Abstract
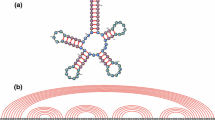
We present and study the behavior of a simple kinetic model for the melting of RNA secondary structures, given that those structures are known. The model is then used as a map that. assigns structure dependent overall rate constants of melting (or refolding) to a sequence. This induces a “landscape” of reaction rates, or activation energies, over the space of sequences with fixed length. We study the distribution and the correlation structure of these activation energies.
Similar content being viewed by others
References
Beaudry AA, Joyce GF (1992) Directed evolution of an RNA enzyme. Science 257:635–641
Bonhoeffer S, McCaskill JS, Stadler PF, Schuster P (1993) Temperature dependent RNA landscapes. A study based on partition functions. Eur Biophys J 22:13–24
Eigen M, McCaskill JS, Schuster P (1989) The molecular quasispecies. Adv Chem Phys 75:149–263
Ellington AD, Szostak JW (1990) In vitro selection of RNA molecules that bind specific ligands. Nature 346:818–822
Feinberg M (1977) Mathematical aspects of mass action kinetics. In: Lapidus L, Amundson NR (eds) Chemical reactor theory. Prentice Hall, Englewood Cliffs, NJ, pp 1–78
Fernández A, Shakhnovich EI (1990) Activation-energy landscape for metastable RNA folding. Phys Rev A 42:3657–3659
Fontana W, Griesmacher T, Schnabl W, Stadler PF, Schuster P (1991) Statistics of landscapes based on free energies replication and degradation rate constants of RNA secondary structures. Mh Chem 122:795–819
Fontana W, Konings DAM, Stadler PF, Schuster P (1993a) Statistics of RNA secondary structures. SFI preprint 92-02-007. Biopolymers (submitted for publication)
Fontana W Stadler PF, Bornberg-Bauer EG, Griesmacher T, Hofacker IL, Tacker M, Tarazona P, Weinberger ED, Schuster P (1993b) Phys Rev E47:2083–2099
Freier SM, Kierzek R, Jaeger JA, Sugimoto N, Caruthers MH, Neilson T, Turner DH (1986) Improved free-energy parameters for predictions of RNA duplex stability. Biochemistry 83:9373–9377
Horowitz MSZ, Dube DK, Loeb LA (1989) Selection of new biological activities from random nucleotide sequences. Genome 31:112–117
Jaeger JA, Turner DH, Zuker M (1989) Improved predictions of secondary structures for RNA. Biochemistry 86:7706–7710
Joyce GF (1989) Amplification, mutation and selection of catalytic RNA. Gene 82:83–87
Kauffman SA, Levin S (1987) Towards a general theory of adaptive walks on rugged landscapes. J. Theor Biol 128:11–45
Kauffman SA, Weinberger ED, Perelson AS (1988) Maturation of the immune response via adaptive walks on affinity landscapes. Theoretical immunology, Part I Santa Fe Institute Studies in the Sciences of Complexity. Perelson AS (ed),Addison-Wesley, Reading, Mass
Macken CA, Perelson AS (1989) Protein evolution on rugged landscapes. Proc Nat Acad Sci 86:6191–6195
McCaskill JS (1990) The equilibrium partition function and base pair binding probabilities for RNA secondary structures. Biopolymers 29:1105–1119
Peritz AE, Kierzek R, Sugimoto N, Turner DH (1991) Thermodynamic study of internal loops in oligoribonucleotides: symmetric loops are more stable than asymmetric loops. Biochemistry 30: 6428–6436
Piccirrilli JA, Krauch T, Moroney SE, Brenner SA (1990) Enzymatic incorporation of a new base pair into DNA and RNA extends the genetic alphabet. Nature 343:33–37
Pörschke D (1971) Cooperative nonenzymic base recognition II. Thermodynamics of the helix-coil transition of oligoadenylic + oligouridylic acids. Biopolymers 10:1989–2013
Pörschke D (1974) Thermodynamic and kinetic parameters of an oligonucleotide hairpin helix. Biophys Chem 1:381–386
Pörschke D, Eigen M (1971) Co-operative non-enzymic base recognition III. Kinetics of the oligoribouridylic-oligoriboadenylic acid system and of oligoriboadenylic acid alone at acidic pH. J Mol Biol 62:361–381
Press WH, Flannery BP, Teukolsky SA, Vetterling WT (1988) Numerical recipes. Cambridge University Press, New York
Schuster P (1991) Complex optimization in an artificial RNA world. In: Farmer D, Langton C, Rasmussen S, Taylor C ed.: Artificial Life II, SFI Studies in the Science of Complexity XII. Addison-Wesley, Reading, Mass
Stadler PF, Happel R (1992) Correlation structure of the landscape of the graph-bipartitioning-problem. J Phys A: Math Gen 25: 3103–3110
Stadler PF, Schnabl W (1992) The landscape of the travelling salesman problem. Phys Lett A 161:337–344
Tuerk CTC, Gold L (1990) Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science 249:505–510
Waterman MS, Smith TF (1978) RNA secondary structure: a complete mathematical analysis. Math Biosci 42:257–266
Weinberger ED (1990) Correlated and uncorrelated fitness landscapes and how to tell the difference. Biol Cybern 63:325–336
Weinberger ED (1991) Local properties the N-k model, a tuneably rugged energy landscape. Phys Rev A44:6399–6413
We inberger ED, Stadler PF (1992) Why some fitness landscapes are fractal. J Theor Biol (submitted for publication)
Zuker M (1989) The use of dynamic programming algorithms in RNA secondary structure prediction. In: Waterman MS, Mathematical methods for DNA sequences. CRC Press, Boca Raton
Zuker M, Stiegler P (1981) Optimal computer folding of large RNA sequences using thermodynamic and auxiliary information. Nucl Acid Res 9:133–148
Zuker M, Sankoff D (1984) RNA secondary structures and their prediction. Bull Math Biol46:591–621
Author information
Authors and Affiliations
Additional information
Correspondence to: P. Schuster
Rights and permissions
About this article
Cite this article
Tacker, M., Fontana, W., Stadler, P.F. et al. Statistics of RNA melting kinetics. Eur Biophys J 23, 29–38 (1994). https://doi.org/10.1007/BF00192203
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00192203