Summary
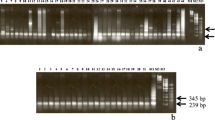
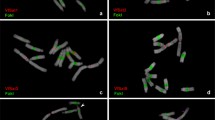
Another satellite DNA repeat (type IV) in the genome of Cucumis sativus (cucumber) was found and investigated with respect to DNA sequence, methylation, and evolution. This satellite shows a repeat length of 360 bp and a GC-content of 47%. The repeats of type IV are highly conserved among each other. Evidence for CG and CNG methylation is presented. By comparison to the previously described satellites (type I/II and type III) from cucumber, it is evident that this repeat is created by an insertion of a 180 bp DNA sequence similar to type I–III into another DNA sequence (or vice versa), and subsequent amplification forming a new satellite repeat. The different satellites of the type I/II, type III, and the 180 bp insert of type IV show a sequence homology of 60%–70%, indicating that the complex satellite DNA of cucumber is originated from a common progenitor by mutation, additional insertion, and amplification events. Copies of a sequence similar to a part of type IV are present in the genome of the related species Cucumis melo (melon).
Similar content being viewed by others
References
Arnheim N (1983) Concerted evolution of multigene families. In: Koehn R Nei M (eds) Evolution of genes and proteins. Sinauer, Sunderland, pp 38–61
Bedbrook JR, Jones J, O'Dell M, Thompson R, Flavell RB (1980) A molecular description of telomeric heterochromatin in Secale species. Cell 19:545–560
Beridze T (1986) Satellite DNA. Springer, Berlin Heidelberg
Brennicke A, Hemleben V (1983) Sequence analysis of the cloned Cucumis melo highly repetitive satellite DNA. Z Naturforsch 38c:1062–1065
Dover G (1982) Molecular drive: A cohesive mode of species evolution. Nature 299:111–117
Dover GA (1986) Molecular drive in multigene families: How biological novelties arise, spread and are assimilated. Trends Genet 2:159–165
Dover GA, Tautz D (1986) Conversion and divergence in multigene families: alternatives to selection and drift. Philos Trans R Soc London, Ser B 312:275–289
Flavell RB (1982) Sequence amplification, deletion, and rearrangement: Major sources of variation during genome evolution. In: Dover GA, Flavell RB (eds) Genome evolution. Academic Press, London, pp 301–323
Flavell R (1985) Repeated sequences and genome change. In: Hohn B, Dennis E (eds) Genetic flux in plants. Springer, Wien New York, pp 139–156
Flavell RB (1986) Repetitive DNA and chromosome evolution in plants. Philos Trans R Soc London, Ser B 312:227–242
Freeling M (1984) Plant transposable elements and insertion sequences. Annu Rev Plant Physiol 35:277–298
Ganal M, Hemleben V (1986a) Different AT-rich satellite DNAs in Cucurbita pepo and Cucurbita maxima. Theor Appl Genet 73:129–135
Ganal M, Hemleben V (1986b) Comparison of the ribosomal RNA genes in four closely related Cucurbitaceae. Plant Syst Evol 154:63–77
Ganal M, Riede I, Hemleben V (1986) Organization and sequence analysis of two related satellite DNAs in cucumber (Cucumis sativus L.). J Mol Evol 23:23–30
Gruenbaum Y, Narveh-Many T, Cedar H, Razin A (1981) Sequence specifity of methylation in higher plant DNA. Nature 292:860–862
Grunstein M, Hogness D (1975) Colony hybridization: A method for the isolation of cloned DNAs that contain a specific gene. Proc Natl Acad Sci USA 72:3961–3965
Hemleben V, Grierson D, Dertmann H (1977) The use of equilibrium centrifugation in actinomycin D-caesium chloride for the purification of ribosomal DNA. Plant Sci Lett 9:129–135
