Abstract
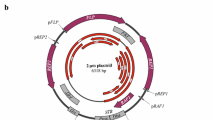
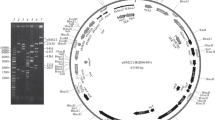
The low-copy-number and broad-host-range pSM19035-derived plasmid pBT233 is stably inherited in Bacillus subtilis cells. Two distinct regions, segA and segB, enhance the segregational stability of the plasmid. Both regions function in a replicon-independent manner. The maximization of random plasmid segregation is accomplished by the recombination proficiency of the host or the presence of the pBT233 segA region. The segA region contains two open reading frames (orϕ) [α and β]. Inactivation or deletion of orϕβ results in SegA− plasmids. Better than random segregation requires an active segB region. The segB region contains two orϕs (orϕɛ and orϕζ). Inactivation of either of the orfs does not lead to an increase in cell death, but orϕζ− plasmids are randomly segregated. These results suggest that pBT233 stabilization relies on a complex system involving resolution of plasmid oligomers (segA) and on the function(s) encoded by the segB region.
Similar content being viewed by others
References
Alonso JC, Lüder G, Tailor RH (1991) Characterization of Bacillus subtilis recombinational pathways. J Bacteriol 173:3977–3980
Austin A, Ziese M, Sternberg N (1981) A novel role for a site-specific recombination in maintenance of bacterial replicons. Cell 25:729–736
Austin S, Nordström K (1990) Partition-mediated incompatibility of bacterial plasmids. Cell 60:351–354
Birnboim HC, Doly J (1979) A rapid alkaline extraction procedure for screening recombinant plasmid DNA. Nucleic Acids Res 7:1513–1523
Biswal N, Kleinschmidt AK, Spatz HC, Trautner TA (1967) Physical properties of the DNA of bacteriophage SP50. Mol Gen Genet 100:39–55
Behnke D, Tomich PK, Clewell DB (1980) Electron microscopic mapping of deletions on a streptococcal plasmid carrying extraordinarily long inverted repeats. Plasmid 4:139–147
Boitsov AS, Golubkov VI (1980) Transformation of Bacillus subtilis by streptococcal plasmids. Dok Acad Nauk SSSR 254:490–492
Boitsov AS, Golubkov VI, Iontova IM, Zaitsev EN, Malke H, Totolian AA (1980) Inverted repeats on plasmids determining resistance to MLS antibiotics in Group A streptococci. FEMS Microbiol Lett 6:11–14
Brantl S, Behnke D (1992) Copy number control of the streptococcal plasmid pIP501 occurs at three levels. Nucleic Acids Res 20:359–400
Brand S, Nowak A, Behnke D, Alonso JC (1989) Revision of the nucleotide sequence of the Streptococcus pyogenes plasmid pSM19035 reps gene. Nucleic Acids Res 17:10110
Brantl S, Behnke D, Alonso JC (1990) Molecular analysis of the replication region of the conjugative Streptococcus agalactiae plasmid pIP501 in Bacillus subtilis. Comparison with plasmids pAMβ1 and pSM19035. Nucleic Acids Res 18:4783–4789
Bron S (1990) Plasmids. In: Harwood CR, Cutting SM (eds) Molecular biological methods for Bacillus, John Wiley and Sons, Sussex, p 75
Bron S, Meijer W, Holsappel S, Haima P (1991) Plasmid instability and molecular cloning in Bacillus subtilis. Res Microbiol 142:875–883
Bruand C, Ehrlich SD, Janniere L (1991) Unidirectional theta replication of the structurally stable Enterococcus faecalis plasmid pAMβ1. EMBO J 10:2171–2177
Canosi U, Morelli G, Trautner TA (1978) The relationship between molecular structure and transformation efficiency of some S. aureus plasmids isolated from B. subtilis. Mol Gen Genet 166:259–267
Ceglowski P, Lurz R, Alonso JC (1993a) Functional analysis of pSM19035-derived replicons in Bacillus subtilis. FEMS Microbiol Lett 109:145–150
