Summary
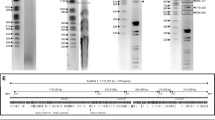
A plasmid-like DNA molecule was isolated from the fungus Ascobolus immersus in strains which exhibited genetic instability. It was found to be inherited maternally. The molecule had a density of 1.708 g/cc and a molecular weight of 4.1×106 daltons. Digestion simultaneously by two restriction enzymes demonstrated the linearity of the molecule. Electron microscopy of alkaline-denatured and self-renatured molecules revealed that the ends are inverted repeated sequences, 1200 base pairs in length.
Similar content being viewed by others
References
Aay C, Borst P (1972) The gel electrophoresis of DNA. Biochem Biophys Acta 269:192–200
Agsteribbe E, Kroon AM, Van Bruggen EFJ (1972) Circular DNA from mitochondria of Neurospora crassa. Biochem Biophys Acta 269:299–303
Buongiorno-Nardelli M, Amaldi F, Lava-Sanchez PA (1976) Electron microscope analysis of amplifying ribosomal DNA from Xenopus laevis. Exp Cell Res 98:95–103
Calos MP, Miller JH (1980) Transposable elements. Cell 20:579–595
Collins RA, Stohl LL, Cole MD, Lambowitz AM (1981) Characterization of a novel plasmid DNA found in mitochondria of Neurospora crassa. Cell 24:443–452
Cummings DJ, Belcour L, Grandchamp C (1979) Mitochondrial DNA from Podospora anserina. II. Properties of mutant DNA and multimeric circular DNA from senescent cultures. Mol Gen Genet 171:239–250
Decaris B (1981) Intragenic location of an unstable insertion element within gene b5 of the fungus Ascobolus immersus. Mol Gen Genet 184:434–439
Decaris B, Francou F, Lefort C, Rizet G (1978) Unstable ascospore color mutants of Ascobolus immersus. I. Temporal occurrence and modalities of back-mutations. Mol Gen Genet 162:69–81
Decaris B, Francou F, Kouassi A, Lefort C, Rizet G (1980) Genetic instability in Ascobolus immersus: Modalities of back-mutations, intragenic mapping of unstable sites and unstable insertion. Preliminary biochemical data. Cold Spring Harbor Symp Quant Biol 45:509–517
Decaris B, Lefort C, Francou F, Rizet G (1979) Unstable ascospore color mutants of Ascobolus immersus. II. Determinism and possible mechanisms of gene instability. Mol Gen Genet 170:191–202
Del Giudice L, Wolf K, Sassone-Corsi P, Mazza A (1979) 2 μm covalently-closed non mitochondrial circular DNA in the petite-negative yeast Schizosaccharomyces pombe. Mol Gen Genet 172:165–169
Flock JI (1977) Deletion mutants of temperate Bacillus subtilis bacteriophage ϕ105. Mol Gen Genet 155:241–247
Guerineau M (1979) Plasmid DNA in yeast. In: Lemke PA (ed) Viruses and plasmids in fungi, vol 1. Dekker M Inc., New York and Basel, p 539
Gunge N, Tamaru A, Ozawa F, Sakagucki K (1981) Isolation and characterization of linear deoxyribonucleic acid plasmids from Kluyveromyces lactis and the plasmid associated killer character. J Bacteriol 145:382–390
Inman RB, Schnös M (1970) Partial denaturation of thymine and 5-bromouracil containing DNA in alkali. J Mol Biol 49:93–98
Kück U, Stahl U, Esser K (1981) Plasmid-like DNA is part of mitochondrial DNA in Podospora anserina. Current Genet 3:151–156
Larionov VL, Grishin AV, Smirnov MN (1980) 3 μm DNA — an extrachromosomal ribosomal DNA in the yeast Saccharomyces cerevisiae. Gene 12:41–49
Le Pecoq JB, Paoletti C (1966) A new fluorometric method for RNA and DNA determination. Annal Biochem 17:100–107
Lewis LA, Decaris B (1974) The induction of apothecial formation in Ascobolus immersus by a spermatization technique. Trans Br Mycol Soc 63:197–199
Meyerink JH, Klootwijk J, Planta RJ (1979) Extrachromosomal circular ribosomal DNA in the yeast Saccharomyces carlsbergensis. Nucleic Acids Res 7:69–76
Potter S, Truett M, Phillips M, Maher A (1980) Eucaryotic transposable genetic elements with inverted terminal repeats. Cell 20:639–647
Rizet G, Engelman N, Lefort C, Lissouba P, Mousseau J (1960) Sur un ascomycète intéressant pour l'étude de certains aspects du problème de la structure du gène. CR Acad Sci (Paris) 250:2050–2052
Rizet G, Lefort C, Decaris B, Francou F, Kouassi A (1979) Unstable ascospore color mutants of Ascobolus immersus. III. Mapping, temporal occurrences and modalities of back-mutations of 34 new unstable mutants. Mol Gen Genet 175:293–303
Roeder GS, Farabaugh PJ, Chaleff DT, Fink GR (1980) The origins of gene unstability in yeast. Science 209:1375–1384
Sinclair JH, Stevens BJ, Sanghavi P, Rabinowitz M (1967) Mitochondrial-satellite and circular DNA filaments in yeast. Science 156:1234–1237
Stanfield SW, Lengyel JA (1979) Small circular DNA of Drosophila melanogaster: chromosomal homology and kinetic complexity. Proc Natl Acad Sci USA 76:6142–6146
Vierny C, Begel O, Keller AM, Raynal A, Belcour L (1980) Senescence specific DNA of Podospora anserina: its variability and its relation with mitochondrial DNA. In: Kroon AM, Saccone C (eds) The organization and expression of the mitochondrial genome. Elsevier North Holland Biomedical Press, Amsterdam
Wesolowski M, Algeri A, Fukuhara H (1981) ADN killer de la levure Kluyveromyces lactis: une version ADN du système killer à ds RNA de Saccharomyces cerevisiae. Congrès de printemps de la Société de chimie biologique. Plasmides, transposons et séquences d'insertion
Wong FY, Wildman SG (1972) Simple procedure for isolation of satellite DNAs from tobacco leaves in high yield and demonstration of minicircles. Biochem Biophys Acta 259:5–12
Zickler D (1973) La méiose et les mitoses au cours du cycle de quelques ascomycètes. Thesis. University of Paris XI, Orsay
Author information
Authors and Affiliations
Additional information
Communicated by W. Gajewski
Rights and permissions
About this article
Cite this article
Francou, F. Isolation and characterization of a linear DNA molecule in the fungus Ascobolus immersus . Mol Gen Genet 184, 440–444 (1981). https://doi.org/10.1007/BF00352519
Received:
Issue Date:
DOI: https://doi.org/10.1007/BF00352519