Summary
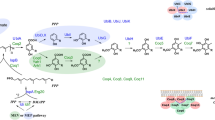
Coenzyme Q (or ubiquinone) is the product of two distinct biosynthetic pathways: the lipid “tail” of coenzyme Q is formed via the isoprene biosynthetic pathway, and the quinone ring derives from the metabolism of either shikimic acid or tyrosine. In general, eukaryotic organisms use the classical mevalonate pathway to form isopentenyl- and dimethylallyl-diphosphate, the five carbon building blocks of the polyisoprenoid tail, and prokaryotes use 1-deoxy-D-xylulose-5-phosphate, formed via the Rohmer pathway. The quinone ring precursor is 4-hydroxybenzoic acid, which is formed directly from chorismate inSaccharomyces cerevisiae andEscherichia coli, or from tyrosine in animal cells. Ring modification steps including prenylation, decarboxylation, and successive hydroxylation and methylation steps form the fully substituted benzoquinone ring of coenzyme Q. Many of the genes and polypeptides involved in coenzyme Q biosynthesis have been isolated and characterized by utilizing strains ofE. coli andS. cerevisiae with mutations in theubi andCOQ genes, respectively. This article reviews recent progress in characterizing the biosynthesis of coenzyme Q inE. coli, S. cerevisiae, and other eukaryotic organisms.
Similar content being viewed by others
References
Alam SS, Nambudiri AMD, Rudney H (1975) 4-Hydroxybenzoate:polyprenyl transferase and the prenylation of 4-aminobenzoate in mammalian tissues. Arch Biochem Biophys 171: 183–190
Ashby MN, Edwards PA (1990) Elucidation of the deficiency in two yeast coenzyme Q mutants: characterization of the structural gene encoding hexaprenyl pyrophosphate synthetase. J Biol Chem 265: 13157–13164
—, Kutsunai SY, Ackerman S, Tzagoloff A, Edwards PA (1992)COQ2 is a candidate for the structural gene encodingpara- hydroxybenzoate:polyprenyltransferase. J Biol Chem 267: 4128–4136
Avelange-Macherel M-H, Joyard J (1998) Cloning and functional expression ofAtCOQ3, theArabidopsis homologue of the yeastCOQ3 gene, encoding a methyltransferase from plant mitochondria involved in ubiquinone biosynthesis. Plant J 14: 203–212
Barkovich RJ, Shtanko A, Shepherd JA, Lee PT, Myles DC, Tzagoloff A, Clarke CF (1997) Characterization of theCOQ5 gene fromSaccharomyces cerevisiae: evidence for a C-methyltransferase in ubiquinone biosynthesis. J Biol Chem 272: 9182–9188
Begley TP, Kinsland C, Taylor S, Tandon M, Nicewonger R, Wu M, Chiu H-J, Kelleher N, Campobasso N, Zhang Y (1998) Cofactor biosynthesis: a mechanistic perspective. Topics Curr Chem 195: 93–142
Boitier E, Degoul F, Desguerre I, Charpentier C, Francois D, Ponsot G, Diry M, Rustin P, Marsac C (1998) A case of mitochondrial encephalomyopathy associated with a muscle coenzyme Q10 deficiency. J Neurol Sci 156: 41–46
Chan A, Reichmann H, Kogel A, Beck A, Gold R (1998) Metabolic changes in patients with mitochondrial myopathies and effects of coenzyme Q10 therapy. J Neurol 245: 681–685
Clarke CF, Williams W, Teruya JH (1991) Ubiquinone biosynthesis inSaccharomyces cerevisiae: isolation and sequence ofCOQ3, the 3,4-dihydroxy-5-hexaprenylbenzoate (DHHB) methyltransferase gene. J Biol Chem 266: 16636–16644
