, Marciniak RA, Wong MA, Gentry MK, Burke DS, Grant JA, Trent DW (1986) Geographic classification of dengue-2 virus strains by antigen signature analysis. Virology 154: 313–324
Putnak JR, Charles PC, Padmanabhan R, Irie K, Hoke CH, Burke DS (1988) Functional and antigenic domains of the dengue-2 virus nonstructural glycoprotein NS 1. Virology 163: 93–103
Repik PM, Dalrymple JM, Brandt WE, McCown JM, Russell PK (1983) RNA fingerprinting as a method for distinguishing dengue 1 virus strains. Am J Trop Med Hyg 32: 577–589
Rice CM, Strauss EG, Strauss JH (1986) Structure of the flavivirus genome. In: Schlesinger S, Schlesinger M (eds) The Togaviridae and Flaviviridae. Plenum, New York, pp 279–327
Rico-Hesse R (1990) Molecular evolution and distribution of dengue viruses type 1 and 2 in nature. Virology 174: 479–493
Rohlf FJ (1989) NTSYS-pc: numerical taxonomy and multivariant analysis system. Exiter, New York
Rohlf FJ, Wooten MC (1988) Evaluation of the restricted maximum-likelihood method for estimating phylogenetic trees using simulated allele-frequency data. Evolution 42: 581–595
Saitou N, Nei M (1987) The neighbour-joining method: a new method for reconstructing phylogentic trees. Mol Biol Evol 4: 406–425
Sanger F, Nicklen S, Coulson AR (1977) DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci USA 74: 5463–5467
Sangkhawibha N (1982) Viral diseases in Thailand — a national overview. In: Mackenzie JS (ed) Viral diseases in South-East Asia and the Western Pacific. Academic Press, New York, pp 217–223
Schlesinger JJ, Brandiss MW, Walsh EE (1987) Protection of mice against dengue-2 virus encephalitis by immunization with dengue-2 non-structural glycoprotein NS 1. J Gen Virol 68: 853–857
Schlesinger RW (1977) Dengue viruses. Springer, Wien New York [Gard S, Hallauer C (eds) Virology monographs, vol 16]
Trent DW, Grant JA, Rosen L, Monath TP (1983) Genetic variation among dengue 2 viruses of different geographic origin. Virology 128: 271–284
Trent DW, Grant JA, Monath TP, Manske CL, Corina M, Fox GE (1989) Genetic variation and microevolution of dengue 2 virus in Southeast Asia. Virology 172: 523–535
Walker PJ, Henchal EA, Blok J, Repik PM, Henchal LS, Burke DS, Robbins SJ, Gorman BM (1988) Variation in dengue type 2 viruses isolated in Bangkok during 1980. J Gen Virol 69: 591–602
Westaway EG, Brinton MA, Gaidamovich SYa, Horzinek MC, Igarashi A, Kääriäinen L, Lvov DK, Porterfield JS, Russell PK, Trent DW (1985) Flaviviridae. Intervirology 24: 183–192
World Health Organization (1975) Technical guides for diagnosis, treatment, surveillance, prevention and control of dengue hemorrgagic fever. WHO, Geneva
Zhao B, Mackow E, Buckler-White A, Markoff L, Chanock RM, Lai C-J, Makino Y (1986) Cloning full-length dengue 4 viral DNA sequences: analysis of genes coding for structural proteins. Virology 155: 77–88
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Blok, J., Gibbs, A.J., McWilliam, S.M. et al. NS 1 gene sequences from eight dengue-2 viruses and their evolutionary relationships with other dengue-2 viruses. Archives of Virology 118, 209–223 (1991). https://doi.org/10.1007/BF01314031
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF01314031