Summary
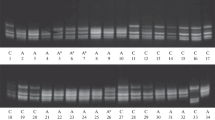
α- and β-amylase isozyme diversity was studied electrophoretically by thin-layer polyacrylamide gel isoelectrofocusing in the tetraploid wild emmer wheat, Triticum dicoccoides, the progenitor of all cultivated wheats. We analyzed 225 plants from 23 populations encompassing the ecological spectrum of T. dicoccoides in Israel. The results were as follows: (a) Band and multilocus genotype polymorphisms abound and vary within and between the four amylase components: malt, green (α-amylases), and dry and germinating seeds (β-amylases). (b) The number of bands of malt, green, and dry and germinating seeds were 20, 6, 11 and 13, respectively, generating 40, 6, 51, and 51 patterns or multilocus genotypes (MGP), respectively. The MGPs vary drastically within and between populations, from monomorphic in some populations with a single pattern to highly polymorphic ones, (c) Mean H e values for malt, green, and germinating and dry seeds are 0.053, 0.055, 0.088, and 0.077, respectively; mean number of bands per individual was 11.8, 4.4, 7.6, and 4.0, respectively, (d) The percentages of 50 bands and 148 multilocus genotype patterns (MGP) (in parenthesis) were classified into widespread, sporadic, and localized: 84.4 (10.8), 8.9 (12.2), 6.7 (77.0), respectively. Notably, 89.2% of the patterns were not widespread, but sporadic and localized, (e) The mean value of genetic distances among populations (Nei's D) for the four amylase groups is D = 0.136, 0.175, 0.288 and 0.307, respectively, not displaying geographical correlates. (f) Most of the α- and β-amylase diversity is between populations (G st = 68–75%). (g) Significant environmental correlates occur between either bands or patterns and climatic diversity (water and primarily temperature factors). (h) Significant associations of multilocus amylase bands occur across Israel. Like-wise, significant gametic phase disequilibria, D, occur within populations and are positively correlated with climatic variables, primarily that of temperature, (i) Discriminant analyses correctly classified (95–100%) the 23 wild emmer populations into their ecogeographical region and soil type. (j) Autocorrelation analysis showed that there is no correlation between bands and geographic distance and excluded migration as a major factor of amylase differentiation.
These results suggest that diversifying climatic and edaphic natural selection rather than stochastisity or migration is the major evolutionary force driving amylase differentiation at both the single and multilocus levels. Furthermore, wild emmer harbors high levels of α- and β-amylase diversity both as single bands and as multilocus adaptive genetic patterns. These are exploitable both as genetic markers for quantitative loci (QTLs) and as adaptive genetic resources to improve wheat germination and growth through classical breeding and/or biotechnology.
Similar content being viewed by others
References
Ainsworth CC, Gale MD, Baird S (1983) The genetics of β-amylase isozymes in wheat. I. Allelic variation among hexaploid varieties and intra-chromosomal gene locations. Theor Appl Genet 66:39–49
Ainsworth CC, Doherty P, Edwards KGK, Martienssen RA, Gale MD (1985) Allelic variation at α-amylase loci in hexaploid wheat. Theor Appl Genet 70:400–406
Ainsworth CC, Miller TE, Gale MD (1987) α-Amylase and β-amylase homoeoloci in species related to wheat. Genet Res 49:93–103
Akazawa T, Hara-Nishimura I (1985) Topographic aspects of biosynthesis, extracellular secretion and intracellular storage of proteins in plant cells. Annu Rev Plant Physiol 36:441–472
Baulcombe DC, Huttly AK, Martienssen RA, Barker RF, Jarvis MG (1987) A novel wheat α-amylase gene (α-Amy-3). Mol Gen Genet 209:33–40
Carver BF, Nevo E (1990) Genetic diversity of photosynthetic characters in native populations of Triticum dicoccoides. Photosynth Res 25:119–128
Dabrowska T (1983) Studies on chromosomal location of genes involved in synthesis of beta-amylase isoenzymes in wheat kernels (Triticum aestivum L). Genet Pol 24:9–19
Feldman M (1976) Wheats. In: Simmonds NW (ed) Evolution of crop plants. jLongman, London, pp 120–128
Gale MD, Ainsworth CC (1984) The relationship between α-amylase species found in developing and germinating wheat grain. Biochem Genet 22:1031–1036
Gale MD, Law CN, Chojecki AJ, Kempton RA (1983) Genetic control of α-amylase production in wheat. Theor Appl Genet 64:309–361
Hart GE (1983) Hexaploid wheat (Triticum aestivum L. em Thell) In: Tanksley SD, Orton TJ (eds) Isozymes in plant genetics and breeding. Eisevier Sci Publ, Amsterdam, pp 35–56
Hart GE, Gale MD (1989) Biochemical/molecular loci of hexaploid wheat (Triticum aestivum, 2n = 42, Genomes AABBDD). In: O'Brien JS (ed) Genetic maps. Locus maps of complex genomes, 5th edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y., pp 28–38
Huang N, Sutliff TD, Litts JC, Rodriguez RL (1990) Classification and characterization of the rice α-amylase multigene family. Plant Mol Biol 14:655–668
Huttly AK, Martienssen RA, Baulcombe DC (1988) Sequence heterogeneity and differential expression of the α-Amy-2 gene family in wheat. Mol Gen Genet 214:232–240