Clausen M, Lamb CJ, Megnet R, Doerner PW (1994)PAD1 encodes phenylacrylic acid decarboxylase which confers resistance to cinnamic acid inSaccharomyces cerevisiae. Gene 142: 107–112
Cox GB, Downie JA (1979) Isolation and characterization of mutants ofEscherichia coli K-12 affected in oxidative phophorylation or quinone biosynthesis. Methods Enzymol 56: 106–117
—, Young IG, McCann LM, Gibson F (1969) Biosynthesis of ubiquinone inEscherichia coli K-12: location of genes affecting the metabolism of 3-octaprenyl-4-hydroxybenzoic acid and 2-octaprenylphenol. J Bacteriol 99: 450–458
Daniels DL, Plunkett G, Burland V, Blattner FR (1992) Analysis of theEscherichia coli genome: DNA sequence of the region from 84.5 to 86.5 minutes. Science 257: 771–778
Dibrov E, Robinson KM, Lemire BD (1997) TheCOQ5 gene encodes a yeast mitochondrial protein necessary for ubiquinone biosynthesis and the assembly of the respiratory chain. J Biol Chem 272: 9175–9181
Do TQ, Schultz JR, Clarke CF (1996) Enhanced sensitivity of ubiquinone deficient mutants ofSaccharomyces cerevisiae to products of autooxidized polyunsaturated fatty acids. Proc Natl Acad Sci USA 93: 7534–7539
Eggink G, Engel H, Vriend G, Terpstra P, Witholt B (1990) Rubredoxin reductase ofPseudomonas oleovorans: structural relationship to other flavoprotein oxidoreductases based on one NAD and two FAD fingerprints. J Mol Biol 212: 135–142
Eppink MHM, Schreuder HA, Van Berkel WJH (1997) Identification of a novel conserved sequence motif in flavoprotein hydroxylases with a putative dual function in FAD/NAD(P)H binding. Protein Sci 6: 2454–2458
Ewbank JJ, Barnes TM, Lakowski B, Lussier M, Bussey H, Hekimi S (1997) Structural and functional conservation of theCaenorhabditis elegans timing geneclk-1. Science 275: 980–983
Friis P, Daves GD, Folkers K (1966) Complete sequence of biosynthesis fromp-hydroxybenzoic acid to ubiquinone. J Am Chem Soc 88: 4754–4756
Gancedo JM (1998) Yeast carbon catabolite repression. Microbiol Mol Biol Rev 62: 334–361
Gibson F (1973) Chemical and genetic studies on the biosynthesis of ubiquinone byEscherichia coli. Biochem Soc Trans 1: 317–326
—, Young IG (1978) Isolation and characterization of intermediates in ubiquinone biosynthesis. Methods Enzymol 53: 600–609
Gibson MI, Gibson F (1964) Preliminary studies on the isolation and metabolism of an intermediate in aromatic biosynthesis: chorismic acid. Biochem J 90: 248–256
Glerum DM, Muroff I, Jin C, Tzagoloff A (1997)COX15 codes for a mitochondrial protein essential for the assembly of yeast cytochrome oxidase. J Biol Chem 272: 19088–19094
Goewert RR (1980) Studies on the biosynthesis of ubiquinone. PhD dissertation, Saint Louis University, Saint Louis, Mo
—, Sippel CJ, Olson RE (1981a) Identification of 3,4-dihydroxy-5-hexaprenylbenzoic acid as an intermediate in the biosynthesis of ubiquinone-6 bySaccharomyces cerevisiae. Biochemistry 20: 4217–4223
— —, Grimm MF, Olson RE (1981b) Identification of 3-methoxy-4-hydroxy-5-hexaprenylbenzoic acid as a new intermediate in ubiquinone biosynthesis bySaccharomyces cerevisiae. Biochemistry 20: 5611–5616
Goldstein JL, Brown MS (1990) Regulation of the mevalonate pathway. Nature 343: 425–430
