Summary
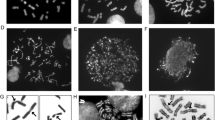
The AT specificity of the fluorochromes DIPI and DAPI and the GC specificity of mithramycin are evidenced by observations in human, mouse, and bovine chromosomes. DIPI and DAPI produce a pattern similar to Hoechst 33258 in all three species, whereas mithramycin results in a reverse pattern. The AT-rich centromeric heterochromatin in mouse is brilliantly stained by DIPI or DAPI and remains nearly invisible after mithramycin staining. In the GC-rich centromeric heterochromatin of cattle the opposite behavior is observed.
Similar content being viewed by others
References
Behr, W., Honikel, K., Hartmann, G.: Interaction of the RNA polymerase inhibitor chromomycin with DNA. Eur. J. Biochem. 9, 82–92 (1969)
Caspersson, T., Zech, L., Johansson, C., Modest, E. J.: Identification of human chromosomes by DNA-binding fluorescent agents. Chromosoma (Berl.) 30, 215–227 (1970)
Comings, D. E.: Mechanisms of chromosome banding. VIII. Hoechst 33 258-DNA interaction. Chromosoma (Berl.) 52, 229–243 (1975)
Comings, D. E., Kovács, B. W., Avelino, E., Harris, D. C.: Mechanisms of chromosome banding. V. Quinacrine banding. Chromosoma (Berl.) 50, 111–145 (1975)
Comings, D. E., Drets, M. E.: Mechanisms of chromosome banding. IX. Are variations in DNA base composition adequate to account for quinacrine, Hoechst 33 258 and daunomycin banding? Chromosoma (Berl.) 56, 199–211 (1976)
Dann, O., Bergen, G., Demant, E., Volz, G.: Trypanocide Diamine des 2-Phenyl-benzoforan, 2-Phenyl-inden und 2-Phenyl-indols. Liebigs Ann. Chem. 749, 68–89 (1971)
Distèche, C., Bontemps, J.: Chromosome regions containing DNAs of known base composition, specifically evidenced by 2,7-di-t-butyl proflavine. Comparison with the Q-banding and relation to dye-DNA interactions. Chromosoma (Berl.) 47, 263–281 (1974)
Distèche, C., Bontemps, J.: Method for the determination of mean densitometric profiles of chromosomes. Chromosoma (Berl.) 54, 39–59 (1976)
Dutrillaux, B., Lejeune, J.: Sur une nouvelle technique d'analyse du caryotype humain. C. R. Acad. Sci. (Paris) 272, série D, 2638–2640 (1971)
Filipski, J., Thiery, J.-P., Bernardi, G.: An analysis of the bovine genome by Cs2SO4−Ag+ density gradient centrifugation. J. Molec. Biol. 80, 177–197 (1973)
Flamm, W. G., Bernheim, N. J., Brubaker, P. E.: Density gradient analysis of newly replicated DNA from synchronized mouse lymphoma cells. Exp. Cell Res. 64, 97–104 (1971)
Hilwig, I., Gropp, A.: Staining of constitutive heterochromatin in mammalian chromosomes with a new fluorochrome. Exp. Cell Res. 75, 122–126 (1972)
Kurnit, D. M., Shafit, B. R., Maio, J. J.: Multiple satellite deoxyribonucleic acids in the calf and their relation to the sex chromosomes. J. molec. Biol. 81, 273–284 (1973)
Lin, C. C., van de Sande, J. H.: Differential fluorescent staining of human chromosomes with daunomycin and adriamycin—the D-bands. Science 190, 61–63 (1975)
Miller, O. J., Miller, D. A., Kouri, R. E., Alderdice, P. W., Dev, V. G., Grewal, M. S., Hutton, J. J.: Identification of the mouse karyotype by quinacrine fluorescence, and tentative assignment of seven linkage groups. Proc. Natl. Acad. Sci. (Wash.) 68, 1530–1533 (1971)
Schnedl, W.: Analysis of the human karyotype by the recent banding techniques. Archiv Genetik 46, 1–34 (1973)
Schnedl, W., Czaker, R.: Centromeric heterochromatin and comparison of G banding in cattle, goat and sheep chromosomes (Bovidae). Cytogenet. Cell Genet. 13, 246–255 (1974)
Schnedl, W., Mikelsaar, A.-V., Breitenbach, M., Dann, O.: DIPI and DAPI: Fluorescence banding with only negligible fading. Hum. Genet. 36 (1977)
Schreck, R. R., Erlanger, B. F., Miller, O. J.: The use of antinucleoside antibodies to probe the organization of chromosomes denatured by ultraviolet irradiation. Exp. Cell Res. 88, 31–39 (1974)
Schweizer, D.: Reverse fluorescent chromosome banding with chromomycin and DAPI. Chromosoma (Berl.) 58, 307–324 (1976a)
Schweizer, D.: DAPI fluorescence of plant chromosomes prestained with actinomycin D. Exp. Cell Res. 102, 408–412 (1976b)
Schweizer, D.: Giemsa and fluorochrome banding of polytene chromosomes in Phaseolus vulgaris and P. coccineus. In: Current chromosome research, K. Jones, P. E. Brandham, eds., pp. 51–56. Amsterdam: Elsevier/North Holland Biomedical Press 1976c
Schweizer, D., Nagl, W.: Heterochromatin diversity in Cymbidium, and its relationship to differential DNA replication. Exp. Cell Res. 98, 411–423 (1976)
Slater, M. L.: Rapid nuclear staining method for Saccharomyces cerevisiae. J. Bact. 126, 1339–1341 (1976)
Vosa, C. G.: Heterochromatin classification in Vicia faba and Scilla sibirica. Chromosomes today 5, 185–192 (1976)
Walker, P. M. B.: How different are the DNAs from related animals? Nature 219, 228–232 (1968)
Ward, D. C., Reich, E., Goldberg, I. H.: Base specificity in the interaction of polynucleotides with antibiotic drugs. Science 149, 1259–1263 (1965)
Weisblum, B.: Why centric regions of quinacrine treated mouse chromosomes show diminished fluorescence. Nature 246, 150–151 (1973)
Weisblum, B., de Haseth, P. L.: Nucleotide specificity of the quinacrine staining reaction for chromosomes. Chromosomes today 4, 35–51 (1973)
Weisblum, B., Haenssler, E.: Fluorometric properties of the bibenzimidazole derivative Hoechst 33 258, a fluorescent probe specific for AT concentration in chromosomal DNA. Chromosoma (Berl.) 46, 255–260 (1974)
Williamson, D. H., Fennell, D. J.: The use of fluorescent DNA binding agent for detecting and separating yeast mitochondrial DNA. In: Methods in cell biology, Vol. XII, Yeast cells, D. M. Prescott, ed., pp. 335–351. New York-San Francisco-London: Academic Press 1975
Yasmineh, W. G., Yunis, J. J.: Satellite DNA in calf heterochromatin. Exp. Cell Res. 64, 41–48 (1971)
Zakharov, A. F., Egolina, N. A.: Differential spiralization along mammalian mitotic chromosomes. I. BUdR-revealed differentiation in chinese hamster chromosomes. Chromosoma (Berl.) 38, 341–365 (1972)
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Schnedl, W., Breitenbach, M., Mikelsaar, A.V. et al. Mithramycin and DIPI: A pair of fluorochromes specific for GC- and AT-rich DNA respectively. Hum Genet 36, 299–305 (1977). https://doi.org/10.1007/BF00446280
Received:
Issue Date:
DOI: https://doi.org/10.1007/BF00446280