Abstract
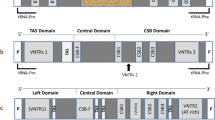
Twenty-one different caprine and 13 ovine MHC-DRB exon 2 sequences were determined including part of the adjacent introns containing simple repetitive (gt)n(ga)m elements. The positions for highly polymorphic DRB amino acids vary slightly among ungulates and other mammals. From man and mouse to ungulates the basic (gt)n(ga)m structure is fixed in evolution for 7 × 107 years whereas ample variations exist in the tandem (gt)n and (ga)m dinucleotides and especially their “degenerated” derivatives. Phylogenetic trees for the α-helices and β-pleated sheets of the ungulate DRB sequences suggest different evolutionary histories. In hoofed animals as well as in humans DRB β-sheet encoding sequences and adjacent intronic repeats can be assembled into virtually identical groups suggesting coevolution of noncoding as well as coding DNA. In contrast a-helices and C-terminal parts of the first DRB domain evolve distinctly. In the absence of a defined mechanism causing specific, site-directed mutations, double-recombination or gene-conversion-like events would readily explain this fact. The role of the intronic simple (gt)n(ga)m repeat is discussed with respect to these genetic exchange mechanisms during evolution.
Similar content being viewed by others
References
Acha-Orbea H, Scarpellino L (1991) Nonobese diabetic and nonobese nondiabetic mice have unique MHC class II haplotypes. Immunogenetics 34: 57–59
Ammer H, Schwaiger F-W, Kammerbauer C, Gomolka M, Arriens A, Lazary S, Epplen JT (1992) Exonic polymorphism versus intronic simple repeat hypervariability in MHC-DRB-genes. Immunogenetics 32: 330–337
Andersson L, Sigurdardottir S, Borsch C, Gustafsson K (1991) Evolution of MHC polymorphism: extensive sharing of polymorphic sequence motifs between human and bovine DRB alleles. Immunogenetics 33: 188–193
Ayane M, Mengle-Gaw L, McDevitt HO, Benoist C, Mathis D (1986) E-alpha-u and E-beta-u chain association: Where lies the anomaly? J Immunol 137: 948–951
Bandelt H-J, Dress AWM (1989) Weak hierarchies associated with similarity measures-an additive clustering technique. Bull Math Biol 51: 133–166
Begovich AB, Vu TH, Jones PP (1990) Characterization of the molecular defects in the mouse E-beta-f and E-beta-q genes: Implications for the origin of MHC polymorphism. J Immunol 144: 1957–1964
Belich MP, Madrigal JA, Hildebrand WH, Zemmour J, Williams RC, Luz R, Peztl-Erler ML, Parham P (1992) Unusual HLA-B alleles in two tribes of Brazilian Indians. Nature 357: 326–329
Bodmer JG, Marsh SGE, Albert E (1990) Nomenclature for factors of the HLA system. Immunol Today 11: 3–10
Braunstein NS, Germain RN (1986) The mouse E-beta-2 gene: a class II MHC beta gene with limited intraspecies polymorphism and an unusual pattern of transcription. EMBO J 5: 2469–2476
Brown JH, Jardetzky T, Saper MA, Samraoui B, Bjorkman PJ, Wiley DC (1988) A hypothetical model of the foreign antigen binding site of class II histocompatibility molecules. Nature 332: 845–850
Cam P, Jouvin-Marche E, LeGuern C, Marche PN (1990) Structure of class II genes in wild mouse Mus saxicola: functional and evolutionary implications. Eur J Immunol 20: 1337–1343
Chao NJ, Timmerman L, McDevitt HO, Jacob CO (1989) Molecular characterization of MHC class II antigens (beta-1 domains) in the bb diabetes-prone and -resistant rat. Immunogenetics 29: 231–234