Hamilton JA, Cox GB (1971) Ubiquinone biosynthesis inEscherichia coli K-12: accumulation of an octaprenol, farnesylfarnesylgeraniol, by a multiple aromatic auxotroph. Biochem J 123: 435–443
Howlett BJ, Bar-Tana J (1980) Polyprenylp-hydroxybenzoate carboxylyase in flagellation ofSalmonella typhimurium. J Bacteriol 143: 644–651
Hsu A, Poon WW, Shepherd JA, Myles DC, Clarke CF (1996) Complementation ofcoq3 mutant yeast by mitochondrial targeting of theE. coli UbiG polypeptide: evidence that UbiG catalyzes both O-methylation steps in ubiquinone biosynthesis. Biochemistry 35: 9797–9806
—, Do TQ, Lee PT, Clarke CF (2000) Genetic evidence for a multisubunit complex in the O-methyl-transferase steps of coenzyme Q biosynthesis. Biochim Biophys Acta 1484: 287–297
Jonassen T, Clarke CF (2000) Isolation and functional expression of humanCOQ3, a gene encoding a methyltransferase required for ubiquinone biosynthesis. J Biol Chem 275: 12381–12387
—, Marbois BN, Kirn L, Chin A, Xia Y-R, Lusis AJ, Clarke CF (1996) Isolation and sequencing of the ratCOQ7 gene and the mapping of mouseCOQ7 to chromosome 7. Arch Biochem Biophys 330: 285–289
—, Proft M, Randez-Gil F, Schultz JR, Marbois BN, Entian K-D, Clarke CF (1998) Yeast Clk-1 homologue (Coq/Cat5): a mitochondrial protein in ubiquinone synthesis. J Biol Chem 273: 3351–3357
Kagan RM, Clarke S (1994) Widespread occurrence of three sequence motifs in diverse S-adenosylmethionine-dependent methyltransferases suggests a common structure for these enzymes. Arch Biochem Biophys 310: 417–427
Kalen A, Norling B, Appelkvist EL, Dallner G (1987) Ubiquinone biosynthesis by the microsomal fraction from rat liver. Biochim Biophys Acta 926: 70–78
—, Appelkvist E-L, Chojnacki T, Dallner G (1990) Nonaprenyl-4-hydroxybenzoate transferase, an enzyme ivolved in ubiquinone biosynthesis, in the endoplasmic reticulum-Golgi system of rat liver. J Biol Chem 265: 1158–1164
Kobayashi T, Kishigami S, Sone M, Inokuchi H, Mogi T, Ito K (1997) Respiratory chain is required to maintain oxidized states of the DsbA-DsbB disulfide bond formation system in aerobically growingEscherichia coli cells. Proc Natl Acad Sci USA 94: 11857–11862
Lawrence J, Cox GB, Gibson F (1974) Biosynthesis of ubiquinone inEscherichia coli K-12: biochemical and genetic characterization of a mutant unable to convert chorismate into 4-hydroxybenzoate. J Bacteriol 118: 41–45
Lee PT, Hsu AY, Ha HT, Clarke CF (1997) A C-methyltransferase involved in ubiquinone biosynthesis: isolation and identification of theEscherichia coli ubiE gene. J Bacteriol 179: 1748–1754
Leppik RA, Stroobant P, Shineberg B, Young IG, Gibson F (1976a) Membrane associated reactions in ubiquinone biosynthesis: 2-octaprenyl-3-methyl-5-hydroxy-6-methoxy-1,4-benzoquinone methyltransferase. Biochim Biophys Acta 428: 146–156
—, Young IG, Gibson F (1976b) Membrane-associated reactions in ubiquinone biosynthesis inEscherichia coli: 3-octaprenyl-4-hydroxybenzoate carboxy-lyase. Biochim Biophys Acta 436: 800–810
Lichtenthaler HK, Schwender J, Disch A, Rohmer M (1997) Biosynthesis of isoprenoids in higher plant chloroplasts proceeds via a mevalonate-independent pathway. FEBS Lett 400: 271–274
