Abstract
Enzyme engineering is a tailoring process that allows the modification of naturally occurring enzymes to provide them with improved catalytic efficiency, stability, or specificity. By introducing partial modifications to their sequence and to their structural features, enzyme engineering can transform natural enzymes into more efficient, specific and resistant biocatalysts and render them suitable for virtually countless industrial processes. Current enzyme engineering methods mostly target the active site of the enzyme, where the catalytic reaction takes place. Nonetheless, the tunnel that often connects the surface of an enzyme with its buried active site plays a key role in the activity of the enzyme as it acts as a gatekeeper and regulates the access of the substrate to the catalytic pocket. Hence, there is an increasing interest in targeting the sequence and the structure of substrate entrance tunnels in order to fine-tune enzymatic activity, regulate substrate specificity, or control reaction promiscuity.
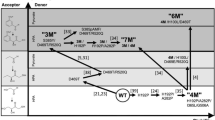
In this chapter, we describe the use of a rational in silico design and screening method to engineer the access tunnel of a fructosyl peptide oxidase with the aim to facilitate access to its catalytic site and to expand its substrate range. Our goal is to engineer this class of enzymes in order to utilize them for the direct detection of glycated proteins in diabetes monitoring devices. The design strategy involves remodeling of the backbone structure of the enzyme , a feature that is not possible with conventional enzyme engineering techniques such as single-point mutagenesis and that is highly unlikely to occur using a directed evolution approach.
The proposed strategy, which results in a significant reduction in cost and time for the experimental production and characterization of candidate enzyme variants, represents a promising approach to the expedited identification of novel and improved enzymes. Rational enzyme design aims to provide in silico strategies for the fast, accurate, and inexpensive development of biocatalysts that can meet the needs of multiple industrial sectors, thus ultimately promoting the use of green chemistry and improving the efficiency of chemical processes.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Pravda L, Berka K, Svobodová Vařeková R et al (2014) Anatomy of enzyme channels. BMC Bioinformatics. https://doi.org/10.1186/s12859-014-0379-x
Prokop Z, Gora A, Brezovsky J et al (2012) Engineering of protein tunnels: the keyhole—lock—key model for catalysis by enzymes with buried active sites. Protein Engineering Handbook, Wiley-VCH, Weinheim, pp. 421–464
Chovancova E, Pavelka A, Benes P et al (2012) CAVER 3.0: a tool for the analysis of transport pathways in dynamic protein structures. PLoS Comput Biol 8. https://doi.org/10.1371/journal.pcbi.1002708
Narayanan C, Bernard DN, Bafna K et al (2018) Ligand-induced variations in structural and dynamical properties within an enzyme superfamily. Front Mol Biosci. https://doi.org/10.3389/fmolb.2018.00054
Kreß N, Halder JM, Rapp LR, Hauer B (2018) Unlocked potential of dynamic elements in protein structures: channels and loops. Curr Opin Chem Biol 47:109–116
Farrugia MA, Wang B, Feig M, Hausinger RP (2015) Mutational and computational evidence that a nickel-transfer tunnel in UreD is used for activation of Klebsiella aerogenes urease. Biochemistry. https://doi.org/10.1021/acs.biochem.5b00942
Lewis-Ballester A, Karkashon S, Batabyal D et al (2018) Inhibition mechanisms of human Indoleamine 2,3 dioxygenase 1. J Am Chem Soc. https://doi.org/10.1021/jacs.8b03691
Kong XD, Yuan S, Li L et al (2014) Engineering of an epoxide hydrolase for efficient bioresolution of bulky pharmaco substrates. Proc Natl Acad Sci U S A. https://doi.org/10.1073/pnas.1404915111
Brezovsky J, Babkova P, Degtjarik O et al (2016) Engineering a de novo transport tunnel. ACS Catal. https://doi.org/10.1021/acscatal.6b02081
