Abstract
Ensembl Plants (http://plants.ensembl.org) offers genome-scale information for plants, with four releases per year. As of release 47 (April 2020) it features 79 species and includes genome sequence, gene models, and functional annotation. Comparative analyses help reconstruct the evolutionary history of gene families, genomes, and components of polyploid genomes. Some species have gene expression baseline reports or variation across genotypes. While the data can be accessed through the Ensembl genome browser, here we review specifically how our plant genomes can be interrogated programmatically and the data downloaded in bulk. These access routes are generally consistent across Ensembl for other non-plant species, including plant pathogens, pests, and pollinators.
You have full access to this open access chapter, Download protocol PDF
Similar content being viewed by others
Key words
1 Introduction
Plants play a central role in the ecology and economy of our planet and are essential to our food security. As the world population increased by 145% in the last 60 years, the yields of cereals increased even more, while not needing much more land [1]. This has been possible as a result of improved agricultural practices and crops. Currently, breeding programs take advantage of inexpensive genomic and phenotypic data. The next steps towards what is being called Breeding 4.0 [2] include adapting crops to changing environments and broadening the diversity pool to compensate for the losses occurred during domestication. For this reason wild relatives of crops are being sequenced increasingly and added to pre-breeding programs [3]. In addition, natural plant populations and model plants are being studied to understand their ecology and the genetic basis of their adaptation mechanisms, which can then be applied to in crops. In this context, genomics is a foundation of plant sciences, as standard approaches such as marker-assisted breeding, QTL analysis, and genome-wide association studies, as well as genomic selection, induced variation experiments, and genome editing, all depend on genomic technologies and databases. These tools are accelerating breeding and helping to untangle complex polyploid genomes, such as that of bread wheat [4].
Ensembl Plants (http://plants.ensembl.org) is the Ensembl portal for plants and red algae [5] and provides a consistent set of interfaces to genomic data, including reference genome sequences, gene and transcript models, genetic variation, gene expression, markers, and comparative genomics. There are up to four releases per year. At the time of writing, the latest release of Ensembl Plants is version 47 (April 2020), which corresponds to Ensembl version 100. This release comprises 79 genomes, containing several cultivars and ecotypes for some species. Ensembl Plants is developed with our long-term partners Gramene [6] and with individual groups that publish plant genomes around the world. This chapter documents how the data at Ensembl Plants can be downloaded in bulk and interrogated programmatically using a variety of approaches. It provides a series of recipes, available as source code at https://github.com/Ensembl/plant-scripts, that can be modified to carry out more complex analyses of plant genomes.
2 Materials
2.1 Database Structure and Data Access
Ensembl Plants is implemented primarily as a collection of MySQL relational databases. The overall data structure is modular, with different data (e.g., core annotation, comparative genomics, functional genomics, variation data) modeled by distinct schemas. The core schema is modeled on the central dogma of molecular biology, linking genome sequence to genes, transcripts, and their translations, each of which can be decorated with functional annotation (see Note 1). Much annotation takes the form of cross-references, which are web links to entries in other resources, such as InterPro [7] or Gene Ontology [8], that either represent the primary source of the biological entity or provide additional information. Cross-references describe functional entities such as domains, reactions, and processes. Some also serve as controlled vocabularies for functional annotation.
The databases can be downloaded for local installation or alternatively accessed via a public MySQL server. Local MySQL databases are an efficient alternative to the public MySQL server, particularly if heavy use is anticipated (see Note 2). Programmatic access is supported by two APIs, which allow data discovery and access through an abstraction layer that hides the detailed structure of the underlying data store. One is a Perl API, while the other uses a language-agnostic REST interface [9]. The REST service allows up to 15 requests per second.
In addition to the primary databases, Ensembl Plants also provides access to denormalized data warehouses, constructed using the BioMart tool kit [10]. These are specialized databases that support efficient gene- and variant-centric queries. Finally, a variety of data selections are exported from the databases in common file formats and made available for download via an FTP site.
These resources are summarized in Table 1. Recipes to query each of them are listed in Table 7.
2.2 Overview of Data Content
2.2.1 Genomes and Core Data
Genome assemblies are typically imported from the European Nucleotide Archive (ENA) [11], which is part of the International Nucleotide Sequence Database Collaboration (http://www.insdc.org, INSDC). Gene model annotations are imported from the ENA [11], Phytozome [12], or provided by community members (see Note 3). For instance, the rice annotation was imported from RAP-DB [13]. After import, various computational analyses are performed for each genome. A summary of these is given in Table 2. In addition, specific datasets are imported and analyzed according to the requirements of individual communities. These datasets typically fall into two classes, markers, and variants across genotype panels.