Loscher R, Heide L (1994) Biosynthesis ofp-hydroxybenzoate fromp-coumarate andp-coumaroyl-coenzyme A in cell-free extracts ofLithospermum erythrorhizon cell cultures. Plant Physiol 106: 271–279
Macinga DR, Cook GM, Poole RK, Rather PN (1998) Identification and characterization ofaarF, a locus required for production of ubiquinone inProvidencia stuartii andEscherichia coli and for expression of 2′-N-acetyltransferase inP. stuartii. J Bacteriol 180: 128–135
Marbois BN, Clarke CF (1996) TheCOQ7 gene encodes a protein inSaccharomyces cerevisiae necessary for ubiquinone biosynthesis. J Biol Chem 271: 2995–3004
—, Hsu A, Pillai R, Colicelli J, Clarke CF (1994a) Cloning of a rat cDNA encoding dihydroxypolyprenylbenzoate methyltransferase by functional complementation of aSaccharomyces cerevisiae mutant deficient in ubiquinone biosynthesis. Gene 138: 213–217
—, Xia Y-R, Lusis AJ, Clarke CF (1994b) Ubiquinone biosynthesis in eukaryotic cells: tissue distribution of mRNA encoding 3,4-dihydroxy-5-polyprenylbenzoate methyltransferase in the rat and mapping of theCOQ3 gene to mouse chromosome 4. Arch Biochem Biophys 313: 83–88
Matthews RT, Yang L, Browne S, Baik M, Beal MF (1998) Coenzyme Q10 administration increases brain mitochondrial concentrations and exerts neuroprotective effects. Proc Natl Acad Sci USA 95: 8892–8897
Meganathan R (1996) Biosynthesis of the isoprenoid quinones menaquinone (vitamin K2) and ubiquinone (coenzyme Q). In: Neidhardt FC (ed)Escherichia coli andSalmonella: cellular and molecular biology, vol 1. American Society for Microbiology Press, Washington, DC, pp 642–656
Melzer M, Heide L (1994) Characterization of polyprenyldiphosphate:4-hydroxybenzoate polyprenyltransferase fromEscherichia coli. Biochim Biophys Acta 1212: 93–102
Momose K, Rudney H (1972) 3-Polyprenyl-4-hydroxybenzoate synthesis in the inner membrane of mitochondria fromp-hydroxybenzoate and isopentenylpyrophosphate. J Biol Chem 247: 3930–3940
Murakami S, Johnson TE (1996) A genetic pathway conferring life extension and resistance to UV stress inCaenorhabditis elegans. Genetics 143: 1207–1218
Nakahigashi K, Miyamoto K, Nishimura K, Inokuchi H (1992) Isolation and characterization of a light-sensitive mutant ofEscherichia coli K-12 with a mutation in a gene that is required for the biosynthesis of ubiquinone. J Bacteriol 174: 7352–7359
Nambudiri AMD, Brockman D, Alam SS, Rudney H (1977) Alternate routes for ubiquinone bisoynthesis. Biochem Biophys Res Commun 76: 282–288
Nichols BP, Green JM (1992) Cloning and sequencing ofEscherichia coli ubiC and purification of chorismate lyase. J Bacteriol 174: 5309–5316
Nishimura K, Nakahigashi K, Inokuchi H (1992) Location of theubiA gene on the physical map ofEscherichia coli. J Bacteriol 174: 5762
Ogasahara S, Engel AG, Frens D, Mack D (1989) Muscle coenzyme Q deficiency in familial mitochondrial encephalomyopathy. Proc Natl Acad Sci USA 86: 2379–2382
Ogura K, Koyama T (1998) Enzymatic aspects of isoprenoid chain elongation. Chem Rev 98: 1263–1276
Okada K, Suzuki K, Kamiya Y, Zhu X, Fujisaki S, Nishimura Y, Nishino T, Nakagawa T, Kawamukai M, Matsuda H (1996) Polyprenyl diphosphate synthase essentially defines the length of the side chain of ubiquinone. Biochim Biophys Acta 1302: 217–223
