Abstract
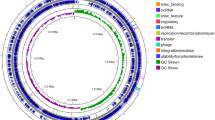
Whole-genome sequencing of bacteria Bacillus velezensis BIM B-439D has revealed that its genome is represented by a single circular chromosome 3 978 954 bp in size and contains 3969 predicted genes. Chromosome has 10 conserved loci that determine the production of antimicrobial compounds of varying chemical nature, namely lipopeptides (surfactin, bacillomycin, fengycin), polyketides (bacillaene, difficidin/oxydifficidin, macrolactin), bacillibactin siderophore, dipeptide bacilysin, a protein/polyketide of uncertain composition, and amylocyclicin bacteriocin. Mutants with impaired surfactin synthesis have been obtained by direct mutagenesis; they were characterized by an increased production of bacillomycin and fengycin. The obtained mutants demonstrated high antimicrobial activity towards a number of bacterial and fungal pathogens (Penicillium expansum, Alternaria tenuis, Botrytis cinerea,Bipolaris sorokiniana), but they inhibited the growth of the Fusarium genus fungi to a lesser degree.




Similar content being viewed by others
REFERENCES
Borriss, R., in Bacteria in Agrobiology: Plant Growth Responses, Maheshwari, D.K., Ed., Heidelberg: Springer, 2011, pp. 41–76.
Chen, X.H., Koumoutsi, A., Scholz, R., and Borriss, R., J. Mol. Microbiol. Biotechnol., 2009, vol. 16, nos. 1–2, pp. 14–24. https://doi.org/10.1159/000142891
Bullock, W.O., Fernandez, J.M., and Short, J.M., BioTechniques, 1987, vol. 5, no. 3, pp. 376–379.
Palmer, B.R. and Marinus, M.G., Gene, 1994, vol. 143, no. 1, pp. 1–12. https://doi.org/10.1016/0378-1119(94)90597-5
Vieira, J. and Messing, J., Gene, 1982, vol. 19, no. 3, pp. 259–268.
Cbambers, S.P., Prior, S.E., Barstow, D.A., and Minton, N.P., Gene, 1988, vol. 68, no. 1, pp. 139–149.
Miller, J., Experiments in Molecular Genetics, New York: Cold Spring Harbor, 1972.
Dudka I. A., Vasser S. P., Ellanskaya I. A., Koval’ E. 3., Gorbik L. T., Nikol’skaya, E.A., et al., Metody eksperimental’noi mikologii: spravochnik (Experimental Mycology Methods: A Handbook), Bilai, V.I., Ed., Kiev: Naukova Dumka, 1982.
Maniatis, T., Fritsch, E. F., and Sambrook, J. Molecular Cloning, Cold Spring Harbor, New York: Cold Spring Harbor Lab. Press, 1982.
Chen, X.H., Liesegang, H., Hitzeroth, G., Franke, P., Vater, J., and Borriss, R., J. Bacteriol., 2004, vol. 186, no. 4, pp. P. 1084–1096.
Richter, M., Rosselló-Móra, R., Oliver Glöckner, F., and Peplies, J., Bioinformatics, 2016, vol. 32, no. 6, pp. 929–931.
Yokota, K., Yatsuda, M., Miwa, E., and Higuchi, K.J., J. ISSAA, 2012, vol. 18, no. 1, pp. 70–75.
Yang, H., Li, X., Li, X., Yu, H., and Shen, Z., Anal. Bioanal. Chem., 2015, vol. 407, no. 9, pp. 2529–2542. https://doi.org/10.1007/s00216-015-8486-8
Segi, I., Metody pochvennoi mikrobiologii (Methods of Soil Microbiology), Moscow: Kolos, 1983.
Darling, A.E., Mau, B., and Perna, N.T., PLoS One, 2010, vol. 5, no. 6. https://doi.org/10.1371/journal.pone.0011147
Fan, B., Blom, J., Klenk, Hans-P., and Borriss, R., Front. Microbiol., 2017, vol. 8, no. 22, pp. 1–15.
Niazi, A., Manzoor, Sh., Asari, S., Bejai, S., Meijer, J., and Bongcam-Rudloff, E., PLoS One, 2014, vol. 9, no. 8. e104651. https://doi.org/10.1371/journal.pone.0104651
Rückert, Ch., Blom, J., Chen, X.H., Revac, O., and Borriss, R.J., Biotechnology, 2011, no. 155, pp. 78– 85.
Chen, X.H., Scholz, R., Eisenreich, A., Schneider, K., Heinemeyer, I., et al., Nat. Biotechnol., 2007, vol. 25, pp. 1007–1014. https://doi.org/10.1038/nbt1325
Borriss, R., Chen, X.H., Ruckert, Ch., Blom, J., Becker, A., Baumgarth, B., Fan, B., Pukall, R., Schumann, P., Spröer, C., Junge, H., Vater, J., Pühler, A., and Klenk, Hans-P., Int. J. Syst. Evol. Microbiol., 2011, pp. 1786–1801.
Dunlap, C.A., Kim, S.-J., Kwon, S.-W., and Rooney, A.P., Int. J. Syst. Evol. Microbiol., 2016, vol. 66, pp. 1212–1217.
Klappenbach, J.A., Dunbar, J.M., and Schmidt, T.M., Appl. Environ. Microbiol., 2000, vol. 66, pp. 1328–1333.
Luo, C., Liu, X., Zhou, H., Wang, Z., and Chen, X.H., Appl. Environ. Microbiol., 2015, vol. 81, no. 1, pp. 422–431.
Reva, O.N. and Tummler, B., BMC Bioinf., 2005, vol. 6, no. 251, pp. 1–12.
Scholz, R., Vater, J., Budiharjo, A., Wang, Z., He, Y., Dietel, K., Schwecke, T., Herfort, S., Lasch, P., and Borriss, R., J. Bacteriol., 2014, vol. 196, no. 10, pp. 1842–1852.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors declare that they have no conflict of interest. This article does not contain any studies involving animals or human participants performed by any of the authors.
Additional information
Translated by E. Martynova
Rights and permissions
About this article
Cite this article
Berezhnaya, A.V., Evdokimova, O.V., Valentovich, L.N. et al. Molecular Genetic and Functional Analysis of the Genome of Bacteria Bacillus velezensis BIM B-439D. Appl Biochem Microbiol 55, 386–396 (2019). https://doi.org/10.1134/S0003683819040033
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0003683819040033