Jourdrier MP (1980) Controle genetique de la β-amylase du grain de blé tendre. C R Acad Sci 291:477–480
Jourdrier MP, Cauderon T (1976) Localization chromosomique de genes controlant la synthese de certains constituants β-amylase du grain de blé tendre. C R Acad Sci 282:115–118
Khursheed B, Rogers JC (1988) Barley α-amylase genes. J Biol Chem 263:18953–18960
Kimber G, Feldman M (1987) Wild wheats. An introduction. Special Report 353. College of Agriculture, University of Missouri, Columbia, Mo., pp 1–142
Knox CAP, Sonthayanon B, Chandra GR, Muthukrishnan S (1987) Structure and organization of two divergent α-amylase genes from barley. Plant Mol Biol 9:3–17
Lazarus CM, Baulcombe DC, Martienssen RA (1985) α-Amylase genes of wheat are two multigene families which are differentially expressed. Plant Mol Biol 5:13–24
Marshall DR, Brown AHD (1975) Optimum sampling strategies in genetic conservation. In: Frankel OH, Hawles JW (eds) Crop genetic resources for today and tomorrow. Cambridge University Press, Cambridge, pp 53–70
Milne DL, McIntosh RA (1989) Triticum aestivum (common wheat). In: O'Brien JS (ed) Genetic maps. Locus maps of complex genomes, 5 edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y., pp 16–27
Nei M (1972) Genetic distance between populations. Am Nat 106:283–292
Nei M (1973) Analysis of gene diversity in subdivided populations. Proc Natl Acad Sci USA 70:3321–3323
Nevo E (1983) Genetic resources of wild emmer wheat: structure, evolution and application in breeding. In: Sakamoto S (ed) Proc 6th Int Wheat Symp. Kyoto, Japan, pp 421–431
Nevo E (1986) Genetic resources of wild cereals and crop improvement: Israel, a natural laboratory. Isr J Bot 35:255–278
Nevo E (1987) Plant genetic resources: prediction by isozyme markers and ecology. In: Rattazzi MC, Scandalios JG, Whitt GS (eds) Isozymes: current topics in biological and medical research vol 16: agriculture, physiology, and medicine. Alan R. Liss, New York, pp 247–267
Nevo E (1988) Genetic resources of wild emmer wheat revisited: genetic evolution, conservation, and utilization. In: Miller TE, Koebner RMD (eds) Proc 7th Int Wheat Genet Symp. Cambridge, pp 121–126
Nevo E, Beiles A (1989) Genetic diversity of wild emmer wheat in Israel and Turkey: structure, evolution, and application in breeding. Theor Appl Genet 77:421–455
Nevo E, Carver BF, Beiles A (1991) Photosynthetic performance in wild emmer wheat Triticum dicoccoides: ecological and genetic predictability. Theor Appl Genet 81:445–460
Nevo E, Snape JW, Lavie B, Beiles A (1992) Herbicide response polymorphisms in wild emmer wheat: Ecological and isozyme correlations. Theor Appl Genet 84:209–216
Nishikawa K (1991) Chromosome mapping by use of aneuploids in wheat. Wheat Inform Serv 72:60–63
Nishikawa K, Nobuhara M (1971) Genetic studies of α-amylase isozymes in wheat. I. Location of genes and variation in tetra- and hexaploid wheat. Jpn J Genet 5:345–353
Nishikawa K, Furuta Y, Goshima H (1975) Genetic studies of α-amylase isozymes in wheat. II. Reconstituted AABB tetraploid, Aegilops squarrosa and their synthetic AABBDD hexaploid. Jpn J Genet 5:409–416
Nishikawa K, Furuta Y, Kudo S, Ujihara K (1979) Differentiation of tetraploid wheat in relation to DNA content of nucleus and alpha-amylase isozymes. Rep Plant Germ plasm Inst Kyoto Univ 4:30–38
Nishikawa K, Furuta, Y, Wada T (1980) Genetic studies on α-amylase isozymes in wheat. III. Intra-specific variation in Aegilops squarrosa and birthplace of hexaploid wheat. Jpn J Genet 55:325–336
Nishikawa K, Furuta Y, Hiraku T, Takayanagi M (1988a) Variation and chromosome arm location of the genes for Gibberelic acid induced isozymes. Genetic studies on α-amylase isozymes in wheat. V. Jpn J Breed 38:187–197
Nishikawa K, Furuta Y, Kudo S (1988b) Genetic studies of α-amylase isozymes in Wheat. VI. Variation and differentiation in tetrapoloid wheat. Jpn J Genet 63:425–434
Schell JST (1987) Transgenic plants as tools to study the molecular organization of plant genes. Science 237:1176–1183
Sharp PJ, Desai S, Gale MD (1988) Isozyme variation and RFLPs at the β-amylase loci in wheat. Theor Appl Genet 76:691–699
Snape JW, Nevo E, Parker BB, Leckie D, Morgunov A (1991) Herbicide responses in wild emmer wheat. Heredity 66:251–257
Sokal RR, Oden NL (1987a) Spatial autocorrelation in biology. I. Methodology. Biol J Linn Soc 10:199–228
Sokal RR, Oden NL (1987b) Spatial autocorrelation in biology. 2. Some biological implications and four applications of evolutionary and ecological interest. Biol J Linn Soc 10:229–249
SPSS-x (1986) User's guide, 2nd edn. McGraw-Hill, New York
Zohary D (1970) Centers of diversity and centers of origin. In: Frankel OH, Bennett E (eds) Genetic resources in plants — their exploration and conservation. Blackwell, Oxford, pp 33–42
Author information
Authors and Affiliations
Additional information
Communicated by H. F. Linskens
Rights and permissions
About this article
Cite this article
Nevo, E., Nishikawa, K., Furuta, Y. et al. Genetic polymorphisms of α- and β-amylase isozymes in wild emmer wheat, Triticum dicoccoides, in Israel. Theoret. Appl. Genetics 85, 1029–1042 (1993). https://doi.org/10.1007/BF00215044
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00215044