—, Kamiya Y, Zhu X, Suzuki K, Tanaka K, Nakagawa T, Matsuda H, Kawamukai M (1997a) Cloning of thesdsA gene encoding solanesyl diphosphate synthase fromRhodobacter capsulatus and its functional expression inEscherichia coli andSaccharomyces cerevisiae. J Bacteriol 179: 5992–5998
—, Minehira M, Zhu X, Suzuki K, Nakagawa T, Matsuda H, Kawamukai M (1997b) Theisp B gene encoding octaprenyl diphosphate synthase is essential for growth ofEscherichia coli. J Bacteriol 179: 3058–3060
—, Kainou T, Matsuda H, Kawamukai M (1998a) Biological significance of the side chain length of ubiquinone inSaccharomyces cerevisiae. FEBS Lett 431: 241–244
— —, Tanaka K, Nakagawa T, Matsuda H, Kawamukai M (1998b) Molecular cloning and mutational analysis of theddsA gene encoding decaprenyl diphosphate synthase fromGluconobacter suboxydans. Eur J Biochem 255: 52–59
Olson RE (1966) Biosynthesis of ubiquinones in animals. Vitamin Horm 24: 551–573
—, Rudney H (1983) Biosynthesis of ubiquinone. Vitamin Horm 40: 1–43
Palfey BA, Ballou DP, Massey V (1995) Oxygen activation by flavins and pterins. In: Valentine JS, Foote CS, Greenberg A, Liebman JF (eds) Active oxygen in biochemistry. Blackie Academic and Professional Press, Glasgow, pp 37–83
Parmryd I, Dallner G (1996) Organization of isoprenoid biosynthesis. Biochem Soc Trans 24: 677–682
Pennock JF, Threlfall DR (1983) Biosynthesis of ubiquinone and related compounds. In: Porter JW, Spurgeon SL (eds) Biosynthesis of isoprenoid compounds. Wiley, New York, pp 191–302
Poon WW, Marbois BN, Faull K, Clarke CF (1995) 3-Hexaprenyl-4-hydroxybenzoic acid forms a predominant intermediate pool in ubiquinone biosynthesis inSaccharomyces cerevisiae. Arch Biochem Biophys 320: 305–314
—, Do TQ, Marbois BN, Clarke CF (1997) Sensitivity to treatment with polyunsaturated fatty acids is a general characteristic of the ubiquinone-deficient yeastcoq mutants. Mol Aspects Med 18: S121-S127
—, Barkovich RJ, Hsu AY, Frankel A, Lee PT, Shepherd JN, Myles DC, Clarke CF (1999) Yeast and Rat Coq3 andE. coli UbiG catalyze both O-methyltransferase steps in coenzyme Q biosynthesis. J Biol Chem 274: 21665–21672
- Davis DE, Ha HT, Jonassen T, Rather PN, Clarke CF (2000) Identification ofE. coli ubiB: a gene required for the first monooxygenase step in ubiquinone biosynthesis. J Bacteriol (in press)
Proft M, Kotter P, Hedges D, Bojunga N, Entian KD (1995)CAT5, a new gene necessary for derepression of gluconeogenic enzymes inSaccharomyces cerevisiae. EMBO J 14: 6116–6126
Ranganathan S, Ramasarma T (1974) The metabolism of phenolic acids in the rat. Biochem J 140: 517–522
Rohmer RH, Seemann M, Horbach S, Bringer-Meyer S, Sahm H (1996) Glyceraldehyde 3-phosphate and pyruvate as precursors of isoprenic units in an alternative non-mevalonate pathway for terpenoid biosynthesis. J Am Chem Soc 118: 2564–2566
Rowland MA, Nagley P, Linnane AW, Rosenfeldt FL (1998) Coenzyme Q10 treatment improves the tolerance of the senescent myocardium to pacing stress in the rat. Cardiovasc Res 40: 165–173
Santos-Ocana C, Cordoba F, Crane FL, Clarke CF, Navas P (1998a) Coenzyme Q6 and iron reduction are responsible for the extracellular ascorbate stabilization at the plasma membrane ofSaccharomyces cerevisiae. J Biol Chem 273: 8099–8105
