Abstract
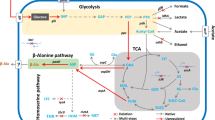
β-Alanine is the only naturally occurring β-type amino acid, with various applications in the pharmaceutical, food, and chemical industries. Given the growing market demand, the study of β-alanine production is important. This study utilized a modified lysine-producing strain as a chassis cell line to further promote β-alanine synthesis through metabolic engineering. In order to reduce the consumption of oxaloacetate, the gene pck was deleted. A promoter mutation library was constructed to screen the original promoter of the stronger promoter replacement gene pyc to enhance the oxaloacetate synthesis pathway and further increase the intracellular supply of oxaloacetate. Next, the gene poxB was deleted, and pyruvate accumulation further promoted β-alanine synthesis. Then, the aspartate kinase-coding gene lysC was weakened by predicting the RBS sequence, thus reducing the synthesis of lysine by-products and improving β-alanine synthesis. Ultimately, the carbon flux in the β-alanine biosynthetic pathways was increased by overexpressing aspartate-α-decarboxylase, aspartate ammonia-lyase, and aspartate aminotransferase using the strong promoter Ptrc. The resulting strain QBA9 was cultured in a 5-L fermenter by fed-batch to produce 70.8 g/L of β-alanine with a productivity of 0.98 g/L/h. These modification strategies demonstrate the potential for efficient β-alanine production by the lysine-producing strain and provide an innovative idea for the developing β-alanine-producing strains.








Similar content being viewed by others
References
Steunenberg P, Könst PM, Scott EL, Franssen MCR, Zuilhof H, Sanders JPM. Polymerisation of β-alanine through catalytic ester–amide exchange. Eur Polymer J. 2013;49(7):1773–81. https://doi.org/10.1016/j.eurpolymj.2013.03.032.
Sílvia ÀC, Míriam PT, Joan A, Pau F. Engineering the synthetic β-alanine pathway in Komagataella phaffii for conversion of methanol into 3-hydroxypropionic acid. Microb Cell Fact. 2023;22(1):237.
Rivera D, Roa-Sanchez P, Bido P, Speckter H, Oviedo J, Stoeter P. Cerebral and cerebellar white matter tract alterations in patients with pantothenate kinase-Associated Neurodegeneration (PKAN). Parkinsonism Relat Disord. 2022;98:1–6. https://doi.org/10.1016/j.parkreldis.2022.03.017.
Lee JW, Na D, Park JM, Lee J, Choi S, Lee SY. Systems metabolic engineering of microorganisms for natural and non-natural chemicals. Nat Chem Biol. 2012;8(6):536–46. https://doi.org/10.1038/nchembio.970.
Hossein MG, Pirzad JG. Biochemical mechanisms of Beneficial effects of Beta-Alanine supplements on Cognition. Biochem (Moscow). 2023;88(8):1181–90.
Wang L, Mao Y, Wang Z, Ma H, Chen T. Advances in biotechnological production of beta-alanine. World J Microbiol Biotechnol. 2021;37(5):79. https://doi.org/10.1007/s11274-021-03042-1.
Deng S, Zhang J, Cai Z, Li Y. [Characterization of L-aspartate-α-decarboxylase from Bacillus subtilis]. Sheng Wu gong Cheng xue bao = Chinese. J Biotechnol. 2015;31(8):1184–93.
Qian Y, Liu J, Song W, Chen X, Luo Q, Liu L. Production of β-Alanine from Fumaric Acid using a dual‐enzyme Cascade. ChemCatChem. 2018;10(21):4984–91. https://doi.org/10.1002/cctc.201801050.
Wu J, Ma B-D, Xu Y. One-Pot synthesis of β-Alanine from Maleic Acid via Three-Enzyme Cascade Biotransformation. Catalysts. 2023;13(2). https://doi.org/10.3390/catal13020267.
Wang L, Piao X, Cui S, Hu M, Tao Y. Enhanced production of beta-alanine through co-expressing two different subtypes of L-aspartate-alpha-decarboxylase. J Ind Microbiol Biotechnol. 2020;47(6–7):465–74. https://doi.org/10.1007/s10295-020-02285-5.
Irem T, Sirin U, Sinem Y, H TA, Petek BK, Zeynep S. Design, synthesis and biological evaluation of some novel Naphthoquinone-Glycine/β-Alanine anilide derivatives as non-covalent proteasome inhibitors. Volume 101. Chemical biology & drug design; 2023. 6.
