Abstract
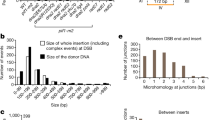
In order to elucidate the mechanisms of illegitimate recombination in eukaryotes, we have studied the structure of DNA fragments integrated by illegitimate recombination into the genome of fission yeast. Nonhomologous recombination was rarely identified when a long region of homology with the chromosomal leu1 + gene was present in the introduced leu1::ura4 + DNA fragment; but a decrease in length of homology leads to an increase in the ratio of nonhomologous to homologous recombination events. The introduced DNA fragments were integrated into different sites in the chromosomes by nonhomologous recombination. The results suggested that there are multiple modes of integration; most events simply involve both ends of the fragments, while in other cases, fragments were integrated in a more complicated manner, probably via circularization or multimerization. To analyze the mechanism of the major type of integration, DNA fragments containing the recombination junctions of three recombinants were amplified by inverted polymerase chain reaction (IPCR) and their nucleotide sequences were determined. There was no obvious homology between introduced DNA and chromosomal DNA at these recombination sites. Furthermore it was found that each terminal region of the introduced DNA was deleted, but that there were no or very small deletions in the target sites of chromosomal DNA. Two models are proposed to explain the mechanism of nonhomologous integration.
Similar content being viewed by others
References
Asch DK, Frederick G, Kinsey JA, Perkins DD (1992) Analysis of junction sequences resulting from integration at nonhomologous loci in Neurospora crassa. Genetics 130:737–748
Bae TS, Kawasaki I, Ikeda H, Liu LF (1988) Illegitimate recombination mediated by calf thymus DNA topoisomerase II in vitro. Proc Natl Acad Sci USA 85:2076–2080
Boeke JD, LaCroute F, Fink GR (1984) A positive selection for mutants lacking orotidine-5′-phosphate decarboxylase activity in yeast: 5-fluoro-orotic acid resistance. Mol Gen Genet 197:345–346
Faugeron G, Goyon C, Gregoire A (1989) Stable allele replacement and unstable non-homologous integration events during transformation of Ascobolus immersus. Gene 76:109–119
Grimm C, Kohli J. (1988) Observation on integrative transformation in Schizosaccharomyces pombe: Mol Gen Genet 215:87–93
Grimm C, Kohli J, Murray J, Maundrell K. (1988) Genetic engineering of Schizosaccharomyces pombe: a system for gene disruption and replacement using the ura4 gene as a selectable marker. Mol Gen Genet 215:81–86
Gutz H, Heslot H, Leupold U, Loprieno N (1974) Schizosaccharomyces pombe. In: King RC (ed) Handbook of genetics. Plenum, New York, pp 395–446
Hasty P, Rivera-Perez J, Bradley A (1991a) The length of homology required for gene targeting in embryonic stem cells. Mol Cell Biol 11:5586–5591
Hasty P, Rivera-Perez J, Chang C, Bradley A (1991b) Target frequency and integration pattern for insertion and replacement vectors in embryonic stem cells. Mol Cell Biol 11:4509–4517
Hattori M, Yoshioka K, Sakaki Y (1992) High-sensitive fluorescent DNA sequencing and its application for detection and massscreening of point mutations. Electrophoresis 13:560–565
Hinnen A, Hicks JB, Fink GR (1978) Transformation of yeast. Proc Natl Acad Sci USA 75:1929–1933
Ikeda H (1986) Bacteriophage T4 DNA topoisomerase mediates illegitimate recombination in vitro. Proc Natl Acad Sci USA 83:922–926
Ikeda H, Shiozaki M (1984) Nonhomologous recombination mediated by Escherichia coli DNA gyrase: possible involvement of DNA replication. Cold Spring Harbor Symp Quant Biol 49:401–409
Ikeda H, Moriya K, Matsumoto T (1981) In vitro study of illegitimate recombination: involvement of DNA gyrase. Cold Spring Harbor Symp Quant Biol 45:399–408