Kokkonen P, Sykora J, Prokop Z et al (2018) Molecular gating of an engineered enzyme captured in real time. J Am Chem Soc. https://doi.org/10.1021/jacs.8b09848
Hamre AG, Frøberg EE, Eijsink VGH, Sørlie M (2017) Thermodynamics of tunnel formation upon substrate binding in a processive glycoside hydrolase. Arch Biochem Biophys. https://doi.org/10.1016/j.abb.2017.03.011
Bao L, Li JJ, Jia C et al (2016) Structure-oriented substrate specificity engineering of aldehyde-deformylating oxygenase towards aldehydes carbon chain length. Biotechnol Biofuels. https://doi.org/10.1186/s13068-016-0596-9
Van Beilen JB, Smits THM, Roos FF et al (2005) Identification of an amino acid position that determines the substrate range of integral membrane alkane hydroxylases. J Bacteriol. https://doi.org/10.1128/JB.187.1.85-91.2005
Subramanian K, Góra A, Spruijt R et al (2018) Modulating D-amino acid oxidase (DAAO) substrate specificity through facilitated solvent access. PLoS One. https://doi.org/10.1371/journal.pone.0198990
Brundiek HB, Evitt AS, Kourist R, Bornscheuer UT (2012) Creation of a lipase highly selective for trans fatty acids by protein engineering. Angew Chem Int Ed. https://doi.org/10.1002/anie.201106126
Li G, Yao P, Gong R et al (2017) Simultaneous engineering of an enzyme’s entrance tunnel and active site: the case of monoamine oxidase MAO-N. Chem Sci. https://doi.org/10.1039/c6sc05381e
David B, Irague R, Jouanneau D et al (2017) Internal water dynamics control the transglycosylation/hydrolysis balance in the Agarase (AgaD) of Zobellia galactanivorans. ACS Catal. https://doi.org/10.1021/acscatal.7b00348
Yan X, Wang J, Sun Y et al (2016) Facilitating the evolution of esterase activity from a promiscuous enzyme (Mhg) with catalytic functions of amide hydrolysis and carboxylic acid perhydrolysis by engineering the substrate entrance tunnel. Appl Environ Microbiol. https://doi.org/10.1128/AEM.01817-16
Wu XL, Palfey BA, Mossine VV, Monnier VM (2001) Kinetic studies, mechanism, and substrate specificity of amadoriase I from Aspergillus sp. Biochemistry 40:12886–12895. https://doi.org/10.1021/Bi011244e
Gan W, Gao F, Xing K et al (2015) Structural basis of the substrate specificity of the FPOD/FAOD family revealed by fructosyl peptide oxidase from Eupenicillium terrenum. Acta Crystallogr F Struct Biol Commun 71:381–387. https://doi.org/10.1107/s2053230x15003921
Rigoldi F, Spero L, Dalle Vedove A et al (2016) Molecular dynamics simulations provide insights into the substrate specificity of FAOX family members. Mol BioSyst 12:2622–2633. https://doi.org/10.1039/C6MB00405A
Ferri S, Miyamoto Y, Sakaguchi-Mikami A et al (2013) Engineering fructosyl peptide oxidase to improve activity toward the fructosyl hexapeptide standard for HbA1c measurement. Mol Biotechnol 54:939–943. https://doi.org/10.1007/s12033-012-9644-2
Miura S, Ferri S, Tsugawa W et al (2008) Development of fructosyl amine oxidase specific to fructosyl valine by site-directed mutagenesis. Protein Eng Des Sel 21:233–239. https://doi.org/10.1093/protein/gzm047
Kim S, Miura S, Ferri S et al (2009) Cumulative effect of amino acid substitution for the development of fructosyl valine-specific fructosyl amine oxidase. Enzym Microb Technol 44:52–56. https://doi.org/10.1016/j.enzmictec.2008.09.001
Miura S, Ferri S, Tsugawa W et al (2006) Active site analysis of fructosyl amine oxidase using homology modeling and site-directed mutagenesis. Biotechnol Lett 28:1895–1900. https://doi.org/10.1007/s10529-006-9173-9
Collard F, Zhang J, Nemet I et al (2008) Crystal structure of the deglycating enzyme fructosamine oxidase (amadoriase II). J Biol Chem 283:27007–27016. https://doi.org/10.1074/jbc.M804885200
Hatada M, Tsugawa W, Kamio E et al (2016) Development of a screen-printed carbon electrode based disposable enzyme sensor strip for the measurement of glycated albumin. Biosens Bioelectron 88:1–7. https://doi.org/10.1016/j.bios.2016.08.005