Laura S, Leo O, Gopal P, Michael L, Paula J, Pekka PJ, et al. Production of biopolymer precursors beta-alanine and L-lactic acid from CO2 with metabolically versatile Rhodococcus opacus DSM 43205. Front Bioeng Biotechnol. 2022;10:989481.
Zhou H-Y, Tang Y-Q, Peng J-B, Wang S-H, Liu Z-Q, Zheng Y-G. Re-designing Escherichia coli for high-yield production of β-alanine by metabolic engineering. Biochem Eng J. 2022;189. https://doi.org/10.1016/j.bej.2022.108714.
Li B, Zhang B, Wang P, Cai X, Chen YY, Yang YF, et al. Rerouting fluxes of the Central Carbon Metabolism and relieving mechanism-based inactivation of l-Aspartate-alpha-decarboxylase for fermentative production of beta-alanine in Escherichia coli. ACS Synth Biol. 2022;11(5):1908–18. https://doi.org/10.1021/acssynbio.2c00055.
Raquel SB, Gabriel LC, Davi BO, Vitor LP, José DDSN, Adilson JS. Glycerol as substrate and NADP+-dependent glyceraldehyde-3-phosphate dehydrogenase enable higher production of 3-hydroxypropionic acid through the β-alanine pathway in E. Coli Bioresource Technol. 2023;393:130142.
Shilong H, Mingyue F, Beibei F, Mingjing Y, Panhong Y, Biao T, et al. Development of probiotic E. Coli Nissle 1917 for β-alanine production by using protein and metabolic engineering. Appl Microbiol Biotechnol. 2023;107(7–8):2277–88.
Zhao N, Qian L, Luo G, Zheng S. Synthetic biology approaches to access renewable carbon source utilization in Corynebacterium glutamicum. Appl Microbiol Biotechnol. 2018;102(22):9517–29. https://doi.org/10.1007/s00253-018-9358-x.
Shen Y, Zhao L, Li Y, Zhang L, Shi G. Synthesis of β-alanine from L-aspartate using L-aspartate-α-decarboxylase from Corynebacterium glutamicum. Biotechnol Lett. 2014;36(8):1681–6. https://doi.org/10.1007/s10529-014-1527-0.
Ghiffary MR, Prabowo CPS, Adidjaja JJ, Lee SY, Kim HU. Systems metabolic engineering of Corynebacterium glutamicum for the efficient production of beta-alanine. Metab Eng. 2022;74:121–9. https://doi.org/10.1016/j.ymben.2022.10.009.
Wang JY, Rao ZM, Xu JZ, Zhang WG. Enhancing beta-alanine production from glucose in genetically modified Corynebacterium glutamicum by metabolic pathway engineering. Appl Microbiol Biotechnol. 2021;105(24):9153–66. https://doi.org/10.1007/s00253-021-11696-y.
Wang YY, Shi K, Chen P, Zhang F, Xu JZ, Zhang WG. Rational modification of the carbon metabolism of Corynebacterium glutamicum to enhance L-leucine production. J Ind Microbiol Biotechnol. 2020;47(6–7):485–95. https://doi.org/10.1007/s10295-020-02282-8.
Song CW, Lee J, Ko YS, Lee SY. Metabolic engineering of Escherichia coli for the production of 3-aminopropionic acid. Metab Eng. 2015;30:121–9. https://doi.org/10.1016/j.ymben.2015.05.005.
Klaffl S, Eikmanns BJ. Genetic and functional analysis of the soluble oxaloacetate decarboxylase from Corynebacterium glutamicum. J Bacteriol. 2010;192(10):2604–12. https://doi.org/10.1128/JB.01678-09.
Petersen S, Mack C, de Graaf AA, Riedel C, Eikmanns BJ, Sahm H. Metabolic consequences of altered phosphoenolpyruvate carboxykinase activity in Corynebacterium glutamicum reveal anaplerotic regulation mechanisms in vivo. Metab Eng. 2001;3(4):344–61. https://doi.org/10.1006/mben.2001.0198.
Chen Z, Bommareddy RR, Frank D, Rappert S, Zeng AP. Deregulation of feedback inhibition of phosphoenolpyruvate carboxylase for improved lysine production in Corynebacterium glutamicum. Appl Environ Microbiol. 2014;80(4):1388–93. https://doi.org/10.1128/AEM.03535-13.