The genomes currently included in Ensembl Plants are listed in Table 3. A summary of UniProt coverage of proteins encoded by genes within these genomes is given in Table 4 [17]. In all cases, genomes are identified by their Ensembl production name, which is usually binomial but can also include a strain name to distinguish particular cultivars or ecotypes, such as malus_domestica_golden. Details of other datasets incorporated can be found through the homepage for each species (see Note 3).
2.2.2 Variation Data
The variation schema can store genetic variants observed in populations or germplasm collections, alleles, and frequencies, alongside sample genotype data. Supported variant types include single nucleotide polymorphisms, indels, and structural variants. The functional consequence of variants on genes is predicted with the Ensembl Variant Effect Predictor (VEP) [14]. Linkage disequilibrium data and statistical associations with phenotypes are available for selected species. The variation datasets of release 47 of Ensembl Plants are described in Table 5. The Ensembl VEP is also a command line tool that can be used to efficiently annotate variants and we provide recipes for it as well (see Table 7).
2.2.3 Comparative Genomics Data
The Ensembl Gene Tree pipeline is used to calculate evolutionary relationships among members of protein families (Table 2). For each gene, the translation of the canonical transcript is selected (see Note 4). Briefly, this pipeline first finds clusters of similar proteins and then, for each cluster, attempts to reconcile the relationship between the sequences with the known species cladogram (Fig. 1), derived from the NCBI Taxonomy database [42]. The analysis also contains a few non-plant outgroups. The TreeBeST software (https://github.com/Ensembl/treebest) is used to construct a consensus tree, which allows the identification of orthologues and paralogues. As polyploid genomes are split into components, homoeologous genes are effectively defined as orthologues among subgenomes. A number of plant genomes are also included in a pan-taxonomic gene tree, containing a representative selection of sequenced genomes from all domains of life. Recipe R2 can be used to check which comparative analyses have been run for a particular species. This information is also displayed in the table at http://plants.ensembl.org/species.html.
Species cladogram of release 47 (April 2020) of Ensembl Plants. Genomes of polyploid species are decomposed into genomic components. This topology is used in the comparative genomic analyses to derive orthologous and paralogous genes. This tree was produced with the Newick file obtained with recipe F12 and visualized with iToL [41]
Other comparative analyses available in Ensembl Plants are pairwise whole-genome alignments and synteny (see Tables 2 and 6).
2.2.4 Baseline Expression Data
Baseline gene expression reports are available as “Gene expression” on the website for selected species. An example for barley is shown at http://plants.ensembl.org/Hordeum_vulgare/Gene/ExpressionAtlas?g=HORVU5Hr1G095630;r=chr5H:599085656-599133086. The underlying curated expression data, produced by Expression Atlas [44], can be browsed and downloaded via the expression widget.
2.2.5 RNA-seq Tracks
RNA-seq datasets from the public INSDC archives are mapped to genome assemblies in Ensembl Plants in every release. They are handled as ENA studies and for each of them CRAM files are created with the RNA-Seq-er pipeline (https://www.ebi.ac.uk/fg/rnaseq/api) [45] and published at ftp://ftp.ensemblgenomes.org/pub/misc_data/Track_Hubs. Each study contains a separate folder for each assembly that was used for mapping. These tracks can be interactively displayed in the browser, but can be of interest for high-throughput studies as well. For instance, study SRP133995 was mapped to tomato assembly SL3.0 and the tracksDb.txt file therein indicates the full path to the relevant CRAM file next to its metadata. CRAM files for a selected assembly can be discovered with recipe C1; note that the assembly name corresponds to column “assembly_default” in recipe R2. As of May 2020 there were 89,355 CRAM files available.
3 Methods
This section describes some of the recipes listed in Table 7 in detail so that the reader can execute or modify any of them. Software dependencies required by these recipes are listed in https://github.com/Ensembl/plant-scripts/blob/master/README.md.
The different approaches are complementary. While the native Perl API is the most powerful and used extensively by Ensembl developers, it also requires some Perl knowledge and the installation of several repositories. Similarly, the Biomart and MySQL examples require knowledge of R and SQL, respectively. However, the REST endpoints can be interrogated with any programming language; however, only a defined set of queries are currently supported. The FTP recipes allow efficient bulk downloads, but with no customization. The source code for all recipes can be found at https://github.com/Ensembl/plant-scripts.
3.1 Clone the GitHub Repository and Install Dependencies
The following steps explain how to obtain a local copy of the recipes and how to test them on Linux/MacOS operating systems (OS).
-
1.
Open a terminal and check whether git is installed by typing: git --version.
-
2.
If required install git if using the appropriate software manager for your OS.
-
3.
