Abstract
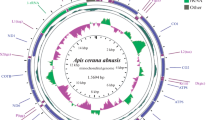
Apis cerana ussuriensis Ilyasov et al., 2019 is the northernmost subspecies of the Asian honey bee A. cerana Fabricius, 1793, common in the forests of Primorsky krai and Khabarovsk krai as far as 47°54′ N. Genetic studies of this subspecies are of great interest for science and apiculture, since all its adaptive traits were formed under the influence of the natural environment without human interference. We sequenced and annotated the complete mitochondrial DNA (mtDNA) sequences of bees of subspecies Apis cerana ussuriensis Ilyasov et al., 2019 (GenBank accession number AP018450) from Primorsky krai and Apis cerana koreana Ilyasov et al., 2019 (AP018431) from South Korea, as well as six exons of the nuclear DNA (nDNA) vitellogenin VG E2–E7 gene of bee subspecies A. c. ussuriensis, A. c. koreana, A. c. japonica Radoszkowski, 1887, A. c. cerana, and A. c. indica Fabricius, 1798. Cluster analysis of the mtDNA and the nDNA VG gene sequences showed the division of bees into two groups, the southern subspecies A. c. indica and the northern subspecies A. c. ussuriensis, A. c. koreana, A. c. japonica, and A. c. cerana. On the basis of the genetic divergence, we showed that subspecies A. c. ussuriensis was genetically closer to subspecies A. c. japonica, A. c. koreana, and A. c. cerana than to subspecies A. c. indica. Values of genetic divergence (0.80–8.00%) and Jukes–Cantor genetic distance (0.005–0.100) for mtDNA and nDNA VG gene between subspecies A. c. ussuriensis, A. c. koreana, A. c. japonica, A. c. cerana, and A. c. indica are within the range of intraspecific differences between insect subspecies. The estimated time of the emergence of the A. cerana subspecies is from two to one million years ago.



Similar content being viewed by others
REFERENCES
Ruttner, F., Breeding Techniques and Selection for Breeding of the Honeybee, Derby, UK: British Isles Bee Breeders Association, 1988.
Proshchalykin, M.Yu., Novomodnyi, E.V., Bezborodov, V.G., and Koshkin, E.S., The first modern findings of a wax bee Apis cerana Fabricius, 1793 (Hymenoptera, Apidae) in Khabarovsk Krai, Evras. Entomol. Zh., 2014, vol. 13, no. 3, pp. 295—298.
Kuznetsov, V.N., Kitaiskaya voskovaya pchela Apis cerana cerana F. (Hymenoptera, Apidae) na Dal’nem Vostoke Rossii (Chinese Wax Bee, Apis cerana cerana F. (Hymenoptera, Apidae) in the Russian Far East), Moscow: KMK, 2005.
Behura, S.K., Analysis of nuclear copies of mitochondrial sequences in honeybee (Apis mellifera) genome, Mol. Biol. Evol., 2007, vol. 24, no. 7, pp. 1492—1505. https://doi.org/10.1093/molbev/msm068
Choi, Y.S., Lee, M.Y., Hong, I.P., et al., Occurrence of sacbrood virus in Korean apiaries from Apis cerana (Hymenoptera: Apidae), J. Apic., 2010, vol. 25, no. 3, pp. 187—191.
Koetz, A.H., Ecology, behaviour and control of Apis cerana with a focus on relevance to the Australian incursion, Insects, 2013, vol. 4, no. 4, pp. 558—592. https://doi.org/10.3390/insects4040558
Vung, N., Lee, M.-L., Lee, M.-Y., et al., Breeding and selection for resistance to sacbrood virus for Apis cerana, J. Apic., 2017, vol. 32, pp. 345—352. https://doi.org/10.17519/apiculture.2017.11.32.4.345
Pesenko, Yu.A., Lelei, A.S., Radchenko, V.G., and Filatkin, G.N., Chinese wax bee Apis cerana cerana F. (Hymenoptera, Apidae) in the Soviet Far East, Entomol. Obozr., 1989, vol. 68, no. 3, pp. 527—548.
Zhuang, D., New subspecies of Apis cerana (in Chinese), Southwest China J. Agric. Sci., 1989, vol. 2, pp. 61—65.
