Summary
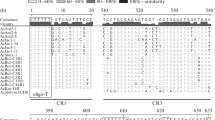
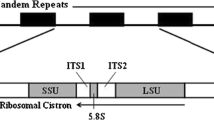
Nontranscribed spacer (NTS) regions of ribosomal (r)RNA genes are non-conserved and are shown to be useful for phylogenetic studies. 32P-labelled N. crassa NTS pCC3400 DNA, was used as a molecular probe to hybridize Southern blots of genomic DNAs obtained from Neurospora, Gelacinospora, Sordaria, bacteria, plants, and animals. Our studies conclude that: (a) the homotahllic species of Neurospora should not belong to genus Gelacinospora (a historical question) and that Neurospora homothallic species are closer to Gelacinospora than to Sordaria; and that (b) all of the filamentous fungal species tested are indeed closer to the higher plant genome than to higher primate animal genome based on shared restriction sites of 12 enzymes. Our studies also demonstrate the usefulness of nontranscribed rRNA gene probes in resolving questions regarding phylogenetic relatedness between widely separated organisms using the parsimony principle based on mutation sites from DNA restriction maps; it has not been possible to do this using DNA: DNA hybridization procedures that involved the total genome.
Similar content being viewed by others
References
Baharaeen S, Vishniac H (1984) Can J Microbiol 30:613–621
Chambers C (1985) PhD Thesis, Howard University, Washington, DC
Chambers C, Dutta SK, Crouch RJ (1986) Gene (in press)
Dutta SK, Bollon AP (1984) The Nucleus 27(1, 2):6–19.
Dutta SK, Frederick L, Austin WL (1981) J Elisha Mitchell Sci Soc 97:126–135
Dutta SK, Sheikh I, Choppala J, Aulakh GS, Nelson WH (1976) Mol Gen Genet 147:325–330
Dutta SK, Williams NP, Mukhopadhyay DK (1983) Mol Gen Genet 189:207–210
Dutta SK (1976) Mycologia LXVIII:388–401
Ferris SK, Wilson AC, Brown WM (1981) Proc Natl Acad Sci USA 78:2432–2436
Gottschalk M, Blanz PA (1984) Nucleic Acids Res 12:3951–3958
Le-John HB (1974) In: Dobzhansky T, Hecht M, Steere WC (eds) Evolutionary biology, vol 7. Plenum, New York, pp 79–125
Maniatis T, Fritsch EF, Sambrook J (1982) Molecular cloning, a laboratory manual. Cold Spring Harbor Laboratory, Cold Spring Harbor, NY
Nei M, Li W (1979) Proc Natl Acad Sci USA 76:5269–5273
Ojha M, Dutta SK (1986) In: Dutta SK (ed) DNA systematics: Plants, vol II. CRC Press, Florida, pp 45–62
Rigby PW, Dieckman M, Rhodes C, Berg P (1977) J Mol Biol 113:237–251
Russell PJ, Wagner S, Rodland KD, Feinbaum RL, Russell JP, Bret-Harte MS, Free SJ, Metzenberg RL (1984) Mol Gen Genet 196:275–282
Storck R, Alexopoulos CJ, Phaff HJ (1969) J Bacteriol 98:1069
Verma M, Ghosh B, Chambers C, Crouch RJ, Dutta SK (1985) Curr Sci 54:1092–1097
Vogel HJ (1960) Biochim Biophys Acta 41:172–173
Vogel HJ (1964) Am Nat 98:435–446
Williams MW, DeSalle R, Strobeck C (1985) Mol Biol Evol 2:338–346
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Verma, M., Dutta, S.K. Phylogenetic implication of heterogeneity of the nontranscribed spacer of rDNA repeating unit in various Neurospora and related fungal species. Curr Genet 11, 309–314 (1987). https://doi.org/10.1007/BF00355405
Received:
Issue Date:
DOI: https://doi.org/10.1007/BF00355405