Clone the repository: git clone https://github.com/Ensembl/plant-scripts.git.
-
4.
Navigate to the scripts directory: cd plant-scripts.
-
5.
Optionally test the scripts: perl demo_test.t.
3.2 Perl API Recipes
The Ensembl Perl API enables access to all types of data from Ensembl Plants (genes, variation, comparative genomics, regulation, etc.) and it is documented extensively (see Note 5). It allows complex queries to be executed without the construction of any explicit SQL queries. The repository contains eight Perl API recipes, of which three are described here (A1, A4, and A8).
3.2.1 Get a BED File with Repeats on Chromosome 4
-
1.
Load the Registry object with details of genomes available from the public Ensembl Genomes servers (recipe A1):
Use Bio::EnsEMBL::Registry; Bio::EnsEMBL::Registry->load_registry_from_db( -USER => 'anonymous', -HOST => 'mysql-eg-publicsql.ebi.ac.uk', -PORT => '4157', );
-
2.
Set species and chromosome of interest and print BED file with repeats (recipe A4). Ensembl uses 1-based inclusive coordinates internally:
my $species = 'arabidopsis_thaliana'; my $chrname = 'chr4'; my $slice_adaptor = Bio::EnsEMBL::Registry-> get_adaptor($species, 'core', 'Slice'); my $slice = $slice_adaptor-> fetch_by_region( 'toplevel', $chrname ); my @repeats = @{ $slice->get_all_RepeatFeatures() }; foreach my $repeat (@repeats) { printf("%s\t%d\t%d\t%s\t%s\t%s\n", $chrname, $repeat->start()-1, $repeat->end(), $repeat->analysis()->logic_name(), $repeat->repeat_consensus()->repeat_class(), $repeat->repeat_consensus()->repeat_type() ); }
3.2.2 Get Markers Mapped on Chromosome 1D of Bread Wheat
Only a few plants have markers loaded. Recipe A8 retrieves wheat KASP markers, with coordinates returned in BED format:
$species = 'triticum_aestivum'; $chrname = '1D'; $slice_adaptor = Bio::EnsEMBL::Registry-> get_adaptor( $species, 'Core', 'Slice' ); $slice = $slice_adaptor-> fetch_by_region( 'chromosome', $chrname ); foreach my $mf (@{ $slice->get_all_MarkerFeatures() }) { my $marker = $mf->marker(); printf("%s\t%d\t%d\t%s\t%s\t%s\t%d\n", $mf->seq_region_name(), $mf->start()-1, $mf->end(), $mf->display_id(), $marker->left_primer(), $marker->right_primer(), $marker->max_primer_dist() ); }
3.3 R Biomart Recipes
The BioMart databases can be queried in many ways (see Note 6). There are five recipes in the repository written in the R language. They all use the BioConductor package BiomaRt [46], which can be installed as follows:
if (!requireNamespace("BiocManager", quietly = TRUE)) install.packages("BiocManager") BiocManager::install("biomaRt")
This example corresponds to recipe R4, which queries sunflower genes to obtain annotated Pfam domains. Dataset names are abbreviations of Ensembl production names. See recipe R5 for an example querying BioMart variation databases:
EPgenes = useMart( biomart="plants_mart", host="plants.ensembl.org", dataset="hannuus_eg_gene") pfam = getBM( attributes=c("ensembl_gene_id", "pfam"), mart=EPgenes)
3.4 FTP Recipes
There are 12 recipes in the repository that query the Ensembl Genomes FTP server. They use shell variables and the wget program to download files. The recipes refer to the Ensembl release and the Ensembl Plants release as RELEASE and EGRELEASE, respectively. Recipe F5 involves a prewritten BioMart query.
3.4.1 Download Soft-Masked Genomic Sequences
Soft-masked sequences are FASTA files with all annotated repeated elements in lower case. Using recipe F4 they can be downloaded for a chosen species and release as follows:
SERVER=ftp://ftp.ensemblgenomes.org/pub DIV=plants EGRELEASE=47 SPECIES=Brachypodium_distachyon FASTASM="${SPECIES}*.dna_sm.toplevel.fa.gz" URL="${SERVER}/release-${EGRELEASE}/${DIV}/fasta/${SPECIES,,}/dna/${FASTASM}" wget -c "$URL"
3.4.2 Download All Homologies in a Single TSV File
Recipe F9 downloads a large file (several GB) with all homologies of a release in TSV format. Sequence identifiers correspond to canonical transcripts (see Note 4):
TSVFILE="Compara.${RELEASE}.protein_default.homologies.tsv.gz" URL="${SERVER}/${DIV}/release-${EGRELEASE}/tsv/ensembl-compara/homologies/${TSVFILE}" wget -c "$URL"
This file can be parsed in the command line in order to extract homologies (see Note 7):
zcat "$TSVFILE" | grep triticum_aestivum | grep oryza_sativa | grep ortholog
Homologies of each species can be retrieved from a smaller, specific file:
TSVFILE="Compara.${RELEASE}.protein_default.homologies.tsv.gz" SPECIES=Triticum_aestivum URL="${SERVER}/${DIV}/release-${EGRELEASE}/tsv/ensembl-compara/homologies/${SPECIES,,}${TSVFILE}" wget -c "$URL" zcat "$TSVFILE" | grep oryza_sativa | grep ortholog
Homologies can also be downloaded in OrthoXML format [47], which renders a smaller file but requires a more complex parser.