Zhen-Ming, J., Yang, G., Huang, S., et al., The advancement of beekeeping science and technology in China, Honeybees in Mountain Agriculture, Verma, L.R., Ed., New Delhi: Oxford and IBH Publ., 1992.
Diniz-Filho, J.A.F., Malapsina, O., and Pignata, M.I.B., Geographic variation in Apis cerana indica F.: a spatial autocorrelation analysis of morphometric patterns, J. Apic. Res., 1993, vol. 32, pp. 65—72. https://doi.org/10.1080/00218839.1993.11101289
Engel, M.S., The taxonomy of recent and fossil honey bees (Hymenoptera, Apidae, Apis), J. Hymenoptera Res., 1999, vol. 8, no. 2, pp. 165—196. https://doi.org/10.1007/978-1-4614-4960-718
Sugawara, M., Feral colonies of Japanese honey bees, Apis cerana japonica and their life history: 2. Natural nests and swarming, Mitsubachi Kagaku (Honeybee Sci.), 2000, vol. 21, no. 1, pp. 35—39.
Hepburn, H.R., Smith, D.R., Radloff, S.E., and Otis, G.W., Infraspecific categories of Apis cerana: morphometric, allozymal and mtDNA diversity, Apidologie, 2001, vol. 32, no. 1, pp. 3—23. https://doi.org/10.1051/apido:2001108
Takahashi, J. and Yoshida, T., The origin of Japanese honey bee Apis cerana japonica inferred from mitochondrial DNA, Mitsubachi Kagaku (Honeybee Sci.), 2003, vol. 24, no. 2, pp. 71—76.
Radloff, S.E., Hepburn, C., Hepburn, R.H., et al., Population structure and classification of Apis cerana, Apidologie, 2010, vol. 41, no. 6, pp. 589—601. https://doi.org/10.1051/apido/2010008
Takahashi, J., Wakamiya, T., Kiyoshi, T., et al., The complete mitochondrial genome of the Japanese honeybee, Apis cerana japonica (Insecta: Hymenoptera: Apidae), Mitochondrial DNA, Part B, 2016, vol. 1, no. 1, pp. 156—157. https://doi.org/10.1080/23802359.2016.1144108
Ilyasov, R.A., Park, J., Takahashi, J., and Kwon, H.W., Phylogenetic uniqueness of honeybee Apis cerana from the Korean peninsula inferred from the mitochondrial, nuclear, and morphological data, J. Apic. Sci., 2018, vol. 62, no. 2, pp. 189—214. https://doi.org/10.2478/Jas-2018-0018
Ilyasov, R.A., Han, G.Y., Lee, M.L., et al., Phylogenetic relationships of Russian Far-East Apis cerana with other north Asian populations, J. Apic. Sci., 2019, vol. 63, no. 2, pp. 297—322. https://doi.org/10.2478/JAS-2019-0024
Cornuet, J.-M., Garnery, L., and Solignac, M., Putative origin and function of the intergenic region between COI and COII of Apis mellifera L. mitochondrial DNA, Genetics, 1991, vol. 128, no. 2, pp. 393—403.
