Summary
We compare the behavior of the genetic distance between individuals in evolving populations for three stochastic models.
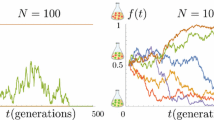
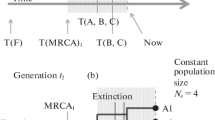
In the first model reproduction is asexual and the distribution of genetic distances reflects the genealogical tree of the population. This distribution fluctuates greatly in time, even for very large populations.
In the second model reproduction is sexual with random mating allowed between any pair of individuals. In this case, the population becomes homogeneous and the genetic distance between pairs of individuals has small fluctuations which vanish in the limit of an infinitely large population.
In the third model reproduction is still sexual but instead of random mating, mating only occurs between individuals which are genetically similar to each other. In that case, the population splits spontaneously into species which are in reproductive isolation from one another and one observes a steady state with a continual appearance and extinction of species in the population. We discuss this model in relation to the biological theory of speciation and isolating mechanisms.
We also point out similarities between these three models of evolving populations and the theory of disordered systems in physics.
Similar content being viewed by others
References
Abbott LF (1988) A model of autocatalytic replication. J Mol Evol 27:114
Amitrano C, Peliti L, Saber M (1989) Population dynamics in a spin-glass model of chemical evolution. J Mol Evol 29:513
Binder K, Young AP (1986) Spin glasses: experimental facts, theoretical concepts and open questions. Rev Mod Phys 58: 801
Bishop MJ, Friday AE (1985) Evolutionary trees from nucleic acid and protein sequences. Proc Roy Soc Lond B226:271
Blaisdell BE (1989) Effectiveness of measures requiring and not requiring prior sequence alignment for estimating the dissimilarity of natural sequences. J Mol Evol 29:526
Crosby JL (1970) The evolution of genetic discontinuity: computer models of the selection of barriers to interbreeding between species. Heredity 25:253
Crow JF, Kimura M (1970) An introduction to population genetics theory. Harper and Row, New York
Derrida B, Bessis D (1988) Statistical properties of valleys in the annealed random map model. J Phys A Math Gen 21:L509
Derrida B, Flyvbjerg H (1987a) The random map model: a disordered system with deterministic dynamics. J Phys France 48:971
Derrida B, Flyvbjerg H (1987b) Statistical properties of randomly broken objects and of multi-valley structures in disordered systems. J Phys A Math Gen 20:5273
Derrida B, Peliti L (1991) Evolution in a flat fitness landscape. Bull Math Biol 53:355
Epstein H, Ruelle D (1989) Test of a probabilistic model of evolutionary success. Physics Reports 184:289
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol 17:368
Fontana W, Schnabl W, Schuster P (1989) Physical aspects of evolutionary optimization and adaptation. Phys Rev A 40: 3301
Fontanari JF (1991) The adaptive map model. J Phys A Math Gen 24:L615
Goodman M (1981) Decoding the pattern of protein evolution. Prog Biophys Mol Biol 38:105
Grant V (1991) The evolutionary process. Columbia University Press, New York
Higgs PG, Derrida B (1991) Stochastic models for species formation in evolving population. J Phys A Math Gen 24:L985
Higgs PG, Orland H (1991) Scaling of polyelectrolytes and polyamphlytes—Simulation by an ensemble growth method. J Chem Phys 95:4506
Kauffman SA (1989) Lectures in the science of complexity. In Stein DL (ed) (Proceedings of the Summer School on Complex Systems, Santa Fe 1988). Addison-Wesley, Reading MA
Kauffman SA, Levin S (1987) Towards a general theory of adaptive walks in rugged fitness landscapes. J Theor Biol 128:11
Kimura M (1983) The neutral theory of molecular evolution. Cambridge University Press, Cambridge
Li WH, Nei M (1975) Drift variances of heterozygosity and genetic distance in transient states. Genet Res Camb 25:229
Maynard Smith J (1966) Sympatric speciation. American Naturalist 100:637
Maynard Smith J (1989) Evolutionary genetics. Oxford University Press, Oxford
Mayr E (1970) Populations, species and evolution. Harvard University Press, Cambridge
Mézard M, Parisi G, Virasoro MA (1987) Spin glass theory and beyond. World Scientific, Singapore
Peliti L (1990) A spin glass model of chemical evolution. Physica A 168:619
Rammal R, Toulouse G, Virasoro MA (1986) Ultrametricity for physicists. Rev Mod Phys 58:765
Rokhsar DS, Anderson PW, Stein DL (1986) Self-organization in prebiological systems: simulation of a model for the origin of genetic information. J Mol Evol 23:119
Sankoff D, Kruskal JB (1983) Time warps, string edits, and macromolecules: theory and practice of sequence comparison. Addison-Wesley, Reading MA
Schuster P, Swetina J (1988) Stationary mutant distributions and evolutionary optimization. Bull Math Biol 50:635
Serva M, Peliti L (1991) A statistical model of an evolving population with sexual reproduction. J Phys A Math Gen 24:L705
Shakhnovich EI, Gutin AM (1989) Formation of a unique structure in polypeptide chains. Theoretical investigation with the aid of a replica approach. Biophys Chem 34:187
Sokal RR, Sneath PHA (1963) Principles of numerical taxonomy. WH Freeman, San Francisco
Stewart FM (1976) Variability in the amount of heterozygosity maintained by neutral mutations. Theor Pop Biol 9:188
Tarazona P (1991) Error thresholds for molecular quasispecies as phase transitions: from simple landscapes to spin glass models. Preprint
Wright S (1931) Evolution in Mendelian populations. Genetics 16:97
Zhang YC, Serva M, Polikarpov M (1990) Diffusion reproduction processes. J Stat Phys 58:849
Author information
Authors and Affiliations
Additional information
Offprint requests to: P.G. Higgs
Rights and permissions
About this article
Cite this article
Higgs, P.G., Derrida, B. Genetic distance and species formation in evolving populations. J Mol Evol 35, 454–465 (1992). https://doi.org/10.1007/BF00171824
Received:
Revised:
Issue Date:
DOI: https://doi.org/10.1007/BF00171824