—, Villalba JM, Cordoba F, Padilla S, Crane FL, Clarke CF, Navas P (1998b) Genetic evidence for coenzyme Q requirement in plasma membrane electron transport. J Bioenerg Biomembr 30: 465–474
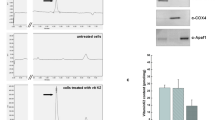
Schnitzler J-P, Madlung J, Rose A, Scitz HU (1992) Biosynthesis ofp-hydroxybenzoic acid in elicitor-treated carrot cell cultures. Plant 188: 594–600
Schultz JR, Clarke CF (199a) Functional roles of ubiquinone. In: Cadenas E, Packer L (eds) Mitochondria, oxidants, and aging. Marcel Dekker, New York, pp 95–118
— — (1999b) Characterization ofSaccharomyces cerevisiae ubiquinone-deficient mutants. BioFactors 9: 121–129
Shepherd JA, Poon WW, Myles DC, Clarke CF (1996) The biosynthesis of ubiquinone: synthesis and enzymatic modification of biosynthetic precursors. Tetrahedron Lett 37: 2395–2398
Siebert M, Bechthold A, Melzer M, May U, Berger U, Schroder G, Schroder J, Severin K, Heide L (1992) Ubiquinone biosynthesis: cloning of the genes coding for chorismate pyruvate-lyase and 4-hydroxybenzoate octaprenyl transferase fromEscherichia coli. FEBS Lett 307: 347–350
—, Severin K, Heide L (1994) Formation of 4-hydroxybenzoate inEscherichia coli: characterization of theubiC gene and its encoded enzyme chorismate pyruvate-lyase. Microbiol 140: 897–904
Sirlin JL (1956)Vacillans, a neurological mutant in the house mouse linked withbrown. J Genet 54: 42–48
Soja AM, Mortensen SA (1997) Treatment of congestive heart failure with coenzyme Q10 illuminated by meta-analyses of clinical trials. Mol Aspects Med 18: S159-S168
Stroobant P, Young IG, Gibson F (1972) Mutants inEscherichia coli K-12 blocked in the final reaction of ubiquinone biosynthesis: characterization and genetic analysis. J Bacteriol 109: 134–139
Suzuki K, Ueda M, Yuasa M, Nakagawa T, Kawamukai M, Matsuda H (1994) Evidence thatEscherichia coli ubiA product is a functional homolog of yeastCOQ2, and the regulation ofubiA gene expression. Biosci Biotech Biochem 58: 1814–1819
—, Okada K, Kamiya Y, Zhu XF, Nakagawa T, Kawamukai M, Matsuda H (1997) Analysis of the decaprenyl diphosphate synthase (dps) gene in fission yeast suggests a role of ubiquinone as an antioxidant. J Biochem 121: 496–505
Teclebrhan H, Jakobsson-Borin A, Brunk U, Dallner G (1995) Relationship between the endoplasmic reticulum-Golgi membrane system and ubiquinone biosynthesis. Biochim Biophys Acta 1256: 157–165
Tzagoloff A, Dieckmann CL (1990)PET genes ofSaccharomyces cerevisiae. Microbiol Rev 54: 211–225
—, Akai A, Needleman RB (1975a) Assembly of the mitochondrial membrane system: isolation of nuclear and cytoplasmic mutants ofSaccharomyces cerevisiae with specific defects in mitochondrial functions. J Bacteriol 122: 826–831
—, Akai A, Needleman RB (1975b) Assembly of the mitochondrial membrane system: characterization of nuclear mutants ofSaccharomyces cerevisiae with defects in mitochondrial ATPase and respiratory enzymes. J Biol Chem 250: 8228–8235
—, Yue J, Jang J, Paul M-F (1994) A new member of a family of ATPases is essential for assembly of mitochondrial respiratory chain and ATP synthetase complexes inSaccharomyces cerevisiae. J Biol Chem 269: 26144–26151