Epplen JT, Ammer H, Epplen C, Kammerbauer C, Mitreiter R, Roewer L, Schwaiger W, Steimle V, Zischler H, Albert E, Andreas A, Beyermann B, Meyer W, Buitkamp J, Nanda I, Schmid M, Nürnberg P, Pena SDJ, Pöche H, Sprecher W, Schartl M, Weising K, Yassouridis A (1991) DNA fingerprinting: Approaches and applications. Birkhäuser-Verlag, Basel
Erlich HA, Gyllenstein UB (1991) Shared epitopes among HLA class II alleles: gene conversion, common ancestry and balancing selection. Immunol Today 12: 411–414
Falk K, Rötzschke O, Stevanovic S, Jung G, Rammensee H (1991) Allele specific motifs revealed by sequencing of selfpeptides eluted from MHC molecules. Nature 351: 290–296
Felsenstein J (1988) Phylogeny from molecular sequences: inference and reliability. Ann Rev Genet 22: 521–565
Figueroa F, Gutknecht J, Tichy H, Klein J (1990) Class II Mhc genes in rodent evolution. Immunol Rev 113: 207–226
Fitch DHA, Mainone C, Goodman M, Slightom JL (1990) Molecular history of gene conversion in the primate fetal τ-globin genes. J Biol Chem 265: 781–793
Groenen MAM, van der Poel JJ, Dijkhof RJM, Giphart RJ (1990) The nucleotide sequence of bovine MHC class 11 DQB and DRB genes. Immunogenetics 31: 37–44
Gorski J, Mach B (1986) Polymorphism of human la antigens: gene conversion between two DR β loci results in a new HLA-D/DR specifity. Nature 322: 67–70
Gorski J, Hayes CE (1990) The I-J-disparate mouse strains B10.A(3R) and B10.A(5R) have identical I-E beta sequences, Immunogenetics 39: 127–129
Gyllenstein U, Sundvall M, Ezcurra I, Erlich HA (1991a) Genetic diversity at class II DRB Loci of the primate MHC. J Immunol 146: 4368–4376
Gyllenstein U, Sundvall M, Erlich HA (1991b). Allelic diversity is generated by intraexon sequence exchange at the DRB1 locus of primates. Proc Natl Acad Sci USA 88: 3686–3690
Hughes AL, Nei M (1989) Nucleotide substitution at major histocompatibility complex class II loci: Evidence for overdominant selection. Proc Natl Acad Sci USA 86: 958–962
Kasahara M, Klein D, Weimin F, Gutknecht J (1990) Evolution of the class II major histocompatibility complex alleles in higher primates. Immunol Rev 113: 65–82.
Kappes D, Strominger JL (1988) Human class II major histocompatibility complex genes and proteins. Annu Rev Biochem 57: 991–1028
Klein J (1980) Generation of diversity at MHC loci: implication for T cell receptor repertoires. In: Fougereau M, Dausset D (eds) Immunology 80. Academic Press, London, pp 239–253
Klein J (1987) Origin of the major histocompatibility complex polymorphism: the transspecies hypothesis. Hum Immunol 19: 155–162
Kuhner MK, Peterson MJ (1992) Genetic exchange in the evolution of the human MHC class II loci. Tissue Antigens 39: 209–215
Legel S (1989) Nutztiere der Tropen and Subtropen (Legel S, (ed) Hirzel-Verlag, Leipzig
Madden DR, Gorga JC, Strominger JL, Wiley DC (1991) The structure of HLA-B27 reveals nonamer self-peptides bound in an extended conformation. Nature 353: 321–325
Marsh SGE, Bodmer J (1993) HLA class II nucleotide sequences, 1992. Immunogenetics 37: 79–94
Mäueler W, Muller M, Köhne AC, Epplen JT (1992) A gel retardation assay system for studying protein binding to simple repetitive DNA sequences. Electrophoresis 13: 7–10
Mengle-Gaw L, Conner S, McDevitt HO, Fathman CG (1984) Gene conversion between murine class II major histocompatibility complex loci. J Exp Med 160: 1184–1194
