Abstract
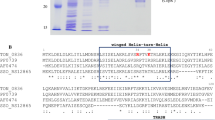
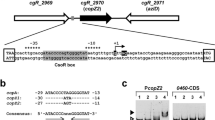
The copper resistance (cop) operon promoter (P cop ) of Pseudomonas syringae is copper-inducible, and requires the regulatory genes copRS. Sequence analysis revealed that CopR has significant homology with other known activator proteins from bacterial two-component regulatory systems. In the present study we characterized P cop and its interaction with CopR. We found that crude protein extracts from copper-resistant and-sensitive strains of P. syringae contain a P cop -specific DNA-binding protein. We hypothesized that this DNA-binding protein was the product of copR. A 27-kDa protein, which corresponded to the predicted copR product, was expressed from this gene in Escherichia coli. CopR was purified, and the first eight amino acids were sequenced to confirm its relationship to copR. Specific binding of purified CopR to the plasmid-borne P coP and the chromosomally encoded cop homolog promoter (P copH), identified in this report, was demonstrated using specific and non-specific promoter competitors in DNA mobility shift assays. DNase I footprinting identified a conserved CopR binding region (cop box) on P cop and P copH . The cop box contains an inverted repeat within a stretch of 16 bp, which shares approximately 75% identity with the PhoB binding region from several phosphate regulon gene promoters in E. coli. Primer extension analysis identified the transcriptional initiation site of P cop 59 by 5′ to the translational start site of copA, and the transcriptional initiation site of P copH 88 by 5′ to the translational start site of the chromosomal homolog of copA. The cop box was localized to between positions -54 and -35 relative to the transcriptional initiation site of P cop and P copH . Deletion analysis of P cop delimited copper-inducible activity to a 104-bp region. P cop and P copH do not share a sequence consensus with other characterized promoters from P. syringae or E. coli. The results presented delineate important regions on two copper-inducible promoters from P. syringae.
Similar content being viewed by others
References
Ausubel FM, Brent R, Kingston RE, Moore DD, Siedman JG, Smith JA, Struhl K (1987) Current protocols in molecular biology. John Wiley & Sons, New York
Bender CL, Cooksey DA (1986) Indigenous plasmids in Pseudomonas syringae pv. tomato: conjugative transfer and role in copper resistance. J Bacteriol 165:534–541
Bender CL, Cooksey DA (1987) Molecular cloning of copper resistance genes from Pseudomonas syringae pv. tomato. J Bacteriol 169:470–474
Boyer HW Roulland-Dussoix D (1969) A complementation analysis of the restriction and modification of DNA in Escherichia coli. J Mol Biol 41:459–472
Cass AEG, Hill HAO (1980) Copper proteins and copper enzymes. CIBA Found Symp 79:71–85
Cha J-S, Cooksey DA (1991) Copper resistance in Pseudomonas syringae mediated by periplasmic and outer membrane proteins. Proc Natl Acad Sci USA 88:8915–8919
Cha J-S, Cooksey DA (1993) Copper hypersensitivity and uptake in Pseudomonas syringae containing cloned components of the copper resistance operon. Appl Environ Microbiol 59:1671–1674
Charles TC, Jin S, Nester EW (1992) Two-component sensory transduction systems in phytobacteria. Annu Rev Phytopathol 30:463–484
Chou PY, Fasman GD (1978) Prediction of the secondary structure of proteins from their amino acid sequence. Adv Enzymol 47:45–147
Collado-Vides J, Magasanik B, Gralla JD (1991) Control site location and transcriptional regulation in Escherichia coli. Microbiol Rev 55:371–394
Comeau DE, Ikenaka K, Tsung K, Inouye M (1985) Primary characterization of the protein products of the Escherichia coli ompB locus: structure and regulation of synthesis of the OmpR and EnvZ proteins. J Bacteriol 164:578–584
Cooksey DA (1987) Characterization of a copper resistance plasmid conserved in copper-resistant strains of Pseudomonas syringae pv. tomato. Appl Environ Microbiol 53:454–456
Cooksey DA (1990) Genetics of bactericide resistance in plant pathogenic bacteria. Annu Rev Phytopathol 28:201–219
Cooksey DA (1993) Copper uptake and resistance in bacteria. Mol Microbiol 7:1–5
Cooksey DA, Azad HR, Cha J-S, Lim C-K (1990) Copper resistance gene homologs in pathogenic and saprophytic bacterial species from tomato. Appl Environ Microbiol 56:431–435
Devereux J, Haeberli P, Smithies O (1984) A comprehensive set of sequence analysis programs for the VAX. Nucleic Acids Res 12:387–395
Garnier J, Osguthorpe DJ, Robson B (1978) Analysis of the accuracy and implications of simple methods for predicting secondary structure of globular proteins. J Mol Biol 120:97–120
Keane PJ, Kerr A, New PB (1970) Crown gall of stone fruit. II. Identification and nomenclature of Agrobacterium isolates. Aust J Biol Sci 23:585–595