Vajo Z, King LM, Jonassen T, Wilkin DJ, Ho N, Munnich A, Clarke CF, Francomano CA (1999) Conservation of theC. elegans timing geneclk-1 from yeast to human: a gene required for ubiquinone biosynthesis with potential implications for aging. Mammalian Genome 10: 1000–1004
Vidgren J, Svensson LA, Liijas A (1994) Crystal structure of catechol O-methyltransferase. Nature 368: 354–358
Walker GA, Walker DW, Lithgow GJ (1998) A relationship between thermotolerance and longevity inCaenorhabditis elegans. J Invest Dermatol Symp Proc 3: 6–10
Wallace BJ, Young IG (1977) Role of quinones in electron transport to oxygen and nitrate inEscherichia coli: studies with aubiA − menA − double quinone mutant. Biochim Biophys Acta 461: 84–100
White RH (1996) Biosynthesis of isoprenoids in bacteria. In: Neidhardt FC (ed)Escherichia coli andSalmonella: cellular and molecular biology, vol 1. American Society for Microbiology Press, Washington, DC, pp 637–641
Wierenga RK, Terpstra P, Hol WGJ (1986) Prediction of the occurrence of the ADP-binding βαβ-fold in proteins, using an amino acid sequence fingerprint. J Mol Biol 187: 101–107
Wissenbach U, Ternes D, Unden G (1992) AnEscherichia coli mutant containing only demethylmenaquinone but no menaquinone: effects on fumarate, dimethylsulfoxide, trimethylamine N-oxide and nitrate respiration. Arch Microbiol 158: 68–73
Wong A, Boutis P, Hekimi S (1995) Mutations in theclk-1 gene ofCaenorhabditis elegans affect developmental and behavioral timing. Genetics 139: 1247–1259
Wu G, Williams HD, Zamanian M, Gibson F, Poole RK (1992) Isolation and characterization ofEscherichia coli mutants affected in aerobic respiration: the cloning and nucleotide sequence ofubiG. J Gen Microbiol 138: 2101–2112
— —, Gibson F, Poole RK (1993) Mutants ofEscherichia coli affected in respiration: the cloning and nucleotide sequence ofubiA, encoding the membrane-bound p-hydroxybenzoate:octaprenyl-transferase. J Gen Microbiol 139: 1795–1805
Yazaki K, Heide L, Tabata M (1991) Formation ofp-hydroxybenzoic acid fromp-coumaric acid by cell free extract ofLithospermum erythrorhizon cell cultures. Phytochemistry 30: 2233–2236
Young IG, McCann IM, Stroobant P, Gibson F (1971) Characterization and genetic analysis of mutant strains ofEscherichia coli K-12 accumulating the ubiquinone precursors 2-octaprenyl-6-methoxy-1,4-benzoquinone and 2-octaprenyl-3-methyl-6-methoxy-1,4-benzoquinone. J Bacteriol 105: 679–778
—, Stroobant P, MacDonald CG, Gibson F (1973) Pathway for ubiquinone biosynthesis inEscherichia coli K-12: gene-enzyme relationships and intermediates. J Bacteriol 114: 42–52
Zeng H, Snavely I, Zamorano P, Javor GT (1998) Low ubiquinone content inEscherichia coli causes thiol hypersensitivity. J Bacteriol 180: 3681–3685
Zhang Y, Aberg F, Appelkvist E-L, Dallner G, Ernster L (1995) Uptake of dietary coenzyme Q supplement is limited in rats. J Nutr 125: 446–453
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Clarke, C.F. New advances in coenzyme Q biosynthesis. Protoplasma 213, 134–147 (2000). https://doi.org/10.1007/BF01282151
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF01282151