Rigoldi F, Gautieri A, Dalle Vedove A et al (2016) Crystal structure of the deglycating enzyme amadoriase i in its free form and substrate-bound complex. Proteins 84:744–758. https://doi.org/10.1002/prot.25015
Ogawa N, Kimura T, Umehara F et al (2019) Creation of haemoglobin A1c direct oxidase from fructosyl peptide oxidase by combined structure-based site specific mutagenesis and random mutagenesis. Sci Rep 9:942. https://doi.org/10.1038/s41598-018-37806-x
Sievers F, Wilm A, Dineen D et al (2011) Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal omega. Mol Syst Biol 7:539. https://doi.org/10.1038/Msb.2011.75
The PyMOL Molecular Graphics System, Version 2.0 Schrödinger, LLC
Humphrey W, Dalke A, Schulten K (1996) VMD: visual molecular dynamics. J Mol Graph 14:33–38. https://doi.org/10.1016/0263-7855(96)00018-5
Mandell DJ, Coutsias EA, Kortemme T (2009) Sub-angstrom accuracy in protein loop reconstruction by robotics-inspired conformational sampling. Nat Methods 6:551–552. https://doi.org/10.1038/nmeth0809-551
Huang PS, Ban YEA, Richter F et al (2011) Rosettaremodel: a generalized framework for flexible backbone protein design. PLoS One. https://doi.org/10.1371/journal.pone.0024109
Nelson MT, Humphrey W, Gursoy A et al (1996) NAMD: a parallel, object oriented molecular dynamics program. Int J Supercomput Appl High Perform Comput 10:251–268. https://doi.org/10.1177/109434209601000401
Harvey MJ, Giupponi G, De Fabritiis G (2009) ACEMD: accelerating biomolecular dynamics in the microsecond time scale. J Chem Theory Comput 5:1632–1639. https://doi.org/10.1021/Ct9000685
Goldenzweig A, Goldsmith M, Hill SE et al (2016) Automated structure- and sequence-based design of proteins for high bacterial expression and stability. Mol Cell 63:337–346. https://doi.org/10.1016/j.molcel.2016.06.012
Shimasaki T, Yoshida H, Kamitori S, Sode K (2017) X-ray structures of fructosyl peptide oxidases revealing residues responsible for gating oxygen access in the oxidative half reaction. Sci Rep 7:2790. https://doi.org/10.1038/s41598-017-02657-5
Case DA, Cheatham TE, Darden T et al (2005) The Amber biomolecular simulation programs. J Comput Chem 26:1668–1688
Wang JM, Wolf RM, Caldwell JW et al (2004) Development and testing of a general amber force field. J Comput Chem 25:1157–1174. https://doi.org/10.1002/Jcc.20035
Battye TGG, Kontogiannis L, Johnson O et al (2011) iMOSFLM: a new graphical interface for diffraction-image processing with MOSFLM. Acta Crystallogr D Biol Crystallogr. https://doi.org/10.1107/S0907444910048675
McCoy AJ, Grosse-Kunstleve RW, Adams PD et al (2007) Phaser crystallographic software. J Appl Crystallogr. https://doi.org/10.1107/S0021889807021206
Adams PD, Afonine PV, Bunkoczi G et al (2010) PHENIX: a comprehensive python-based system for macromolecular structure solution. Acta Crystallogr D Biol Crystallogr 66:213–221. https://doi.org/10.1107/S0907444909052925
Terwilliger TC, Grosse-Kunstleve RW, Afonine PV et al (2007) Iterative model building, structure refinement and density modification with the PHENIX AutoBuild wizard. Acta Crystallogr D Biol Crystallogr 55:753–762
Murshudov GN, Skubak P, Lebedev AA et al (2011) REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr D Biol Crystallogr 67:355–367. https://doi.org/10.1107/S0907444911001314
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Gautieri, A., Rigoldi, F., Torretta, A., Redaelli, A., Parisini, E. (2022). In Silico Engineering of Enzyme Access Tunnels. In: Magnani, F., Marabelli, C., Paradisi, F. (eds) Enzyme Engineering. Methods in Molecular Biology, vol 2397. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-1826-4_11
Download citation
DOI: https://doi.org/10.1007/978-1-0716-1826-4_11
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-1825-7
Online ISBN: 978-1-0716-1826-4
eBook Packages: Springer Protocols