Abstract
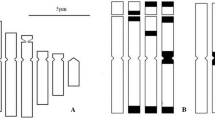
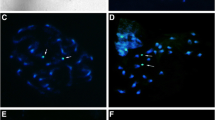
Karyotypes, DNA amounts, and meiotic behaviour were examined in population samples of two closely related species, Gibasis venustula and G. heterophylla, and their F1 hybrids. All samples were diploid (2n=12). DNA amount was similar in G. heterophylla, G. venustula ssp. robusta, and some populations of G. venustula ssp. venustula but in 13 samples of the latter, it showed as much as a 60% difference despite karyotypic uniformity. Although DNA levels were related to the altitude of the habitat, there was some variation within two morphological and ecological races of subspecies venustula. Analysis of hybrid meiosis and pollen fertility demonstrated the cytogenetical discreteness of populations, races, and subspecies. Populations and races were concluded still to be evolving following ecological isolation. The significance of quantitative and qualitative genome changes in evolution and their bearing on the classification of these species is discussed.
Similar content being viewed by others
References
Bennett MD (1972) Nuclear DNA content and minimum generation time in herbaceous plants. Proc R Soc Lond B 181:109–135
Bennett MD (1976) DNA amount, latitude and crop plant distribution. Environ Exp Bot 16:93–108
Brandham PE (1969) Chromosome behaviour in the Aloineae. I. The nature and significance of E-type bridges. Chromosoma 27:201–215
Brandham PE (1970) The consequences of meiotic crossing-over in pericentric inversions in acrocentric chromosomes. Heredity 25:125–129
Brandham PE (1977) The meiotic behaviour of inversions in the Aloineae. I. Paracentric inversions. Chromosoma 62:69–84
Brandham PE (1983) Evolution in a stable chromosome system. In: Brandham PE, Bennett MD (eds) Proc Kew Chromosome Conference II. George Allen and Unwin, London, pp 251–260
Cavalier-Smith T (1978) Nuclear volume control by nucleoskeletal DNA, selection for cell volume and cell growth rate, and the solution of the DNA C-value paradox. J Cell Sci 34:247–278
Dover GA (1977) Variation in genome organization in related species: an annotation. In: de la Chapelle A, Sorsa M (eds) Chromosomes today 6. Elsevier/North-Holland, Amsterdam, pp 105–115
Finch RA, Bennett MD (1980) Mitotic and meiotic chromosome behaviour in new hybrids of Hordeum with Triticum and Secale. Heredity 44:201–209
Gosalvez J, Lopez-Fernandez C, Sañudo A (1981) The meiotic consequences of side-arm bridges. Genetica 54:233–235
Hutchinson J, Rees H, Seal AG (1979) An assay of the activity of the supplementary DNA in Lolium. Heredity 43:411–421
Jenkins G, Rees H (1983) Synaptonemal complex formation in a Festuca hybrid. In: Brandham PE, Bennett MD (eds) Proc Kew Chromosome Conference II. George Allen and Unwin, London, pp 233–242
Jones GH (1968) Meiotic errors in rye related to chiasma formation. Mutat Res 5:385–395
Jones K, Kenton A (1984) Mechanisms of chromosome change in the evolution of the tribe Tradescantieae (Commelinaceae). In: Sharma A, Sharma AK (eds) Chromosomes in evolution of eukaryotic groups. CRC Press, Florida, pp 143–168
Jones K, Bhattarai S, Hunt DR (1981) Contributions to the cytotaxonomy of the Commelinaceae. Gibasis linearis and its allies. Bot J Linn Soc 83:141–156
Jones RN, Rees H (1968) Nuclear DNA variation in Allium. Heredity 23:591–605
Kenton A (1981) Chromosome evolution in the Gibasis linearis alliance (Commelinaceae) I. The Robertsonian differentiation of G. venustula and G. speciosa. Chromosoma 84:291–304
Kenton A (1982) Chromosome evolution in Mexican species of Gibasis (Commelinaceae). PhD Thesis, Reading University
Kenton A (1983) Qualitative and quantitative chromosome change in the evolution of Gibasis. In: Brandham PE, Bennett MD (eds) Proc Kew Chromosome Conference II. George Allen and Unwin, London, pp 273–281
Kenton A (1984) Robertsonian differentiation and preferential pairing revealed in species and F1 hybrids. Chromosome evolution in the Gibasis linearis group II. Plant Syst Evol 144:221–240
Nagl W, Ehrendorfer F (1974) DNA content, heterochromatin, mitotic index and growth in perennial and annual Anthemidae (Asteraceae). Plant Syst Evol 123:35–54
Narayan RKJ (1982) Discontinuous DNA variation in the evolution of plant species. Evolution 36:877–891
Paroda RS, Rees H (1971) Nuclear DNA variation in eu-sorghums. Chromosoma 32:353–363
Price HJ (1976) Evolution of DNA content in higher plants. Bot Rev 42:27–57
Price HJ, Chambers KL, Bachmann K (1981a) Geographic and ecological distribution of genomic DNA content variation in Microseris douglasii (Asteraceae). Bot Gaz 142:415–426
Price HJ, Chambers KL, Bachmann K (1981b) Genome size variation in diploid Microseris bigelovii (Asteraceae). Bot Gaz 142:156–159
Price HJ, Chambers KL, Bachmann K, Riggs J (1983) Inheritance of nuclear 2C DNA content variation in intraspecific and interspecific hybrids of Microseris (Asteraceae). Am J Bot 70:1133–1138
Raina SN, Rees H (1983) DNA variation between and within chromosome complements of Vicia species. Heredity 51:335–346
Seal AG (1983) The distribution and consequences of changes in nuclear DNA content. In: Brandham PE, Bennett MD (eds) Proc Kew Chromosome Conference II. George Allen and Unwin, London, pp 225–232
Seal AG, Rees H (1982) The distribution of quantitative DNA changes associated with the evolution of diploid Festuceae. Heredity 49:179–190
Sherwood SW, Patton JL (1982) Genome evolution in pocket gophers. II. Variation in cellular DNA content. Chromosoma 85:163–179
Smith JB, Gustafson JP, Bennett MD (1976) Variation in DNA amount and heterochromatin patterns in Secale. In: Jones K, Brandham PE (eds) Current chromosome research, North-Holland, Amsterdam, pp 232–233
Tobgy HA (1943) A cytological study of Crepis fuliginosa, C. neglecta and their F1 hybrid, and its bearing on the mechanism of phylogenetic reduction in chromosome number. J Genet 45:67–111
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Kenton, A. Chromosome evolution in the Gibasis linearis group (Commelinaceae). Chromosoma 90, 303–310 (1984). https://doi.org/10.1007/BF00287039
Received:
Revised:
Issue Date:
DOI: https://doi.org/10.1007/BF00287039