3.5 MySQL Recipes
Direct access to the public MySQL server requires knowledge of the schemas (see Notes 1 and 8). While this approach supports complex queries with high-performance, the schemas may change in a new release and thus some queries might stop working. For this reason, API access is recommended. Three recipes are shown here, they all require the mysql-client to be installed.
3.5.1 Count Protein-Coding Genes of a Particular Species
This is recipe S2. The source code works out the current release number, but it can also be set manually as in this example:
SERVER=mysql-eg-publicsql.ebi.ac.uk USER=anonymous PORT=4157 EGRELEASE=47 RELEASE=$((EGRELEASE + 53)) SPECIES=arabidopsis_thaliana SPECIESCORE=$(mysql --host $SERVER --user $USER --port $PORT \ -e "show databases" | grep \ "${SPECIES}_core_${EGRELEASE}_${RELEASE}") mysql --host $SERVER --user $USER --port $PORT \ $SPECIESCORE -e "SELECT COUNT(*) FROM gene \ WHERE biotype='protein_coding'"
3.5.2 Get stable_ids of Transcripts Used in Compara Analyses
Recipe S3 gets a list of identifiers of all transcript used in the comparative genomics gene tree analysis (see Note 4):
SERVER=mysql-eg-publicsql.ebi.ac.uk USER=anonymous PORT=4157 EGRELEASE=47 RELEASE=$((EGRELEASE + 53)) SPECIES=arabidopsis_thaliana mysql --host $SERVER --user $USER --port $PORT \ "ensembl_compara_plants_${EGRELEASE}_${RELEASE}" \ -e "SELECT sm.stable_id \ FROM seq_member sm, gene_member gm, genome_db gdb \ WHERE sm.seq_member_id = gm.canonical_member_id \ AND sm.genome_db_id = gdb.genome_db_id \ AND gdb.name = '$SPECIES'"
See recipe F3 to obtain the corresponding sequences.
3.5.3 Get Variants Significantly Associated with Phenotypes
Recipe S4 queries several tables of the variation schema (see Note 8):
SPECIESVAR=$(mysql --host $SERVER --user $USER --port $PORT \ -e "show databases" | \ grep "${SPECIES}_variation_${EGRELEASE}_${RELEASE}") mysql --host $SERVER --user $USER --port $PORT \ $SPECIESVAR<<SQL SELECT f.object_id, s.name, f.seq_region_start, f.seq_region_end, p.description FROM phenotype p JOIN phenotype_feature f ON p.phenotype_id = f.phenotype_id JOIN seq_region s ON f.seq_region_id = s.name WHERE f.type = 'Variation' AND f.is_significant=1 SQL
3.6 REST Recipes
The following recipes, written in Python, can also be found in R and Perl languages in the repository. They communicate with the Ensembl REST service at https://rest.ensembl.org (see Note 9) using the functions get_json and get_json_post, defined in file exampleREST.py.
