Abstract
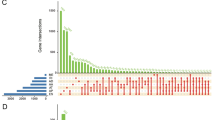
Alternative splicing (AS) in the tumor biological process has provided a novel perspective on carcinogenesis. However, the clinical significance of individual AS patterns of adrenocortical carcinoma (ACC) has been underestimated, and in-depth investigations are lacking. We selected 76 ACC samples from the Cancer Genome Atlas (TCGA) SpliceSeq and SpliceAid2 databases, and 39 ACC samples from Fudan University Shanghai Cancer Center (FUSCC). Prognosis-related AS events (PASEs) and survival analysis were evaluated based on prediction models constructed by machine-learning algorithm. In total, 23,984 AS events and 3,614 PASEs were detected in the patients with ACC. The predicted risk score of each patient suggested that eight PASEs groups were significantly correlated with the clinical outcomes of these patients (p < 0.001). Prognostic models produced AUC values of 0.907 in all PASEs’ groups. Eight splicing factors (SFs), including BAG2, CXorf56, DExD-Box Helicase 21 (DDX21), HSPB1, MBNL3, MSI1, RBMXL2, and SEC31B, were identified in regulatory networks of ACC. DDX21 was identified and validated as a novel clinical promoter and therapeutic target in 115 patients with ACC from TCGA and FUSCC cohorts. In conclusion, the strict standards used in this study ensured the systematic discovery of profiles of AS events using genome-wide cohorts. Our findings contribute to a comprehensive understanding of the landscape and underlying mechanism of AS, providing valuable insights into the potential usages of DDX21 for predicting prognosis for patients with ACC.







Similar content being viewed by others
Availability of data and materials
The datasets analyzed in this study were obtained from the corresponding author upon reasonable request or open-access online databases.
Code availability
The R code is publicly available, and it can be found by name on the GitHub site listed in the Data and Material Availability (https://github.com/) section.
Abbreviations
- ACC:
-
Adrenocortical carcinoma
- AS:
-
Alternative splicing
- AS:
-
Alternative splicing
- AA:
-
Alternate acceptor site
- AD:
-
Alternate donor site
- AP:
-
Alternate promotor
- AT:
-
Alternate terminator
- ES:
-
Exon skip
- ME:
-
Mutually exclusive exons
- RI:
-
Retained intron
- GO:
-
Gene ontology
- BP:
-
Biological process
- CC:
-
Cellular components
- MF:
-
Molecular functions
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- PPI:
-
Protein–protein interaction
- TCGA:
-
The Cancer Genome Atlas
- SFs:
-
Splicing factors
- OS:
-
Overall survival
- DFS:
-
Disease-free survival
- CI:
-
Confidence interval
- PSI:
-
Percent-spliced-in
- PASEs:
-
Prognosis-related AS events
- GSEA:
-
Gene set enrichment analysis
- IHC:
-
Immunohistochemical
References
Bates SE (2020) Epigenetic therapies for cancer. N Engl J Med 383(7):650–663. https://doi.org/10.1056/NEJMra1805035
Cao J, Wu N, Han Y, Hou Q, Zhao Y, Pan Y, Xie X, Chen F (2018) DDX21 promotes gastric cancer proliferation by regulating cell cycle. Biochem Biophys Res Commun 505(4):1189–1194. https://doi.org/10.1016/j.bbrc.2018.10.060
Chen T, Zheng W, Chen J, Lin S, Zou Z, Li X, Tan Z (2019) Systematic analysis of survival-associated alternative splicing signatures in clear cell renal cell carcinoma. J Cell Biochem. https://doi.org/10.1002/jcb.29590
Climente-González H, Porta-Pardo E, Godzik A, Eyras E (2017) The functional impact of alternative splicing in cancer. Cell Rep 20(9):2215–2226. https://doi.org/10.1016/j.celrep.2017.08.012
Consortium SEQC/MAQC-III (2014) A comprehensive assessment of RNA-seq accuracy reproducibility and information content by the Sequencing Quality Control Consortium. Nat Biotechnol 32(9):903–914. https://doi.org/10.1038/nbt.2957
Feng H, Qin Z, Zhang X (2013) Opportunities and methods for studying alternative splicing in cancer with RNA-Seq. Cancer Lett 340(2):179–191. https://doi.org/10.1016/j.canlet.2012.11.010