Hemleben V, Leweke B, Roth A, Stadler I (1982) Organization of highly repetitive satellite DNA of two Cucurbitaceae species (Cucumis melo and Cucumis sativus). Nucleic Acids Res 10:631–644
Ingle J, Timmis JN, Sinclair (1975) The relationship between satellite deoxyribonucleic acid, ribosomal ribonucleic acid gene redundancy, and genome size in plants. Plant Physiol 55:496–501
Kessler C, Neumaier PS, Wolf W (1985) Recognition sequences of restriction endonucleases and methylases — a review. Gene 33:1–102
Larson R, Messing J (1982) Apple II software for M13 shotgun sequencing. Nucleic Acids Res 10:39–49
Maniatis T, Fritsch EF, Sambrook J (1982) Molecular cloning. Cold Spring Harbor Laboratory
Maxam A, Gilbert W (1977) A new method for sequencing DNA. Proc Natl Acad Sci USA 71:2461–2465
Messing J, Vieira J (1982) A new pair of M13 vectors for selecting either DNA strand or double-digest restriction fragments. Gene 19:269–276
Miklos GLG (1986) Localized highly repetitive DNA sequences in vertebrate and invertebrate genomes. In: MacIntyre RJ (ed) Molecular evolutionary genetics. Plenum Press, New York London, pp 241–322
Pech M, Streeck RE, Zachau HG (1979) Patchwork structure of a bovine satellite DNA. Cell 18:883–893
Perl-Treves R, Galun E (1985) The Cucumis plastome: physical map, intrageneric variation and phylogenetic relationships. Theor Appl Genet 71:417–429
Perl-Treves R, Zamir D, Navot N, Galun E (1985) Phytogeny of Cucumis based on isozyme variability and its comparison with plastome phytogeny. Theor Appl Genet 71:430–436
Pöschl E, Streeck RE (1980) Prototype sequence of bovine 1.720 satellite DNA. J Mol Biol 143:147–153
Ramachandran C, Brandenburg WA, den Nijs APM (1985) Intraspecific variation in C-banded karyotype and chiasma frequency in Cucumis sativus (Cucurbitaceae). Plant Syst Evol 151:31–41
Roizes GP, Pages M (1982) Conserved and divergent sequences of bovine satellite DNAs. In: Dover GA, Flavell RB (eds) Genome evolution. Academic Press, London, pp 301–323
Saedler H, Nevers P (1985) Transposition in plants: a molecular model. EMBO J 4:585–590
Sanger F, Nicklen S, Coulson AR (1977) DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci USA 74:5463–5476
Shmookler-Reis R, Timmis JN, Ingle J (1981) Divergence, differential methylation, and interspersion of melon satellite DNA sequences. Biochem J 195:723–734
Singer MF (1982) Highly repeated sequences in mammalian genomes. Int Rev Cytol 76:67–112
Smith GP (1976) Evolution of repeated DNA sequences by unequal crossover. Science 191:528–535
Streeck RE (1982) A multicopy insertion sequence in the bovine genome with structural homology to the long terminal repeats of retroviruses. Nature 298:767–769
Taparowsky EJ, Gerbi SA (1982) Structure of 1.711 g/cm3 bovine satellite DNA: evolutionary relationship to satellite I. Nucleic Acids Res 10:5503–5515
Timmis JN, Ingle J (1977) Variation in satellite DNA from some higher plants. Biochem Genet 15:1159–1173
Vieira J, Messing J (1982) The pUC plasmids, an M13mp7-derived system for insertion mutagenesis and sequencing with universal primers. Gene 19:259–268
Author information
Authors and Affiliations
Additional information
Communicated by F. Mechelke
Rights and permissions
About this article
Cite this article
Ganal, M., Hemleben, V. Insertion and amplification of a DNA sequence in satellite DNA of Cucumis sativus L. (cucumber). Theoret. Appl. Genetics 75, 357–361 (1988). https://doi.org/10.1007/BF00303977
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00303977