Garnery, L., Cornuet, J.-M., and Solignac, M., Evolutionary history of the honey bee Apis mellifera inferred from mitochondrial DNA analysis, Mol. Ecol., 1992, vol. 1, no. 3, pp. 145—154. https://doi.org/10.1111/j.1365-294x.1992.tb00170.x
Garnery, L., Mosshine, E.H., Oldroyd, B.P., and Cornuet, J.-M., Mitochondrial DNA variation in Moroccan and Spanish honey bee populations, Mol. Ecol., 1995, vol. 4, pp. 465—472. https://doi.org/10.1111/j.1365-294X.1995.tb00240.x
Arias, M.C. and Sheppard, W.S., Molecular phylogenetics of honey bee subspecies (Apis mellifera L.) inferred from mitochondrial DNA sequence, Mol. Phylogenet. Evol., 1996, vol. 5, no. 3, pp. 557—566. https://doi.org/10.1006/mpev.1996.0050
Songrarn, O., Sittipraneed, S., and Klinbunga, S., Mitochondrial DNA diversity and genetic differentiation of the honeybee (Apis cerana) in Thailand, Biochem. Genet., 2006, vol. 44, nos. 5—6, pp. 256—269. https://doi.org/10.1007/s10528-006-9030-5
Tan, H.W., Liu, G.H., Dong, X., et al., The complete mitochondrial genome of the Asiatic cavity-nesting honeybee Apis cerana (Hymenoptera: Apidae), PLoS One, 2011, vol. 6, no. 8. e23008. https://doi.org/10.1371/journal.pone.0023008
Kent, C.F., Issa, A., Bunting, A.C., and Zayed, A., Adaptive evolution of a key gene affecting queen and worker traits in the honey bee, Apis mellifera, Mol. Ecol., 2011, vol. 20, no. 24, pp. 5226—5235. https://doi.org/10.1111/j.1365-294X.2011.05299.x
Bernt, M., Donath, A., Jühling, F., et al., MITOS: improved de novo metazoan mitochondrial genome annotation, Mol. Phylogenet. Evol., 2013, vol. 69, no. 2, pp. 313—319. https://doi.org/10.1016/j.ympev.2012.08.023
Lowe, T.M. and Eddy, S.R., tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence, Nucleic Acids. Res., 1997, vol. 25, no. 5, pp. 955—964. https://doi.org/10.1093/nar/25.5.0955
Sanger, F., Nicklen, S., and Coulson, A.R., DNA sequencing with chain-terminating inhibitors, Proc. Natl. Acad. Sci. U.S.A., 1977, vol. 74, no. 12, pp. 5463—5467. https://doi.org/10.1073/pnas.74.12.5463
Jukes, T.H. and Cantor, C.R., Evolution of protein molecules, in Mammalian Protein Metabolism, Munro, H.N., Ed., New York: Acad. Press, 1969, pp. 21—132.
Tamura, K., Battistuzzi, F.U., Billing-Ross, P., et al., Estimating divergence times in large molecular phylogenies, Proc. Natl. Acad. Sci. U.S.A., 2012, vol. 109, no. 47, pp. 19333—19338. https://doi.org/10.1073/pnas.1213199109
Nei, M. and Kumar, S., Molecular Evolution and Phylogenetics, New York: Oxford Univ. Press, 2000.
Kumar, S., Stecher, G., and Tamura, K., MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets, Mol. Biol. Evol., 2016, vol. 33, no. 7, pp. 1870—1874. https://doi.org/10.1093/molbev/msw054
Saitou, N. and Nei, M., The neighbor-joining method: a new method for reconstructing phylogenetic trees, Mol. Biol. Evol., 1987, vol. 4, no. 4, pp. 406—425. https://doi.org/10.1093/oxfordjournals.molbev.a040454
Tamura, K. and Nei, M., Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees, Mol. Biol. Evol., 1993, vol. 10, no. 3, pp. 512—526. https://doi.org/10.1093/oxfordjournals.molbev.a040023
Crozier, R.H. and Crozier, Y.C., The mitochondrial genome of the honeybee Apis mellifera: complete sequence and genome organization, Genetics, 1993, vol. 133, no. 1, pp. 97—117. https://doi.org/10.1111/j.1365-2583.1993.tb00131.x
Okuyama, H., Tingek, S., and Takahashi, J., The complete mitochondrial genome of the cavity-nesting honeybee, Apis cerana (Insecta: Hymenoptera: Apidae) from Borneo, Mitochondrial DNA, Part B, 2017, vol. 2, no. 2, pp. 475—476. https://doi.org/10.1080/23802359.2017.1361344
Kent, C.F., Minaei, S., Harpur, B.A., and Zayed, A., Recombination is associated with the evolution of genome structure and worker behavior in honey bees, Proc. Natl. Acad. Sci. U.S.A., 2012, vol. 109, no. 44, pp. 18012—18017. https://doi.org/10.1073/pnas.1208094109
Lee, J.Y., Wang, A.R., Choi, Y.S., et al., Mitochondrial DNA variations in Korean Apis cerana (Hymenoptera: Apidae) and development of another potential marker, Apidologie, 2016, vol. 47, no. 1, pp. 123—134. https://doi.org/10.1007/s13592-015-0381-y
Tan, Y.D., Wan, C.L., Zhu, Y.F., et al., An amplified fragment length polymorphism map of the silkworm, Genetics, 2001, vol. 157, no. 3, pp. 1277—1284.