Ikeda H, Aoki K, Naito A (1982) Illegitimate recombination mediated in vitro by DNA gyrase of Escherichia coli: structure of recombinant DNA molecules. Proc Natl Acad Sci USA 79:3724–3728
Katinka MD, Bourgain FM (1992) Interstitial telomeres are hotspots for illegitimate recombination with DNA molecules injected into the macronucleus of Paramecium primaurelia. EMBO J 11:725–732
Kikuchi Y, Kitazawa Y, Shimatake H, and Yamamoto M (1988) The primary structure of the leu1 + gene of Schizosaccharomyces pombe. Curr Genet 14:375–379
Lin FL, Sperle K, Sternberg N (1985) Recombination in mouse L cells between DNA introduced into cells and homologous chromosomal sequences. Proc Natl Acad Sci USA 82:1391–1395
Meuth M (1989) Illegitimate recombination in mammalian cells. In: Berg DE, Howe MM (eds) Mobile DNA. American Society for Microbiology, Washington, D.C., pp 833–860
Miura-Masuda A, Ikeda H (1990) The DNA gyrase of Escherichia coli participates in the formation of a spontaneous deletion by recA − independent recombination in vivo. Mol Gen Genet 220:345–352
Moreno S, Klar A, Nurse P (1991) Molecular genetic analysis of fission yeast Schizosaccharomyces pombe. Methods Enzymol 194:794–823
Okazaki K, Okazaki N, Kume K, Jinno S, Tanaka K, Okayama H (1990) High-frequency transformation method and library transducing vectors for cloning mammalian cDNAs by transcomplementation of Schizosaccharomyces pombe. Nucleic Acids Res 18:6485–6489
Pfeiffer P, Vielmetter W (1988) Joining of nonhomologous DNA double strand breaks in vitro. Nucleic Acids Res 16:907–924
Rose MD, Winston F, Hieter P (1990) Methods in yeast genetics: a laboratory course manual. Cold Spring Harbor Laboratory, Cold Spring Harbor, New York, pp 128–129
Roth DB, Wilson JH (1986) Nonhomologous recombination in mammalian cells: role for short sequence homologies in the joining reaction. Mol Cell Biol 6:4295–4304
Roth DB, Wilson JH (1988) Illegitimate recombination in mammalian cells. In: Kucherlapati R, Smith GR (eds) Genetic recombination. American Society for Microbiology, Washington, D.C., pp 621–653
Roth DB, Porter TN, Wilson JH (1985) Mechanisms of nonhomologous recombination in mammalian cells. Mol Cell Biol 5:2599–2607
Rothstein RJ (1983) One-step gene disruption in yeast. Methods Enzymol 101:202–211
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory, Cold Spring Harbor, New York
Schiestl RH, Petes TD (1991) Integration of DNA fragments by illegitimate recombination in Saccharomyces cerevisiae. Proc Natl Acad Sci USA 88:7585–7589
Schiestl RH, Dominska M, Petes TD (1993) Transformation of Saccharomyces cerevisiae with nonhomologous DNA: illegitimate integration of transforming DNA into yeast chromosomes and in vivo ligation of transforming DNA to mitochondrial DNA sequences. Mol Cell Biol 13:2697–2705
Thode S, Schafer A, Pfeiffer P, Vielmetter W (1990) A novel pathway of DNA end-to-end joining. Cell 60:921–928
Thomas KR, Capecchi MR (1986) Introduction of homologous DNA sequences into mammalian cells induces mutations in the cognate gene. Nature 324:34–38
Thomas KR, Folger KR, Capecchi MR (1986) High frequency targeting of genes to specific sites in the mammalian genome. Cell 44:419–428
Wright APH, Maundrell K, Shall S (1986) Transformation of Schizosaccharomyces pombe by non-homologous, unstable integration of plasmids in the genome. Curr Genet 10:503–508
Author information
Authors and Affiliations
Additional information
Communicated by M. Sekiguchi
Rights and permissions
About this article
Cite this article
Tatebayashi, K., Kato, Ji. & Ikeda, H. Structural analyses of DNA fragments integrated by illegitimate recombination in Schizosaccharomyces pombe . Molec. Gen. Genet. 244, 111–119 (1994). https://doi.org/10.1007/BF00283511
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00283511