Abstract
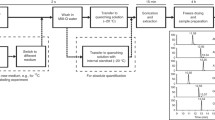
Metabolome analysis in micro physiological models is a challenge due to the low volume of the cell culture medium (CCM). Here, we report a LC-MS-based untargeted metabolomics protocol for the detection of hepatocyte extracellular metabolites from micro-scale samples of CCM. Using a single LC-MS method we have detected 57 metabolites of which 27 showed >2-fold shifts after 72-hour incubation. We demonstrate that micro-scale CCM samples can be used for modelling micro-physiological temporal dynamics in metabolite intensities.
Similar content being viewed by others
References
E. Trefts, M. Gannon, and D. H. Wasserman, Curr. Biol., 2017, 27, R1147.
J. V. Castell, R. Jover, C. P. Martnez-Jimnez, and M. J. Gmez-Lechn, Expert Opin. Drug Metab. Toxicol., 2006, 2, 183.
S. Wilkening, F. Stahl, and A. Bader, Drug Metab. Dispos., 2003, 31, 1035.
S. R. Khetani and S. N. Bhatia, Nat. Biotechnol., 2008, 26, 120.
Y.-C. Toh, T. C. Lim, D. Tai, G. Xiao, D.. van Noort, and H. Yu, Lab Chip, 2009, 9, 2026.
K. Domansky, W. Inman, J. Serdy, A. Dash, M. H. M. Lim, and L. G. Griffith, Lab Chip, 2010, 10, 51.
K. Kamei, M. Yoshioka, S. Terada, Y. Tokunaga, and Y. Chen, Biomed. Microdevices, 2019, 21, 73.
J. G. Camp, K. Sekine, T. Gerber, H. Loeffler-Wirth, H. Binder, M. Gac, S. Kanton, J. Kageyama, G. Damm, D. Seehofer, L. Belicova, M. Bickle, R. Barsacchi, R. Okuda, E. Yoshizawa, M. Kimura, H. Ayabe, H. Taniguchi, T. Takebe, and B. Treutlein, Nature, 2017, 546, 533.
T. Takebe, K. Sekine, M. Kimura, E. Yoshizawa, S. Ayano, M. Koido, S. Funayama, N. Nakanishi, T. Hisai, T. Kobayashi, T. Kasai, R. Kitada, A. Mori, H. Ayabe, Y. Ejiri, N. Amimoto, Y. Yamazaki, S. Ogawa, M. Ishikawa, Y. Kiyota, Y. Sato, K. Nozawa, S. Okamoto, Y. Ueno, and H. Taniguchi, Cell Rep., 2017, 21, 2661.
S. Halldorsson, E. Lucumi, R. Gómez-Sjöberg, and R. M. T. Fleming, Biosens. Bioelectron., 2015, 63, 218.
S. Naz, D. C. Moreira dos Santos, A. García, and C. Barbas, Bioanalysis, 2014, 6, 1657.
S. Takashina, Y. Igarashi, M. takahashi, Y. Kondo, and K. Inoue, Anal. Sci., 2020, 36, 1427.
S. Dong, Z. Yan, and H. Yang, Anal. Sci., 2016, 32, 1333.
K. D. Duncan, J. Fyrestam, and I. Lanekoff, Analyst, 2019, 144, 782.
E. Daskalaki, N. J. Pillon, A. Krook, C. E. Wheelock, and A. Checa, Anal. Chim. Acta, 2018, 1037, 338.
S. Naz, H. Gallart-Ayala, S. N. Reinke, C. Mathon, R. Blankley, R. Chaleckis, and C. E. Wheelock, Anal. Chem., 2017, 89, 7933.
R. Chaleckis, S. Naz, I. Meister, and C. E. Wheelock, “Methods in Molecular Biology (Clifton, N.J.)”, 2018, Vol. 1730, United States, 45.
I. Tada, H. Tsugawa, I. Meister, P. Zhang, R. Shu, R. Katsumi, C. E. Wheelock, M. Arita, and R. Chaleckis, Metabolites, 2019, 9, 251.
H. Tsugawa, T. Cajka, T. Kind, Y. Ma, B. Higgins, K. Ikeda, M. Kanazawa, J. VanderGheynst, O. Fiehn, and M. Arita, Nat. Methods, 2015, 12, 523.
I. Tada, R. Chaleckis, H. Tsugawa, I. Meister, P. Zhang, N. Lazarinis, B. Dahlén, C. E. Wheelock, and M. Arita, Anal. Chem., 2020, 92, 11310.
K. Haug, K. Cochrane, V. C. Nainala, M. Williams, J. Chang, K. V. Jayaseelan, and C. O’Donovan, Nucleic Acids Res., 2020, 48, D440.
J. Demšar, T. Curk, A. Erjavec, Č. Gorup, T. Hočevar, M. Milutinovič, M. Možina, M. Polajnar, M. Toplak, A. Starič, M. Štajdohar, L. Umek, L. Žagar, J. Žbontar, M. Žitnik, and B. Zupan, J. Mach. Learn. Res., 2013, 35, 2349.
J. Chong, O. Soufan, C. Li, I. Caraus, S. Li, G. Bourque, D. S. Wishart, and J. Xia, Nucleic Acids Res., 2018, 46, W486.
L. W. Sumner, A. Amberg, D. Barrett, M. H. Beale, R. Beger, C. A. Daykin, T. W. M. Fan, O. Fiehn, R. Goodacre, J. L. Griffin, T. Hankemeier, N. Hardy, J. Harnly, R. Higashi, J. Kopka, A. N. Lane, J. C. Lindon, P. Marriott, A. W. Nicholls, M. D. Reily, J. J. Thaden, and M. R. Viant, Metabolomics, 2007, 3, 211.
R. Boon, M. Kumar, T. Tricot, I. Elia, L. Ordovas, F. Jacobs, J. One, J. De Smedt, G. Eelen, M. Bird, P. Roelandt, G. Doglioni, K. Vriens, M. Rossi, M. A. Vazquez, T. Vanwelden, F. Chesnais, A. El Taghdouini, M. Najimi, E. Sokal, D. Cassiman, J. Snoeys, M. Monshouwer, W. S. Hu, C. Lange, P. Carmeliet, S. M. Fendt, and C. M. Verfaillie, Nat. Commun., 2020, 11, 1393.
Acknowledgements
This work was supported by JSPS KAKENHI Grant Number 17H02083, 20K20168 and 19K17662. Kyoto University GAP fund program (207010). We acknowledge the support from the Gunma University Initiative for Advanced Research (GIAR). I. M. was supported by Japan Society for the Promotion of Science (JSPS) postdoctoral fellowship (P17774). Authors thank Dr. Satoshi Imamura for assisting in HepG2 cells culture.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Abdalkader, R., Chaleckis, R., Meister, I. et al. Untargeted LC-MS Metabolomics for the Analysis of Micro-scaled Extracellular Metabolites from Hepatocytes. ANAL. SCI. 37, 1049–1052 (2021). https://doi.org/10.2116/analsci.20N032
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.2116/analsci.20N032