Ceglowski P, Boitsov A, Chai S, Alonso JC (1993b) Analysis of the stabilization system of the pSM19035-derived plasmid pBT233 in Bacillus subtilis. GENE in press
Clewell DB (1981) Plasmids, drug resistance, and gene transfer in the genus Streptococcus. Microbiol Rev 45:409–436
Davis MA, Martin KA, Austin SJ (1992) Biochemical activities of the ParA partition protein of the P1 plasmid. Mol Microbiol 6:1141–1147
Gruss A, Ehrlich SD (1989) The family of highly interrelated singlestranded deoxyribonucleic acid plasmids. Microbiol Rev 53:231–241
Haima P, Bron S, Venema G (1987) The effect of restriciton on shotgun cloning and plasmid stability in Bacillus subtilis Marburg. Mol Gen Genet 209:335–342
Jannière L, Bruand C, Ehrlich SD Structurally stable Bacillus subtilis cloning vectors. Gene 87:53–59
Kolodner R (1980) Genetic recombination of bacterial plasmid DNA: electron microscopic analysis of in vitro intramolecular recombination. Proc Natl Acad Sci USA 77:4847–4851
Laban A, Cohen A (1981) Interplasmidic and intraplasmidic recombination in Escherichia coli K-12. Mol Gen Genet 184:200–207
Maciag IE, Viret J-F, Alonso JC (1988) Replication and incompatibility properties of plasmid pUB110 in Bacillus subtilis. Mol Gen Genet 212:232–240
Motallebi-Veshareh M, Rouch DA, Thomas CM (1990) A family of ATPases involved in active partitioning of diverse bacterial plasmids. Mol Microbiol 4:1455–1463
Nordström K, Austin SJ (1989) Mechanisms that contribute to the stable segregation of plasmids. Annu Rev Genet 23:37–69
Novick RP, Edelman I, Lofdahl S (1986) Small Staphylococcus aureus plasmids are transduced as linear multimers that are formed and resolved by replicative processes. J Mol Biol 192:209–220
Rottländer E, Trautner TA (1970) Genetic and transfection studies with B. subtilis phage SP50. I. Phage mutants with restricted growth on B. subtilis strains. Mol Gen Genet 108:47–60
Sambrook J, Maniatis T, Fritsch EF (1989) Molecular cloning: A laboratory manual, second edition. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York
Swinfield T-J, Oultram JD, Thompson DE, Brehn JK, Minton NP (1990) Physical characterization of the Streptococcus faecalis plasmid pAMßI. Gene 87:79–90
Swinfield T-J, Jannière L, Ehrlich SD, Minton NP (1991) Characterization of a region of the Enterococcus faecalis plasmid pAMß1 which enhances the segregational stability of pAM β1-derived cloning vectors in Bacillus subtilis. Plasmid 26:209–221
Walker JE, Saraste M, Runswich MJ, Gay NJ (1982) Distantly related sequences in the α- and β-subunits of ATP synthase, myosin, kinase and other ATP-requiring enzymes and a common nucleotide binding fold. EMBO J 1:945–951
Way JC, Davis MA, Morisato D, Roberts DE, Kleckner N (1984) New Tn10 derivatives for transposon mutagenesis and for construction of lacZ operon fusions by transposition. Gene 32:369–379
Willians DR, Thomas CM (1992) Active partitioning of bacterial plasmids. J Gen Microbiol 138:1–16
Yanisch-Perron C, Vieira J, Messing J (1985) Improved M13 phage cloning vectors and host strains: nucleotide sequences of the M13mp18 and pUC19 vectors. Gene 33:103–119
Author information
Authors and Affiliations
Additional information
Communicated by H. Böhme
Rights and permissions
About this article
Cite this article
Ceglowski, P., Boitsov, A., Karamyan, N. et al. Characterization of the effectors required for stable inheritance of Streptococcus pyogenes pSM19035-derived plasmids in Bacillus subtilis . Molec. Gen. Genet. 241, 579–585 (1993). https://doi.org/10.1007/BF00279900
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00279900