Smith, C.R., Smith, C.D., Robertson, H.M., et al., Draft genome of the red harvester ant Pogonomyrmex barbatus, Proc. Natl. Acad. Sci. U.S.A., 2011, vol. 108, no. 14, pp. 5667—5672. https://doi.org/10.1073/pnas.1007901108
Tian, D., Wang, Q., Zhang, P., et al., Single-nucleotide mutation rate increases close to insertions/deletions in eukaryotes, Nature, 2008, vol. 455, no. 7209, pp. 105—108. https://doi.org/10.1038/nature07175
McDonald, M.J., Wang, W.C., Huang, H.D., and Leu, J.Y., Clusters of nucleotide substitutions and insertion/deletion mutations are associated with repeat sequences, PLoS Biol., 2011, vol. 9, no. 6. e1000622. https://doi.org/10.1371/journal.pbio.1000622
Koren, A., Polak, P., Nemesh, J., et al., Differential relationship of DNA replication timing to different forms of human mutation and variation, Am. J. Hum. Genet., 2012, vol. 91, no. 6, pp. 1033—1040. https://doi.org/10.1016/j.ajhg.2012.10.018
Seplyarskiy, V.B., Kharchenko, P., Kondrashov, A.S., and Bazykin, G.A., Heterogeneity of the transition/transversion ratio in Drosophila and Hominidae genomes, Mol. Biol. Evol., 2012, vol. 29, no. 8, pp. 1943—1955. https://doi.org/10.1093/molbev/mss071
Han, T., Lee, W., Lee, S., et al., Reassessment of species diversity of the subfamily Denticollinae (Coleoptera: Elateridae) through DNA barcoding, PLoS One, 2016, vol. 11, no. 2. e0148602. https://doi.org/10.1371/journal.pone.0148602
Eimanifar, A., Kimball, R.T., Braun, E.L., et al., The complete mitochondrial genome of the Egyptian honey bee, Apis mellifera lamarckii (Insecta: Hymenoptera: Apidae), Mitochondrial DNA, Part B, 2017, vol. 2, no. 1, pp. 270—272. https://doi.org/10.1080/23802359.2017.1325343
DeSalle, R., Freedman, T., Prager, E.M., and Wilson, A.C., Tempo and mode of sequence evolution in mitochondrial DNA of Hawaiian Drosophila, J. Mol. Evol., 1987, vol. 26, nos. 1—2, pp. 157—164. https://doi.org/10.1007/BF02111289
Johns, G.C. and Avise, J.C., A comparative summary of genetic distances in the vertebrates from the mitochondrial cytochrome b gene, Mol. Biol. Evol., 1998, vol. 15, pp. 1481—1490. https://doi.org/10.1093/oxfordjournals.molbev.a025875
ACKNOWLEDGMENTS
We are grateful to Dr. Hisashi Okuyama for the assistance with sequencing as well as the e-ASIA JRP (The East Asia Science and Innovation Area Joint Research Program) Foundation for support in the implementation of the project (https://www.the-easia.org).
Funding
The present study was supported by the government assignment (registration no. AAAA-A21-121011990120-7) (IR), by the Russian Foundation for Basic Research (no. 19-54-70002 e-Asia_t) (NA)), and by the postdoctoral research programs of Incheon National University 2019 (IR).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest. The authors declare that they have no conflict of interest.
Statement on the welfare of animals. All applicable international, national, and/or institutional guidelines for the care and use of animals were followed.
Additional information
Translated by A. Lisenkova
Rights and permissions
About this article
Cite this article
Ilyasov, R.A., Han, G.Y., Lee, M.L. et al. Genetic Properties and Evolution of Asian Honey Bee Apis cerana ussuriensis from Primorsky Krai, Russia. Russ J Genet 57, 568–581 (2021). https://doi.org/10.1134/S1022795421050033
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S1022795421050033