Abstract
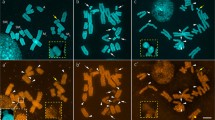
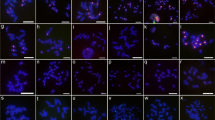
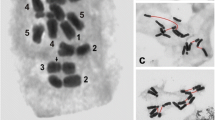
A satellite sequence repeat ofAllium cepa was tested by fluorescent in situ hybridization (FISH) for cross-hybridization to chromosomes of 27 species (in 37 accessions) belonging to 14 sections of four subgenera ofAllium. All investigated species of sect.Cepa, with the two subsects.Cepa andPhyllodolon, revealed clear satellite-specific hybridization signals mainly at their chromosome termini. The tested species belonging to other sections/subgenera revealed no hybridization signals. An exception wasA. roylei, assigned to sect.Oreiprason. Its chromosomes also showed strong terminal hybridization signals. This and other features suggest a close relationship ofA. roylei to the species of sect.Cepa in spite of deviating morphological characters. The divergence between the satellite repeats to species to which theA. cepa repeat cross-hybridized was determined and revealed high degrees of similarity. Therefore, we conclude that this satellite sequence had evolved already in progenitor forms of sect.Cepa and remained unusually well conserved during speciation. This might indicate selection pressure exerted on a secondarily acquired telomere function of the satellite sequence.
Similar content being viewed by others
References
Bark, O. H., Havey, M. J., 1994: Similarities and relationships among populations of the bulb onion as estimated by nuclear RFLPs. — Theor. Appl. Genet.90: 407–414.
Barnes, S. R., James, A. M., Jamieson, G., 1985: The organisation, nucleotide sequence, and chromosomal distribution of a satellite DNA fromAllium cepa. — Chromosoma92: 185–192.
McCollum, G., 1974: Chromosome behavior and sterility of hybrids between the common onion,Allium cepa, and the related wildA. oschaninii. — Euphytica23: 699–709.
Friesen, N. V., 1988: Lukovye Sibiri. Sistematika, kariologija, chorologija. — Novosibirsk: Nauka.
Fuchs, J., Schubert, I., 1995: Localization of seed protein genes on metaphase chromosomes ofVicia faba via fluorescent in situ hybridization. — Chromosome Res.3: 94–100.
—, 1995: Telomere sequence localization and karyotype evolution in higher plants. — Pl. Syst. Evol.196: 227–241.
Hanelt, P., 1985: Zur Taxonomie, Chorologie und Ökologie der Wildarten vonAllium L., sect.Cepa (Mill.)Prokh. — Flora176: 99–116.
—, 1990: Taxonomy, evolution, and history. — InRabinowitch, H., Brewster, J., (Eds): Onions and allied crops1. — Boca Raton: CRC Press.
—, 1992: Infrageneric grouping ofAllium — the Gatersleben Approach. — InHanelt, P., Hammer, K., Knüpffer, H., (Eds): The genusAllium — taxonomic problems and genetic resources, pp. 107–123. — Gatersleben, Germany: IPK.
Havey, M. J., 1991: Phylogenetic relationships among cultivatedAllium species from restriction enzyme analysis of the chloroplast genome. — Theor. Appl. Genet.81: 752–757.
—, 1992a: Restriction enzyme analysis of the nuclear 45s ribosomal DNA of six cultivated Alliums (Alliaceae). — Pl. Syst. Evol.181: 45–55.
—, 1992b: Restriction enzyme analysis of the chloroplast and nuclear 45s ribosomal DNA ofAllium sectionsCepa andPhylodollon (Alliaceae). — Pl. Syst. Evol.183: 17–31.
Hatch, F. T., Bodner, A. J., Mazrimas, J. A., Moore, D. H., 1976: Satellite DNA and cytogenetic evolution: DNA quantity, satellite DNA and karyotypic variation in kangaroo rats (genusDipodomys). — Chromosoma58: 155–168.
Irifune, K., Hirai, K., Zheng, J., Tanaka, R., Morikawa, H., 1995: Nucleotide sequence of a highly repeated DNA sequence and its chromosomal localization inAllium fistulosum. — Theor. Appl. Genet.90: 312–316.
Jones, J. D. G., Flavell, R. B., 1982: The structure, amount, and chromosomal localisation of defined repeated DNA sequences in species of the genus Secale. — Chromosoma86: 613–641.