3.6.1 Find Features Overlapping a Genomic Region
Recipe R3 queries the endpoint overlap/region and returns all features overlapping a selected genomic region:
def get_overlapping_features(species,region): overlap_url = ("/overlap/region/" + species + "/" + region) # repeat or variation could have been used instead of gene ext = (overlap_url + "?feature=gene;content-type=application/json") overlap_data = get_json(ext) for overlap_feat in overlap_data: print("%s\t%s\t%s" % (overlap_feat['id'], overlap_feat['start'], overlap_feat['end'])) species = 'triticum_aestivum'; region = '3D:379400000-379540000'; get_overlapping_features(species,region)
3.6.2 Check Consequences of SNPs Within CDS Sequences
Recipe R8 queries two endpoints (map/cds/ and info/vep/:species/region). The first one translates CDS to genomic coordinates, the second one retrieves the predicted consequences of the SNP in the coding sequence. This recipe can be used to annotate genomic variants in a given gene across germplasm panels, as done in [48]:
def check_snp_consequences(species,transcript_id,SNPCDScoord,SNPbase): # convert CDS coords to genomic coords ext = ("/map/cds/" + transcript_id + "/" + SNPCDScoord + ".." + SNPCDScoord + "?content-type=application/json;species=" + species) map_cds = get_json(ext) if map_cds['mappings'][0]['seq_region_name']: mapping = map_cds['mappings'][0] # fetch VEP consequences for this region SNPgenome_coord = ( mapping['seq_region_name'] + ':' + str(mapping['start']) + '-' + str(mapping['end']) ) ext = ("/vep/"+ species + "/region/" + SNPgenome_coord + "/" + SNPbase + "?content-type=application/json") conseq = get_json(ext) # Print all the relevant info for the given variant if conseq[0]['allele_string']: for tcons in conseq[0]['transcript_consequences']: #... some lines omitted, check exampleREST.py values = (transcript_id, SNPCDScoord, conseq[0]['allele_string'], tcons['biotype'], tcons['codons'], tcons['amino_acids'], tcons['protein_start'], tcons['impact'], tcons['sift_prediction'], tcons['sift_score']) for val in values: print (val, end="\t") print() species = 'triticum_aestivum' transcript_id = 'TraesCS4B02G042700.1' SNPCDScoord = '812' SNPbase = 'T' check_snp_consequences(species,transcript_id,SNPCDScoord,SNPbase)
3.7 Annotate the Effect of Variants with the Ensembl Variant Effect Predictor
The Ensembl VEP tool can be used to predict the effect of variants on genes, transcripts, and protein sequences (see Note 10). As mentioned in Table 2, this analysis is run for all genomic variants imported into Ensembl (see Table 5). While the Ensembl VEP is available through a web interface, the advantage of a local installation is that it can be used to analyze variation sets of any species, including species that are not in Ensembl Plants. If variants are mapped to a reference genome supported in Ensembl Plants, using a cache file increases performance. However, as shown in recipe V4, it is possible to use other reference FASTA files together with the corresponding GFF/GTF annotation files. The next steps summarize how the software is installed and used following recipes F8, V1, V2, and V3.
-
1.
Clone the repository: git clone https://github.com/Ensembl/ensembl-vep.git.
-
2.
Navigate to the Ensembl VEP directory: cd ensembl-vep.
-
3.
Install Ensembl VEP: perl INSTALL.pl.
-
4.
Download cache file with recipe F8
SPECIES=arabidopsis_thaliana VEPCACHE="${SPECIES,,}*.tar.gz*" URL="${SERVER}/${DIV}/release-${EGRELEASE}/variation/vep/${VEPCACHE}" wget -c "$URL"
-
5.
Unpack downloaded cache file and check SIFT support:
tar xfz $VEPCACHE grep sift "${SPECIES}/${EGRELEASE}_*/info.txt"
-
6.
Predict effect of variants, see Note 11:
EGRELEASE=47 VCFILE=ensembl-vep/examples/arabidopsis_thaliana.TAIR10.vcf VEPOPTIONS=( --genomes # Ensembl Genomes, for Plants --species $SPECIES --cache # use local cache file, opposed to --database --dir_cache ./ # path of unpacked cache $SPECIES folder --cache_version $EGRELEASE --input_file $VCFILE --output_file ${VCFILE}.vep --check_existing # co-located known variants --distance 5000 # max dist between variant and transcript --biotype # show biotype of neighbor transcript ) ensembl-vep/vep "${VEPOPTIONS[@]"
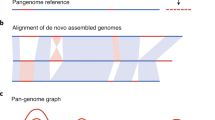
3.8 Querying Plant Pangenomes
Upcoming Ensembl Plants releases will have an increasing number of species with multiple cultivars or ecotypes as additional assemblies are added in collaboration with the relevant communities. On the website these cultivars can be browsed from the appropriate reference genome page such as http://plants.ensembl.org/Triticum_aestivum/Info/Strains?db=core (see Note 12). Starting with several UK cultivars in release 48 (August 2020), Ensembl will host all cultivars of the first assembled wheat pangenome [49] from release 50 planned for early 2021 (see example Fig. 2). Note that related noncultivated species are often included in the pangenomes of crops. For example, Ensembl Plants hosts 11 Oryza species plus the outgroup plant Leersia perrieri. Both types of genome sets can be considered pangenomes.