Giulietti M, Piva F, D’Antonio M, D’Onorio De Meo P, Paoletti D, Castrignanò T, D’Erchia AM, Picardi E, Zambelli F, Principato G, Pavesi G, Pesole G (2013) SpliceAid-F: a database of human splicing factors and their RNA-binding sites. Nucleic Acids Res 41(Database issue):D125-131. https://doi.org/10.1093/nar/gks997
Jasim S, Habra MA (2019) Management of adrenocortical carcinoma. Curr Oncol Rep 21(3):20. https://doi.org/10.1007/s11912-019-0773-7
Jonasch E, Walker CL, Rathmell WK (2021) Clear cell renal cell carcinoma ontogeny and mechanisms of lethality. Nat Rev Nephrol 17(4):245–261. https://doi.org/10.1038/s41581-020-00359-2
Kim DS, Camacho CV, Nagari A, Malladi VS, Challa S, Kraus WL (2019) Activation of PARP-1 by snoRNAs controls ribosome biogenesis and cell growth via the RNA helicase DDX21. Mol Cell 75(6):1270-1285 e1214. https://doi.org/10.1016/j.molcel.2019.06.020
Kostiainen I, Hakaste L, Kejo P, Parviainen H, Laine T, Löyttyniemi E, Pennanen M, Arola J, Haglund C, Heiskanen I, Schalin-Jäntti C (2019) Adrenocortical carcinoma: presentation and outcome of a contemporary patient series. Endocrine 65(1):166–174. https://doi.org/10.1007/s12020-019-01918-9
Lee Y, Rio DC (2015) Mechanisms and regulation of alternative Pre-mRNA splicing. Annu Rev Biochem 84:291–323. https://doi.org/10.1146/annurev-biochem-060614-034316
Li ZX, Zheng ZQ, Wei ZH, Zhang LL, Li F, Lin L, Liu RQ, Huang XD, Lv JW, Chen FP, He XJ, Guan JL, Kou J, Ma J, Zhou GQ, Sun Y (2019) Comprehensive characterization of the alternative splicing landscape in head and neck squamous cell carcinoma reveals novel events associated with tumorigenesis and the immune microenvironment. Theranostics 9(25):7648–7665. https://doi.org/10.7150/thno.36585
Libe R (2015) Adrenocortical carcinoma (ACC): diagnosis prognosis and treatment. Front Cell Dev Biol 3:45. https://doi.org/10.3389/fcell.2015.00045
Liu C, Guo T, Xu G, Sakai A, Ren S, Fukusumi T, Ando M, Sadat S, Saito Y, Khan Z, Fisch KM, Califano J (2018) Characterization of alternative splicing events in HPV-negative head and neck squamous cell carcinoma identifies an oncogenic DOCK5 variant. Clin Cancer Res 24(20):5123–5132
Maniatis T, Tasic B (2002) Alternative pre-mRNA splicing and proteome expansion in metazoans. Nature 418(6894):236–243. https://doi.org/10.1158/1078-0432.CCR-18-0752
Meng T, Huang R, Zeng Z, Huang Z, Yin H, Jiao C, Yan P, Hu P, Zhu X, Li Z, Song D, Zhang J, Cheng L (2019) Identification of prognostic and metastatic alternative splicing signatures in kidney renal clear cell carcinoma. Front Bioeng Biotechnol 7:270. https://doi.org/10.3389/fbioe.2019.00270
Nilsen TW, Graveley BR (2010) Expansion of the eukaryotic proteome by alternative splicing. Nature 463(7280):457–463. https://doi.org/10.1038/nature08909
Rahman MA, Krainer AR, Abdel-Wahab O (2020) SnapShot: splicing alterations in cancer. Cell 180(1):208-208 e201. https://doi.org/10.1016/j.cell.2019.12.011
Ryan MC, Cleland J, Kim R, Wong WC, Weinstein JN (2012) SpliceSeq: a resource for analysis and visualization of RNA-Seq data on alternative splicing and its functional impacts. Bioinformatics 28(18):2385–2387. https://doi.org/10.1093/bioinformatics/bts452
Ryan M, Wong WC, Brown R, Akbani R, Su X, Broom B, Melott J, Weinstein J (2016) TCGASpliceSeq a compendium of alternative mRNA splicing in cancer. Nucleic Acids Res 44(D1):D1018-1022. https://doi.org/10.1093/nar/gkv1288
Schteingart DE, Doherty GM, Gauger PG, Giordano TJ, Hammer GD, Korobkin M, Worden FP (2005) Management of patients with adrenal cancer: recommendations of an international consensus conference. Endocr Relat Cancer 12(3):667–680. https://doi.org/10.1677/erc.1.01029
Sun W, Duan T, Ye P, Chen K, Zhang G, Lai M, Zhang H (2018) TSVdb: a web-tool for TCGA splicing variants analysis. BMC Genom 19(1):405. https://doi.org/10.1186/s12864-018-4775-x
Tierney JF, Chivukula SV, Poirier J, Pappas SG, Schadde E, Hertl M, Kebebew E, Keutgen X (2019) National treatment practice for adrenocortical carcinoma: have they changed and have we made any progress? J Clin Endocrinol Metab 104(12):5948–5956. https://doi.org/10.1210/jc.2019-00915
