Abstract
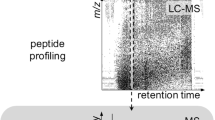
Bottom-up and top-down strategies are two commonly used methods for mass spectrometry (MS) based protein identification; each method has its own advantages and disadvantages. In this chapter, we describe an integrated top-down and bottom-up approach facilitated by concurrent liquid chromatography-mass spectrometry (LC-MS) analysis and fraction collection for comprehensive high-throughput intact protein profiling. The approach employs a high resolution reversed phase (RP) LC separation coupled with LC eluent fraction collection and concurrent on-line MS with a high field (12 T) Fourier-transform ion cyclotron resonance (FTICR) mass spectrometer. Protein elusion profiles and tentative modified protein identification are made using detected intact protein mass in conjunction with bottom-up protein identifications from the enzymatic digestion and analysis of corresponding LC fractions. Specific proteins of biological interest are incorporated into a target ion list for subsequent off-line gas-phase fragmentation that uses an aliquot of the original collected LC fraction, an aliquot of which was also used for bottom-up analysis.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Aebersold, R., Mann, M. (2003) Mass spectromety-based proteomics. Nature 422, 198–207.
Kelleher, N. L. (2004) Top-down proteomics. Anal. Chem. 76, 197A–203A.
McLafferty, F. W., Breuker, K., Jin, M., Han, X., Infusini, G., Jiang, H., Kong, X., Begley, T. P. (2007) Top-down MS, a powerful complement to the high capabilities of proteolysis proteomics. FEBS J. 274, 6256–6268.
Parks, B. A., Jiang, L., Thomas, P. M., Wenger, C. D., Roth, M. J., Ii, M. T., Burke, P. V., Kwast, K. E., Kelleher, N. L. (2007) Top-down proteomics on a chromatographic time scale using linear ion trap Fourier transform hybrid mass spectrometers. Anal. Chem. 79, 7984–7991.
Siuti, N., Kelleher, N. L. (2007) Decoding protein modifications using top-down mass spectrometry. Nat. Methods 4, 817–821.
Zabrouskov, V., Whitelegge, J. P. (2007) Increased coverage in the transmembrane domain with activated-ion electron capture dissociation for top-down Fourier-transform mass spectrometry of integral membrane proteins. J. Proteome Res. 6, 2205–2210.
Zubarev, R. A., Kelleher, N. L., McLafferty, F. W. (1998) Electron capture dissociation of multiply charged proteins cations. A nonergodic process. J. Am. Chem. Soc. 120, 3265–3266.
Coon, J. J., Ueberheide, B., Syka, J. E. P., Dryhurst, D. D., Ausio, J., Shabanowitz, J., Hunt, D. F. (2005) Interpreting the protein language using proteomics. Proc. Natl. Acad. Sci. USA 102, 9463–9468.
Sharma, S., Simpson, D. C., Tolic, N., Jaitly, N., Mayampurath, A. M., Smith, R. D., Pasa-Tolic, L. (2007) Probing proteomes using capillary isoelectric focusing-electrospray ionization Fourier transform ion cyclotron resonance mass spectrometry. J. Proteome Res. 6, 602–610.
Wu, S., Lourette, N. M., Tolic, N., Zhao, R., Robinson, E. W., Tolmachev, A. V., Smith, R. D., Pasa-Tolic, L. (2009) An integrated top-down and bottom-up strategy for broadly characterizing protein isoforms and modifications. J. Proteome Res. 8, 1347–1357.
Shen, Y., Tolic, N., Masselon, C., Pasa-Tolic, L., Camp, D. G., Hixson, K. K., Zhao, R., Anderson, G. A., Smith, R. D. (2004) Ultrasensitive proteomics using high-efficiency on-line micro-SPE-nanoLC-nanoESI MS and MS/MS. Anal. Chem. 76, 144–154.
Tolmachev, A. V., Robinson, E. W., Wu, S., Kang, H., Pasa-Tolic, L., Smith, R. D. (2008) Trapped-ion cell with improved DC potential harmonicity for FT-ICR MS. J. Am. Soc. Mass Spectrom. 19, 586–597.
Yates, J. R., Eng, J. K., McCormack, A. L., Schieltz, D. (1995) Method to correlate tandem mass spectra of modified peptides to amino acid sequences in the protein database. Anal. Chem. 67, 1426–1436.
Washburn, M. P., Wolters, D., Yates, J. R. (2001) Large-scale analysis of the yeast proteome by multidimensional protein identification technology. Nat. Biotechnol. 19, 242–247.
Monroe, M. E., Tolic, N., Jaitly, N., Shaw, J. L., Adkins, J. N., Smith, R. D. (2007) VIPER: an advanced software package to support high-throughput LC-MS peptide identification. Bioinformatics 23, 2021–2023.
LeDuc, R. D., Taylor, G. K., Kim, Y. B., Januszyk, T. E., Bynum, L. H., Sola, J. V., Garavelli, J. S., Kelleher, N. L. (2004) ProSight PTM: an integrated environment for protein identification and characterization by top-down mass spectrometry. Nucleic Acids Res. 32, W340–W345.
Zamdborg, L., LeDuc, R. D., Glowacz, K. J., Kim, Y. B., Viswanathan, V., Spaulding, I. T., Early, B. P., Bluhm, E. J., Babai, S., Kelleher, N. L. (2007) ProSight PTM 2.0: improved protein identification and characterization for top down mass spectrometry. Nucleic Acids Res. 35, W701–W706.
Shen, Y., Tolic, N., Hixson, K. K., Purvine, S. O., Anderson, G. A., Smith, R. D. (2008) De novo sequencing of unique sequence tags for discovery of post-translational modifications of proteins. Anal. Chem. 80, 7742–7754.
Acknowledgments
Portions of this work were supported by the National Center for Research Resources (RR 018522), the National Institute of Allergy and Infectious Diseases (NIH/DHHS through interagency agreement Y1-AI-4894-01), the National Institute of General Medical Sciences (NIGMS, R01 GM063883), and the U. S. Department of Energy (DOE) Office of Biological and Environmental Research. Work was performed in the Environmental Molecular Science Laboratory, a DOE national scientific user facility located on the campus of Pacific Northwest National Laboratory (PNNL) in Richland, Washington. PNNL is a multi-program national laboratory operated by Battelle for the DOE under Contract DE-AC05-76RLO 1830.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2011 Springer Science+Business Media, LLC
About this protocol
Cite this protocol
Wu, S., Tolić, N., Tian, Z., Robinson, E.W., Paša-Tolić, L. (2011). An Integrated Top-Down and Bottom-Up Strategy for Characterization of Protein Isoforms and Modifications. In: Wu, C., Chen, C. (eds) Bioinformatics for Comparative Proteomics. Methods in Molecular Biology, vol 694. Humana Press. https://doi.org/10.1007/978-1-60761-977-2_18
Download citation
DOI: https://doi.org/10.1007/978-1-60761-977-2_18
Published:
Publisher Name: Humana Press
Print ISBN: 978-1-60761-976-5
Online ISBN: 978-1-60761-977-2
eBook Packages: Springer Protocols