The RFL gene (TraesCS1B02G038500) lifted over from the reference landrace Chinese Spring to three wheat cultivars (CDC Landmark, Julius and Jagger). The genes are displayed in the Ensembl Plants genome browser. While in the first cultivar there are three annotated transcript isoforms including one with two exons, the others have a single transcript with one exon. Furthermore, the locus is annotated as a pseudogene in Julius
Currently, some pangenomes in Ensembl can be interrogated using gene trees and whole-genome alignments (WGAs; see Tables 2 and 6). For example, recipe A9 can be used to retrieve syntenic orthologous genes in rice or Brassicaceae species. These analyses will be available for wheat as well once de novo gene annotation and WGAs are produced.
3.9 Getting Help
Documentation for Ensembl Plants, including FAQs, tutorials, and detailed information about the project, datasets, and pipelines that we run can be found under the “Documentation” and “Website help” links at the top of every page. Detailed information for each species can be found on the species homepage. The EMBL-EBI train online website has several free courses on Ensembl, including the recently updated “Ensembl Genomes (non-chordates): Quick tour” (https://www.ebi.ac.uk/training/online/course/ensembl-genomes-non-chordates-quick-tour) and “Ensembl REST API” courses (https://www.ebi.ac.uk/training-beta/online/courses/ensembl-rest-api). Any data problems are reported on our blog http://www.ensembl.info/known-bugs. If the available documentation cannot answer your question, a helpdesk is provided (mail helpdesk@ensemblgenomes.org with your query).
4 Notes
-
1.
The core schema is fully described at https://www.ensembl.org/info/docs/api/core/core_schema.html. There are similar documents for variation, comparative genomics, and regulation schemas: https://www.ensembl.org/info/docs/api/variation/variation_schema.html
https://www.ensembl.org/info/docs/api/compara/compara_schema.html
https://www.ensembl.org/info/docs/api/funcgen/funcgen_schema.html
The schema for the metadata database can be found at https://github.com/Ensembl/ensembl-metadata.
-
2.
Instructions to set up a local Ensembl database are provided at http://plants.ensembl.org/info/docs/webcode/mirror/install/ensembl-data.html.
-
3.
Check the annotation page for each species in Ensembl Plants. For Arabidopsis thaliana, this is http://plants.ensembl.org/Arabidopsis_thaliana/Info/Annotation/#genebuild.
-
4.
Gene trees use canonical transcripts, defined at http://plants.ensembl.org/info/website/glossary.html. In plant species, the canonical transcript of a protein-coding gene is the transcript with the longest translation with no stop codons. This does not necessarily reflect the most biologically relevant transcript of a gene. The script https://github.com/Ensembl/plant_tools/blob/master/phylogenomics/ens_sequences.pl can be used to obtain sequences of canonical transcripts in FASTA format.
-
5.
Check http://plants.ensembl.org/info/docs/Doxygen. See also debugging instructions and tutorials at http://plants.ensembl.org/info/docs/api.
- 6.
-
7.
See http://plants.ensembl.org/info/genome/compara/homology_method.html for the definitions of the different homology types.
-
8.
The variation schema is described at http://plants.ensembl.org/info/docs/api/variation/variation_schema.html.
-
9.
Training material to learn more about the Ensembl REST interface can be found at https://www.ebi.ac.uk/training/online/course/ensembl-rest-api and https://mybinder.org/v2/gh/Ensembl/rest-api-jupyter-course/master. The different endpoints are documented at https://rest.ensembl.org/documentation.
-
10.
Ensembl VEP functionality can be extended to utilize additional data or run additional analyses using plugins, see https://www.ensembl.org/info/docs/tools/vep/script/vep_plugins.html.
-
11.
The full list of options of Ensembl VEP is described at http://www.ensembl.org/info/docs/tools/vep/script/vep_options.html, there are examples at http://www.ensembl.org/info/docs/tools/vep/script/vep_example.html.
-
12.
Reference genomes have binomial production names when possible, such as oryza_sativa (rice) or triticum_aestivum (bread wheat). Additional cultivars or ecotypes have longer trinomial names such as oryza_sativa_indica or triticum_aestivum_cadenza. Following this convention, theobroma_cacao_matina and panicum_hallii_hal2 will be renamed to theobroma_cacao and panicum_hallii by release 50.