Neuner A, Wagner I, Sieker T, Ulber R, Schneider K, Peifer S, et al. Production of L-lysine on different silage juices using genetically engineered Corynebacterium glutamicum. J Biotechnol. 2013;163(2):217–24. https://doi.org/10.1016/j.jbiotec.2012.07.190.
Liu J, Liu M, Shi T, Sun G, Gao N, Zhao X, et al. CRISPR-assisted rational flux-tuning and arrayed CRISPRi screening of an L-proline exporter for L-proline hyperproduction. Nat Commun. 2022;13(1):891. https://doi.org/10.1038/s41467-022-28501-7.
Zelle RM, de Hulster E, van Winden WA, de Waard P, Dijkema C, Winkler AA, et al. Malic acid production by Saccharomyces cerevisiae: engineering of pyruvate carboxylation, oxaloacetate reduction, and malate export. Appl Environ Microbiol. 2008;74(9):2766–77. https://doi.org/10.1128/AEM.02591-07.
Sasaki Y, Eng T, Herbert RA, Trinh J, Chen Y, Rodriguez A, et al. Engineering Corynebacterium glutamicum to produce the biogasoline isopentenol from plant biomass hydrolysates. Biotechnol Biofuels. 2019;12:41. https://doi.org/10.1186/s13068-019-1381-3.
Yang M, Zhang X. Construction of pyruvate producing strain with intact pyruvate dehydrogenase and genome-wide transcription analysis. World J Microbiol Biotechnol. 2017;33(3):59. https://doi.org/10.1007/s11274-016-2202-5.
Liu J, Xu JZ, Rao ZM, Zhang WG. Industrial production of L-lysine in Corynebacterium glutamicum: Progress and prospects. Microbiol Res. 2022;262:127101. https://doi.org/10.1016/j.micres.2022.127101.
Dong X, Zhao Y, Zhao J, Wang X. Characterization of aspartate kinase and homoserine dehydrogenase from Corynebacterium glutamicum IWJ001 and systematic investigation of L-isoleucine biosynthesis. J Ind Microbiol Biotechnol. 2016;43(6):873–85. https://doi.org/10.1007/s10295-016-1763-5.
Kalinowski J, Cremer J, Bachmann B, Eggeling L, Sahm H, Puhler A. Genetic and biochemical analysis of the aspartokinase from Corynebacterium glutamicum. Mol Microbiol. 1991;5(5):1197–204. https://doi.org/10.1111/j.1365-2958.1991.tb01893.x.
Salis HM, Mirsky EA, Voigt CA. Automated design of synthetic ribosome binding sites to control protein expression. Nat Biotechnol. 2009;27(10):946–50. https://doi.org/10.1038/nbt.1568.
Lin L, Wang Y, Wu M, Zhu L, Yang L, Lin J. Enhancing the thermostability of fumarase C from Corynebacterium glutamicum via molecular modification. Enzyme Microb Technol. 2018;115:45–51. https://doi.org/10.1016/j.enzmictec.2018.04.010.
Tomoko N, Kenji T, Mitsuhiro I, et al. Metabolic engineering of carotenoid biosynthesis in Escherichia coli by ordered gene assembly in Bacillus subtilis[J]. Appl Environ Microbiol. 2007;73(4):1355–61. https://doi.org/10.1128/AEM.02268-06.
Li H, Lu X, Chen K, Yang J, Zhang A, Wang X, et al. β-alanine production using whole-cell biocatalysts in recombinant Escherichia coli. Mol Catal. 2018;449:93–8. https://doi.org/10.1016/j.mcat.2018.02.008.
Acknowledgements
The research has been funded by the “Important Amino Acid Industrial Strain System Transformation and Industrial Demonstration” project [grant numbers 2021YFC2100900].
Author information
Authors and Affiliations
Contributions
YS, FZ, JX and WZ conceived and designed the research. YS conducted experiments. YS, JL, JX and WZ analyzed the data and wrote the manuscript. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare that they have no confict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Song, YQ., Zhang, F., Liu, J. et al. Metabolic engineering of Corynebacterium glutamicum for the efficient production of β-Alanine from glucose. Syst Microbiol and Biomanuf (2024). https://doi.org/10.1007/s43393-024-00268-6
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s43393-024-00268-6