Jorgensen, R., Cluster, P., 1988: Modes and tempos in the evolution of nuclear ribosomal DNA: new characters for evolutionary studies and new markers for genetic and population studies. — Ann. Missouri Bot. Gard.75: 1238–1247.
Keller, E. R. J., Schubert, I., Fuchs, J., Meister, A., 1996: Interspecific crosses of onion with distantAllium species and characterization of the presumed hybrids by means of flow cytometry, karyotype analysis, and genomic in situ hybridization. — Theor. Appl. Genet. (in press).
Kofoet, A., Kik, W., Wietsma, W., de Vries, J., 1990: Inheritance of resistance of downy mildew fromAllium roylei Stearn in the backcrossAllium cepa L. × (A. roylei ×A. cepa). — Pl. Breed.105: 144–149.
Kruse, J., 1994: Rasterelektronenmikroskopische Untersuchungen an Samen der GattungAllium L. IV. — Feddes Repert.105: 457–471.
Labani, R. M., Elkington, T. T., 1987: Nuclear DNA variation in the genusAllium L. (Liliaceae). — Heredity59: 119–128.
Linne von Berg, G., Samoylov, A., Klaas, M., Hanelt, P., 1996: Chloroplast DNA restriction analysis and the infrageneric grouping ofAllium L. — Pl. Syst. Evol.200: 253–261.
Maggini, F., Carmona, M., 1981: Sequence heterogeneity of the ribosomal DNA inAllium cepa (Liliaceae). — Protoplasma108: 163–171.
Meer, Q. van der, Vries, J. de, 1990: An interspecific cross betweenAllium roylei Stearn andAllium cepa L. and its backcross toA. cepa. — Euphytica47: 29–31.
Nasir, E., 1975:Alliaceae. — InNasir, E., Ali, S. I., (Eds): Flora of West Pakistan83.
Pich, U., Schubert, I., 1993: Polymorphism of legumin genes in inbred lines ofVicia faba. — Biol. Zentralbl.112: 342–350.
-Fuchs, J., Schubert, I., 1996: How doAlliaceae stabilize their chromosome ends in the absence of TTTAGGG-sequences? — Chromosome Res. (in press).
Raamsdonk, L. W. D. van, Vries, T. de, 1992: Systematics and phylogeny ofAllium cepa L. and allies. — InHanelt, P., Hammer, K., Knüpffer, H., (Eds): The genusAllium. — Taxonomic problems and genetic resources, pp. 257–263. — Gatersleben, Germany: IPK.
Regel, E., 1875: Alliorum adhuc cognitorum monographia. — Acta Horti. Petrop.3 (II: 1–266.
Richards, E. J., Ausubel, F. M., 1988: Isolation of a higher eukaryotic telomere fromArabidopsis thaliana. — Cell53: 127–136.
Schaal, B., Learn, G., 1988: Ribosomal DNA variation within and among plant populations. — Ann. Missouri Bot. Gard.75: 1207–1216.
Schubert, I., Ohle, H., Hanelt, P., 1983: Phylogenetic conclusions from Giemsa banding and NOR staining in top onions (Liliaceae). — Pl. Syst. Evol.143: 245–256.
Thompson, R. D., Bartels, D., Haberd, N. P., Flavell, R. B., 1983: Characterization of the multigene family coding for HMW glutenin subunits in wheat using cDNA clones. — Theor. Appl. Genet.67: 87–96.
Vries, J. N. de, 1992: What's in a name? In quest of the origin and proper name of C502, known asAllium roylei Stearn. — InHanelt, P., Hammer, K., Knüpffer, H., (Eds): The genusAllium. — Taxonomic problems and genetic resources, pp. 327–330. — Gatersleben, Germany: IPK.
Wendelbo, P., 1971:Alliaceae. — InRechinger, K. H. Jr., (Ed.): Flora Iranica76, pp. 1–99. — Graz: Akadem. Druck- und Verlagsanstalt.
Author information
Authors and Affiliations
Additional information
Dedicated to Dr habil.Peter Hanelt on the occasion of his 65th birthday.
Rights and permissions
About this article
Cite this article
Pich, U., Fritsch, R. & Schubert, I. Closely relatedAllium species (Alliaceae) share a very similar satellite sequence. Pl Syst Evol 202, 255–264 (1996). https://doi.org/10.1007/BF00983386
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00983386