References
Ritchie H, Roser M (2013) Crop yields. https://ourworldindata.org/crop-yields. Accessed 1 Jul 2020
Wallace JG, Rodgers-Melnick E, Buckler ES (2018) On the road to breeding 4.0: unraveling the good, the bad, and the boring of crop quantitative genomics. Annu Rev Genet 52:421–444
Arora S, Steuernagel B, Gaurav K et al (2019) Resistance gene cloning from a wild crop relative by sequence capture and association genetics. Nat Biotechnol 37:139–143
Adamski NM, Borrill P, Brinton J et al (2020) A roadmap for gene functional characterisation in crops with large genomes: lessons from polyploid wheat. elife 9:55646. https://doi.org/10.7554/eLife.55646
Howe KL, Contreras-Moreira B, De Silva N et al (2020) Ensembl Genomes 2020-enabling non-vertebrate genomic research. Nucleic Acids Res 48:D689–D695
Tello-Ruiz MK, Naithani S, Stein JC et al (2018) Gramene 2018: unifying comparative genomics and pathway resources for plant research. Nucleic Acids Res 46:D1181–D1189
Mitchell AL, Attwood TK, Babbitt PC et al (2019) InterPro in 2019: improving coverage, classification and access to protein sequence annotations. Nucleic Acids Res 47:D351–D360
The Gene Ontology Consortium, The Gene Ontology Consortium (2019) The Gene Ontology Resource: 20 years and still going strong. Nucleic Acids Res 47:D330–D338
Yates A, Beal K, Keenan S et al (2015) The ensembl REST API: ensembl data for any language. Bioinformatics 31:143–145
Kasprzyk A (2011) BioMart: driving a paradigm change in biological data management. Database 2011:bar049
Amid C, Alako BTF, Balavenkataraman Kadhirvelu V et al (2020) The European Nucleotide Archive in 2019. Nucleic Acids Res 48:D70–D76
Goodstein DM, Shu S, Howson R et al (2012) Phytozome: a comparative platform for green plant genomics. Nucleic Acids Res 40:D1178–D1186
Sakai H, Lee SS, Tanaka T et al (2013) Rice Annotation Project Database (RAP-DB): an integrative and interactive database for rice genomics. Plant Cell Physiol 54:e6
McLaren W, Gil L, Hunt SE et al (2016) The ensembl variant effect predictor. Genome Biol 17(1):122
Naithani S, Gupta P, Preece J et al (2020) Plant Reactome: a knowledgebase and resource for comparative pathway analysis. Nucleic Acids Res 48:D1093–D1103
Herrero J, Muffato M, Beal K et al (2016) Ensembl comparative genomics resources. Database 2016:baw053. https://doi.org/10.1093/database/baw053
Consortium TU, The UniProt Consortium (2019) UniProt: a worldwide hub of protein knowledge. Nucleic Acids Res 47:D506–D515
1001 Genomes Consortium (2016) 1,135 Genomes reveal the global pattern of polymorphism in Arabidopsis thaliana. Cell 166:481–491
Atwell S, Huang YS, Vilhjálmsson BJ et al (2010) Genome-wide association study of 107 phenotypes in Arabidopsis thaliana inbred lines. Nature 465:627–631
Fox SE, Preece J, Kimbrel JA et al (2013) Sequencing and de novo transcriptome assembly of Brachypodium sylvaticum (Poaceae). Appl Plant Sci 1:1200011. https://doi.org/10.3732/apps.1200011
International Barley Genome Sequencing Consortium, Mayer KFX, Waugh R et al (2012) A physical, genetic and functional sequence assembly of the barley genome. Nature 491:711–716
Mascher M, Muehlbauer GJ, Rokhsar DS et al (2013) Anchoring and ordering NGS contig assemblies by population sequencing (POPSEQ). Plant J 76:718–727
Ariyadasa R, Mascher M, Nussbaumer T et al (2014) A sequence-ready physical map of barley anchored genetically by two million single-nucleotide polymorphisms. Plant Physiol 164:412–423
Kersey PJ, Allen JE, Allot A et al (2018) Ensembl Genomes 2018: an integrated omics infrastructure for non-vertebrate species. Nucleic Acids Res 46:D802–D808
Bianco L, Cestaro A, Linsmith G et al (2016) Development and validation of the Axiom(®) Apple480K SNP genotyping array. Plant J 86:62–74
Sherry ST (2001) dbSNP: the NCBI database of genetic variation. Nucleic Acids Res 29:308–311
3,000 Rice Genomes Project (2014) The 3,000 rice genomes project. GigaScience 3:7
Duitama J, Silva A, Sanabria Y et al (2015) Whole genome sequencing of elite rice cultivars as a comprehensive information resource for marker assisted selection. PLoS One 10:e0124617
Zhao K, Wright M, Kimball J et al (2010) Genomic diversity and introgression in O. sativa reveal the impact of domestication and breeding on the rice genome. PLoS One 5:e10780
McNally KL, Childs KL, Bohnert R et al (2009) Genomewide SNP variation reveals relationships among landraces and modern varieties of rice. Proc Natl Acad Sci U S A 106:12273–12278
Yamamoto E, Yonemaru J-I, Yamamoto T, Yano M (2012) OGRO: the overview of functionally characterized Genes in Rice online database. Rice 5:26