Tomczak K, Czerwińska P, Wiznerowicz M (2015) The Cancer Genome Atlas (TCGA): an immeasurable source of knowledge. Contemp Oncol (pozn) 19(1A):A68-77. https://doi.org/10.5114/wo.2014.47136
Treiber T, Treiber N, Plessmann U, Harlander S, Daiß JL, Eichner N, Lehmann G, Schall K, Urlaub H, Meister G (2017) A compendium of RNA-binding proteins that regulate microRNA biogenesis. Mol Cell 66(2):270-284 e213. https://doi.org/10.1016/j.molcel.2017.03.014
Urbanski LM, Leclair N, Anczuków O (2018) Alternative-splicing defects in cancer: splicing regulators and their downstream targets guiding the way to novel cancer therapeutics. Wiley Interdiscip Rev RNA 9(4):e1476. https://doi.org/10.1002/wrna.1476
Valdez BC, Yang H, Hong E, Sequitin AM (2002) Genomic structure of newly identified paralogue of RNA helicase II/Gu: detection of pseudogenes and multiple alternatively spliced mRNAs. Gene 284(1–2):53–61. https://doi.org/10.1016/s0378-1119(01)00888-5
Wang ET, Sandberg R, Luo S, Khrebtukova I, Zhang L, Mayr C, Kingsmore SF, Schroth GP, Burge CB (2008) Alternative isoform regulation in human tissue transcriptomes. Nature 456(7221):470–476. https://doi.org/10.1038/nature07509
Wang J, Xu W, Wang B, Lin G, Wei Y, Abudurexiti M, Zhu W, Liu C, Qin X, Dai B, Wan F, Zhang H, Zhu Y, Ye D (2020) GLUT1 is an AR target contributing to tumor growth and glycolysis in castration-resistant and enzalutamide-resistant prostate cancers. Cancer Lett 485:45–55. https://doi.org/10.1016/j.canlet.2020.05.007
Xing YH, Yao RW, Zhang Y, Guo CJ, Jiang S, Xu G, Dong R, Yang L, Chen LL (2017) SLERT regulates DDX21 rings associated with pol I transcription. Cell 169(4):664-678 e616. https://doi.org/10.1016/j.cell.2017.04.011
Xu WH, Shi SN, Xu Y, Wang J, Wang HK, Cao DL, Shi GH, Qu YY, Zhang HL, Ye DW (2019a) Prognostic implications of Aquaporin 9 expression in clear cell renal cell carcinoma. J Transl Med 17(1):363. https://doi.org/10.1186/s12967-019-2113-y
Xu WH, Wu J, Wang J, Wan FN, Wang HK, Cao DL, Qu YY, Zhang HL, Ye DW (2019b) Screening and identification of potential prognostic biomarkers in adrenocortical carcinoma. Front Genet 10:821. https://doi.org/10.3389/fgene.2019.00821
Xu WH, Xu Y, Wang J, Wan FN, Wang HK, Cao DL, Shi GH, Qu YY, Zhang HL, Ye DW (2019c) Prognostic value and immune infiltration of novel signatures in clear cell renal cell carcinoma microenvironment. Aging (albany NY) 11(17):6999–7020. https://doi.org/10.18632/aging.102233
Xu N, Ke ZB, Lin XD, Lin F, Chen SH, Wu YP, Chen YH, Wei Y, Zheng QS (2020) Identification of survival-associated alternativesplicing events and signatures in adrenocortical carcinoma based on TCGA SpliceSeq data. Aging (Albany NY) 12(6):4996–5009. https://doi.org/10.18632/aging.102924
Xu W, Tian X, Liu W, Anwaier A, Su J, Zhu W, Wan F, Shi G, Wei G, Qu Y, Zhang H, Ye D (2021) m6A regulator-mediated methylation modification model predicts prognosis tumor microenvironment characterizations and response to immunotherapies of clear cell renal cell carcinoma. Front Oncol. https://doi.org/10.3389/fonc.2021.709579
Zhang H, Zhang Y, Chen C, Zhu X, Zhang C, Xia Y, Zhao Y, Andrisani OM, Kong L (2018) A double-negative feedback loop between DEAD-box protein DDX21 and Snail regulates epithelial-mesenchymal transition and metastasis in breast cancer. Cancer Lett 437:67–78. https://doi.org/10.1016/j.canlet.2018.08.021
Zhao N, Zhang J (2018) Role of alternative splicing of VEGF-A in the development of atherosclerosis. Aging (albany NY) 10(10):2695–2708. https://doi.org/10.18632/aging.101580
Zuberi K, Franz M, Rodriguez H, Montojo J, Lopes CT, Bader GD, Morris Q (2013) GeneMANIA prediction server 2013 update. Nucleic Acids Res 41(Web Server issue):W115-122. https://doi.org/10.1093/nar/gkt533
Acknowledgements
We are grateful to all patients for their dedicated participation in the current study. We expressed our sincere gratitude to Ms. Zoo for editing figures and drawing the schematic diagram for this study.