Mengle-Gaw L, McDevitt HO (1985) Predicted protein sequence of the murine I-E-S-beta polypeptide chain from cDNA and genomic clones. Proc Natl Acad Sci USA 82: 2910–2914
Miller SA, Dykes DD, Polesky HF (1988) A simple salting out procedure for extracting DNA from human nucleated cells. Nucleic Acids Res 16: 1215
Nei M, Gobori T (1986) Simple methods for estimating the numbers of synonymous and nonsynonymous nucleotide substitutions. Mol Biol Evol 3: 418–426
Ogawa S, Nishimura H, Awaji M, Nozawa S, Hirose S, Shirai T (1990) Nucleotide sequence analysis of MHC Class II genes in autoimmune disease-prone (NZB X NZW)Fl mice. Immunogenetics 32: 63–67
Potts WK, Wakeland EK (1990) Evolution of diversity at the major histocompatibility complex. Tree 5(6): 181–186
Rie\ O, Kammerbauer C, Roewer L, Steimle V, Andreas A, Albert E, Nagai T, Epplen JT (1990) Hypervariability of intronic simple (gt)n(ga)m repeats in HLA-DRB genes. Immunogenetics 32: 110–116
Saito H, Maki RA, Clayton LK, Tonegawa S (1983) Complete primary structures of the e-beta chain and gene of the mouse major histocompatibility complex. Proc Natl Acad Sci USA 80: 5520–5524
Saitou N, Nei M (1987) The neighbor joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4(4): 406–425
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning. 2nd ed. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY
Schwaiger FW, Buitkamp J, Weyers E, Epplen JT (1993) Typing of MHC-DRB genes with the help of intronic simple repeated DNA sequences. Mol Ecol 2: 55–59
Scott PC, Maddox JF, Gogolin-Ewens KJ, Brandon MR (1991) The nucleotide sequence and evolution of ovine MHC class II B Genes: DQB and DRB. Immunogenetics 34: 80–87
Sigurdardottir S, Borsch C, Gustafsson K, Andersson L (1992) Exon encoding the antigen-binding site of MHC class II betachains is divided into two subregions with different evolutionary histories. J Immunol 148: 968–973
Sneath PHA, Sokal RR (1973) Numerical taxonomy. The principles and practice of numerical classification. Freeman and Co, San Francisco
Thenius E (1979) Die Evolution der Säugetiere. Gustav Fischer Verlag, Stuttgart, New York
Vu TH, Tacchini-Cottier FM, Day CE, Begovich AB, Jones PP (1988) Molecular basis for the defective expression of the mouse E-w17-Beta gene. J Immunol 141: 3654–3661
Wada K, Wada Y, Doi H, Ishibashi F, Gojobori T, Ikemura T (1991) Codon usage tabulated from the GenBank genetic sequence data. Nucleic Acid Res 19 Suppl: 1981–1985
Wakeland EK, Boehme S, She JX (1990) The generation and maintenance of MHC class II gene polymorphism in rodents. Immunol Rev 113: 39–46
Watkins DI, McAdam SN, Liu X, Strang CR, Milford EL, Levine CG, Garber TL, Dogon AL, Lord CI, Ghim SH, Troup GM, Hughes AL, Letvin NL (1992) New recombinant HLA-B alleles in a tribe of South American Amerindians indicate rapid evolution of MHC class I loci. Nature 357: 329–333
Zharkikh AA, Rzhetsky A, Morosov PS, Sitnikova TL, Krushkal JS (1991) VOSTORG: a package of microcomputer programs for sequence analysis and construction of phylogenetic trees. Gene 101: 251–254
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Schwaiger, FW., Weyers, E., Epplen, C. et al. The paradox of MHC-DRB exon/intron evolution: α-helix and β-sheet encoding regions diverge while hypervariable intronic simple repeats coevolve with β-sheet codons. J Mol Evol 37, 260–272 (1993). https://doi.org/10.1007/BF00175503
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00175503