Summary
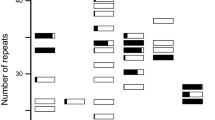
Allozyme surveys of cultivated plant species generally report little within-cultivar variation, but considerable among-cultivar variation. This trend contrasts with natural plant populations in which most allozyme variation resides within, rather than among, populations. The difference may be an artifact of the extreme inbreeding techniques used to develop and propagate these crops, rather than a consequence of domestication per se. To test this hypothesis, we compared the population genetic structure of 24 lines of radish cultivars — a domesticated species developed and maintained as open-pollinated, outcrossed populations — with four wild radish populations in California. Although the wild populations displayed more overall allozyme variation than the cultivars, most of the allozyme variation in the cultivars remains partitioned within, rather than among, lines. Apparently, how a crop is developed and maintained can have a profound influence on the organization of genetic variation of that species.
Similar content being viewed by others
References
Brown AHD (1978) Isozymes, plant population genetic structure and genetic conservation. Theor Appl Genet 52:145–157
Brown AHD (1979) Enzyme polymorphism in plant populations. Theor Popul Biol 15:1–42
Crisp P (1976) Trends in the breeding and cultivation of cruciferous crops. In: Vaughan JG, MacLeod AJ, Jones BMG (eds) The biology and chemistry of the Cruciferae. Academic Press, London New York, pp 69–117
Ellstrand NC (1984) Multiple paternity within fruits of wild radish,Raphanus sativus. Am Nat 123:819–828
George RAT, Evans DR (1981) A classification of winter radish cultivars. Euphytica 30:483–492
Heywood JS (1980) Genetic correlates of edaphic differentiation and endemism inGaillardia. PhD Dissertation, University of Texas, Austin
Levin DA (1976) Consequences of long-term artificial selection, inbreeding and isolation inPhlox. 2. The organization of allozymic variability. Evolution 30:463–472
Levin DA (1978) Genetic variation in annual Phlox: self-compatible versus self-incompatible species. Evolution 32:245–263
Nei M (1973) Analysis of gene diversity in subdivided populations. Proc Natl Acad Sci USA 70:3321–3323
Nei M (1978) Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 89:583–590
Panetsos CA, Baker HG (1967) The origin of variation in “wild”Raphanus sativus (Cruciferae) in California. Genetica 38:243–274
Pichersky E, Gottlieb LD (1983) Evidence for duplication of the structural genes coding plastid and cytosolic isozymes of triose phosphate isomerase in diploid species ofClarkia. Genetics 105:421–436
Shaw CR, Prasad P (1970) Starch gel electrophoresis of enzymes — a compilation of recipes. Biochem Genet 4:297–320
Simmonds NW (1979) Principles of crop improvement. Longman, New York
Sneath PHA, Sokal RR (1973) Numerical taxonomy. WH Freeman, San Francisco
Swofford DL, Selander RB (1981) Biosys-1: a FORTRAN program for the comprehensive analysis to electrophoretic data in population genetics and systematics. J Hered 72:281–283
Tanksley SD, Orton TJ (1983) Isozymes in plant genetics and breeding. Elsevier, Amsterdam
Torres AM (1983) Tree crops. In: Tanksley SD, Orton TJ (eds) Isozymes in plant genetics and breeding. Elsevier, Amsterdam
Wendel JF, Parks CR (1982) Genetic control of isozyme variation inCamellia japonica L. J Hered 73:197–204
Author information
Authors and Affiliations
Additional information
Communicated by P. M. A. Tigerstedt
Rights and permissions
About this article
Cite this article
Ellstrand, N.C., Marshall, D.L. The impact of domestication on distribution of allozyme variation within and among cultivars of radish,Raphanus sativus L.. Theoret. Appl. Genetics 69, 393–398 (1985). https://doi.org/10.1007/BF00570908
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00570908