Funding
This work is supported by National Key Research and Development Program of China (No. 2019YFC1316000), Fuqing Scholar Student Scientific Research Program of Shanghai Medical College, Fudan University (No. FQXZ202112B), Natural Science Foundation of Shanghai (No. 20ZR1413100), and Shanghai Municipal Health Bureau (No. 2020CXJQ03).
Author information
Authors and Affiliations
Contributions
Conceptualization: WX, AA, WL, and HZ. Data curation and formal analysis: WX, AA, WL, JW, XT, and WZ. Funding acquisition: YQ, HZ, and DY. Investigation and methodology: WX, AA, WZ, and WL. Resources and software: WL, WZ, YQ, HZ, and DY. Supervision: YQ, JW, HZ and DY. Validation and visualization: WX, WL, XT, and AA. Original draft: WX, AA, and WL. Editing: YQ, HZ, and DY.
Corresponding authors
Ethics declarations
Conflicts of interest
The authors have no conflicts of interest to declare that are relevant to the content of this article.
Ethics approval and consent to participate
Study procedures were approved by Fudan University Shanghai Cancer Center (Shanghai, China) (ID: 050432-4-1805C). Written informed consents were acquired from open-access online databases.
Consent for publication
Not applicable.
Supplementary Information
Below is the link to the electronic supplementary material.
43657_2021_26_MOESM1_ESM.tif
Fig. S1. LASSO regression analysis was constructed in PASEs in (A) AA, (B) AD, (C) AP, (D) AT, (E) ES, (F) ME, (G) RI and (H) all types of AS events, respectively. Left patterns in each group were LASSO coefficient profiles, and right patterns in each group were the selection of the tuning parameter (λ) in the LASSO model. (TIF 583 KB)
43657_2021_26_MOESM2_ESM.tif
Fig. S2. Survival risk assessment and Cox regression analysis of PASEs. Survival risk assessment identified patients with high-risk score, and suggested markedly relationship between distribution of risk score and survival time in (A) AA, (B) AD, (C) AP, (D) AT, (E) ES, (F) ME, (G) RI and (H) all types of AS events groups. BLOC1S1-AP, TRAFD1-AD, CIRBP-ES and USMG5-ES were identified as significant differential AS event in all types group. (TIF 3612 KB)
43657_2021_26_MOESM3_ESM.tif
Fig. S3. Differential BAG2 expressed and its prognostic value. (A) BAG2 mRNA expressed was decreased in tumor compared with normal tissues, (B) while no statistically significant association between BAG2 expression and prognosis was found in 76 ACC patients from TCGA cohort. (TIF 155 KB)
43657_2021_26_MOESM4_ESM.tif
Fig. S4. Molecular features between AS events and classification the samples according to the cluster of clusters (CoCs) from four platforms (DNA copy number; mRNA expression; DNA methylation; miRNA expression) in TCGA-ACC cohort. (A) The Risk Score constructed using all significant AS event markedly increased with elevated CoCs level using ANOVA test (p=0.012). (B) DDX21 mRNA expression significantly correlated with CoCs level in TCGA-ACC cohort. (TIF 903 KB)
Rights and permissions
About this article
Cite this article
Xu, W., Anwaier, A., Liu, W. et al. Systematic Genome-Wide Profiles Reveal Alternative Splicing Landscape and Implications of Splicing Regulator DExD-Box Helicase 21 in Aggressive Progression of Adrenocortical Carcinoma. Phenomics 1, 243–256 (2021). https://doi.org/10.1007/s43657-021-00026-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s43657-021-00026-x