Keen NT, Tamaki S, Kobayashi D, Trollinger D (1988) Improved broad-host-range plasmids for DNA cloning in gram-negative bacteria. Gene 70:191–197
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Lim C-K, Cooksey DA (1993) Characterization of chromosomal homologs of the plasmid-borne copper resistance operon of Pseudomonas syringae. J Bacteriol 175:4492–4498
Makino K, Shinagawa H, Ammemura M, Nakata A (1986a). Nucleotide sequence of the phoB gene, the positive regulatory gene for the phosphate regulon of Escherichia coli K-12. J Mol Biol 190:37–44
Makino K, Shinagawa H, Ammemura M, Nakata A (1986b) Nucleotide sequence of the phoR gene, a regulatory gene for the phosphate regulon of Escherichia coli. J Mol Biol 192:549–556
Makino K, Shinagawa H, Amemura M, Kimura S, Nakata A, Ishihama A (1988) Regulation of the phosphate regulon of Escherichia coli: activation of pstS transcription by PhoB in vitro. J Mol Biol 203:85–95
Mellano MA (1988) PhD thesis. University of California, Riverside
Mellano MA, Cooksey DA (1988a) Nucleotide sequence and organization of copper resistance genes from Pseudomonas syringae pv. tomato. J Bacteriol 170:2879–2883
Mellano MA, Cooksey DA (1988b) Induction of the copper resistance operon from Pseudomonas syringae. J Bacteriol 170:4399–4401
Miller JH (1972) Experiments in molecular genetics. Cold Spring Harbor Laboratory, Cold Spring Harbor, New York
Mills SD, Jasalavich CA, Cooksey DA (1993) A two-component regulatory system required for copper-Inducible expression of the copper resistance operon of Pseudomonas syringae. J Bacteriol 175:1656–1664
Mohr CD, Leveau JHJ, Krieg DP, Hibler NS, Deretic V (1992) A1gR-binding sites within the algD promoter make up a set of inverted repeats separated by a large intervening segment of DNA. J Bacteriol 174:6624–6633
O'Halloran TV, Walsh C (1987) Metalloregulatory DNA binding protein encoded by the merR gene: isolation and characterization. Science 235:211–214
O'Halloran TV, Frantz B, Shin MK, Ralston DM, Wright JG (1989) The MerR heavy metal receptor mediates positive activation in a topologically novel transcription complex. Cell 56:119–129
Ouzounis C, Sander C (1991) A structure-derived sequence pattern for the detection of type I-copper binding domains in distantly related proteins. FEBS Lett 279:73–78
Parkinson JS, Kofoid EC (1992) Communication modules in bacterial signaling proteins. Annu Rev Genet 26:71–112
Rouch DJ, Camakaris J, Lee BTO, Luke RK (1985) Inducible plasmid mediated copper resistance in Escherichia coli. J Gen Microbiol 131:939–943
Rouch DJ, Camakaris J, Lee BTO (1989a) Copper transport in Escherichia coli. In: Hamer DH, Winge DR (eds) Metal ion homeostasis: molecular biology and chemistry. Liss, New York, pp 469–477
Rouch DJ, Lee BTO, Camakaris J (1989b) Genetic and molecular basis of copper resistance in Escherichia coli. In: Hamer DH, Winge DR (eds) Metal ion homeostasis: molecular biology and chemistry. Liss, New York, pp 439–446
Silver S, Lee BTO, Brown NL, Cooksey DA (1993) Bacterial plasmid resistances to copper, cadmium, and zinc. In: Welch AJ, Chapman SU (eds) The chemistry of the copper and zinc triads. Royal Society of Chemistry, London, pp 38–53
Simon R, Priefer U, Pühler A (1983) A broad host range mobilization system for in vivo genetic engineering: transposon mutagenesis in gram negative bacteria. Bio/Technology 1:784–792
Spaink HP, Okker RJH, Wijffelman CA, Pees E, Lugtenberg B (1986) Promoters and operon structure of the nodulation region of the Rhizobium leguminosarum symbiosis plasmid pRL1JI. In: Lugtenberg B (ed) Recognition in microbe-plant symbiotic and pathogenic interactions (NATO ASI series, vol H4). Springer, Berlin, pp 55–68
Stock JB, Ninfa AJ, Stock AM (1989) Protein phosphorylation and regulation of adaptative responses in bacteria. Microbiol Rev 53:450–490
Wackett LP, Orme-Johnson WH, Walsh CT (1989) Transition metal enzymes in bacterial metabolism. In: Beveridge TJ, Doyle RJ (eds) Metal ions and bacteria. Wiley & Sons. New York, pp 165–206
Yanisch-Perron C, Vieira J, Messing J (1985) Improved M13 phage cloning vectors and host strains: nucleotide sequences of the M13mp18 and pUC19 vectors. Gene 33:103–119
Author information
Authors and Affiliations
Additional information
Communicated by A. Kondorosi
Rights and permissions
About this article
Cite this article
Mills, S.D., Lim, CK. & Cooksey, D.A. Purification and characterization of CopR, a transcriptional activator protein that binds to a conserved domain (cop box) in copper- inducible promoters of Pseudomonas syringae . Molec. Gen. Genet. 244, 341–351 (1994). https://doi.org/10.1007/BF00286685
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00286685