100 Tomato Genome Sequencing Consortium, Aflitos S, Schijlen E et al (2014) Exploring genetic variation in the tomato (Solanum section Lycopersicon) clade by whole-genome sequencing. Plant J 80:136–148
Morris GP, Ramu P, Deshpande SP et al (2013) Population genomic and genome-wide association studies of agroclimatic traits in sorghum. Proc Natl Acad Sci U S A 110:453–458
Mace ES, Tai S, Gilding EK et al (2013) Whole-genome sequencing reveals untapped genetic potential in Africa’s indigenous cereal crop sorghum. Nat Commun 4:2320
Jiao Y, Burke J, Chopra R et al (2016) A sorghum mutant resource as an efficient platform for gene discovery in grasses. Plant Cell 28:1551–1562
Wilkinson PA, Winfield MO, Barker GLA et al (2012) CerealsDB 2.0: an integrated resource for plant breeders and scientists. BMC Bioinformatics 13:219
Krasileva KV, Vasquez-Gross HA, Howell T et al (2017) Uncovering hidden variation in polyploid wheat. Proc Natl Acad Sci U S A 114:E913–E921
Rimbert H, Darrier B, Navarro J et al (2018) High throughput SNP discovery and genotyping in hexaploid wheat. PLoS One 13:e0186329
Myles S, Chia J-M, Hurwitz B et al (2010) Rapid genomic characterization of the genus vitis. PLoS One 5:e8219
Chia J-M, Song C, Bradbury PJ et al (2012) Maize HapMap2 identifies extant variation from a genome in flux. Nat Genet 44:803–807
Letunic I, Bork P (2019) Interactive Tree Of Life (iTOL) v4: recent updates and new developments. Nucleic Acids Res 47:W256–W259
Federhen S (2012) The NCBI Taxonomy database. Nucleic Acids Res 40:D136–D143
Harris RS (2007) Improved pairwise alignment of genomic DNA. The Pennsylvania State University, Pennsylvania
Petryszak R, Keays M, Tang YA et al (2016) Expression Atlas update--an integrated database of gene and protein expression in humans, animals and plants. Nucleic Acids Res 44:D746–D752
Petryszak R, Fonseca NA, Füllgrabe A et al (2017) The RNASeq-er API—a gateway to systematically updated analysis of public RNA-seq data. Bioinformatics 33:2218–2220
Durinck S, Spellman PT, Birney E, Huber W (2009) Mapping identifiers for the integration of genomic datasets with the R/Bioconductor package biomaRt. Nat Protoc 4:1184–1191
Schmitt T, Messina DN, Schreiber F, Sonnhammer ELL (2011) Letter to the editor: SeqXML and OrthoXML: standards for sequence and orthology information. Brief Bioinform 12:485–488
Igartua E, Contreras-Moreira B, Casas AM (2020) TB1: from domestication gene to tool for many trades. J Exp Bot 71:4621–4624
Walkowiak S, Gao L, Monat C et al (2020) Multiple wheat genomes reveal global variation in modern breeding. Nature 588:277. https://doi.org/10.1038/s41586-020-2961-x
Acknowledgements
We would like to thank Magali Ruffier, Ricardo Ramírez-González, Nikolai Adamski, and Marcela Karey Tello-Ruiz for recipe suggestions and Gramene colleagues Andrew Olson, Sharon Wei, Justin Preece, Pankaj Jaiswal, and Doreen Ware for continuous support and cooperation. We also acknowledge all of the members of the Ensembl team for developing and maintaining the front-end and back-end software and infrastructure that underpins Ensembl Plants.
Funding
The UK Biosciences and Biotechnology Research Council [BB/P016855/1 and Ensembl-4-Breeders workshop support], the National Sciences Foundation [1127112], the ELIXIR implementation studies FONDUE and “Apple as a Model for Genomic Information Exchange,” and the European Molecular Biology Laboratory. Funding for open access charge: UK Biosciences and Biotechnology Research Council [BB/P016855/1].
Conflict of Interest Statement
Paul Flicek is a member of the Scientific Advisory Boards of Fabric Genomics, Inc. and Eagle Genomics, Ltd.
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is licensed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license and indicate if changes were made.
The images or other third party material in this chapter are included in the chapter's Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the chapter's Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
Copyright information
© 2022 The Author(s)
About this protocol
Cite this protocol
Contreras-Moreira, B. et al. (2022). Scripting Analyses of Genomes in Ensembl Plants. In: Edwards, D. (eds) Plant Bioinformatics. Methods in Molecular Biology, vol 2443. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2067-0_2
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2067-0_2
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2066-3
Online ISBN: 978-1-0716-2067-0
eBook Packages: Springer Protocols