Abstract
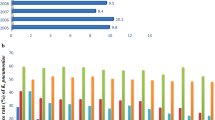
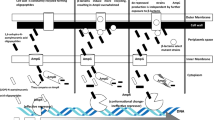
Sichuan is a significant aquaculture province in China, with a total aquaculture output of 1.72 × 106 tons in 2022. One of the most significant microorganisms hurting the Sichuan aquaculture is Aeromonas hydrophila, whose genotype and antibiotic resistance are yet unknown. This study isolated a total of 64 strains of A. hydrophila from various regions during September 2019 to June 2021 within Sichuan province, China. The technique of Multi-Locus Sequence Typing (MLST) was used for the purpose of molecular typing. Meanwhile, identification of antibiotic resistance phenotype and antibiotic resistance gene was performed. The findings of the study revealed that 64 isolates exhibited 29 sequence types (ST) throughout different regions in Sichuan, with 25 of these ST types being newly identified. Notably, the ST251 emerged as the predominant sequence type responsible for the pandemic. The resistance rate of isolated strains to roxithromycin was as high as 98.3%, followed by co-trimoxazole (87.5%), sulfafurazole (87.5%), imipenem (80%), amoxicillin (60%), and clindamycin (57.8%). Fifteen strains of A. hydrophila exhibited resistance to medicines across a minimum of three categories, suggesting the development of multidrug resistance in these isolates. A total of 63 ARGs were detected from the isolates, which mediated a range of antibiotic resistance mechanisms, with deactivation and efflux potentially serving as the primary mechanisms of antibiotic resistance. This study revealed the diversity of A. hydrophila genotypes and the risk of antibiotic resistance in Sichuan, providing reference for scientific and effective control of A. hydrophila infection.





Similar content being viewed by others
Data availability
The data used to support the findings of this study are included within the article that are available from the corresponding author upon request.
References
Stratev D, Odeyemi OA (2016) Antimicrobial resistance of Aeromonas hydrophila isolated from different food sources: a mini-review. J Infect Public Health 9:535–544
Janda JM, Abbott SL (2010) The genus Aeromonas: taxonomy, pathogenicity, and infection. Clin Microbiol Rev 23:35–73
Pianetti A, Battistelli M, Citterio B, Parlani C, Falcieri E, Bruscolini F (2009) Morphological changes of Aeromonas hydrophila in response to osmotic stress. Micron 40:426–433
Rashidian G, Boldaji JT, Rainis S, Prokic MD, Faggio C (2021) Oregano (Origanum vulgare) extract enhances zebrafish (Danio rerio) growth performance, serum and mucus innate immune responses and resistance against Aeromonas hydrophila challenge. Animals (Basel) 11:299
Kaur B, Naveen Kumar BT, Tyagi A, AdmaneHoleyappa S, Singh NK (2021) Identification of novel vaccine candidates in the whole-cell Aeromonas hydrophila biofilm vaccine through reverse vaccinology approach. Fish Shellfish Immunol 114:132–141
Collignon P, Beggs JJ, Walsh TR, Gandra S, Laxminarayan R (2018) Anthropological and socioeconomic factors contributing to global antimicrobial resistance: a univariate and multivariable analysis. The Lancet Planetary Health 2:e398–e405
Elbehiry A, Marzouk E, Abdeen E, Al-Dubaib M, Alsayeqh A, Ibrahem M, Hamada M, Alenzi A, Moussa I, Hemeg HA (2019) Proteomic characterization and discrimination of Aeromonas species recovered from meat and water samples with a spotlight on the antimicrobial resistance of Aeromonas hydrophila. Microbiologyopen 8:e782
Nnadozie CF, Odume ON (2019) Freshwater environments as reservoirs of antibiotic resistant bacteria and their role in the dissemination of antibiotic resistance genes. Environ Pollut 254:113067
Khor WC, Puah SM, Koh TH, Tan J, Puthucheary SD, Chua KH (2018) Comparison of clinical isolates of Aeromonas from Singapore and Malaysia with regard to molecular identification, virulence, and antimicrobial profiles. Microb Drug Resist 24:469–478
Ramadan H, Ibrahim N, Samir M, Abd El-Moaty A, Gad T (2018) Aeromonas hydrophila from marketed mullet (Mugil cephalus) in Egypt: PCR characterization of beta-lactam resistance and virulence genes. J Appl Microbiol 124:1629–1637
Lau TTV, Tan JMA, Puthucheary SD, Puah SM, Chua KH (2020) Genetic relatedness and novel sequence types of clinical Aeromonas dhakensis from Malaysia. Braz J Microbiol 51:909–918
FISHERIES BO (2023) China fishery statistics yearbook. China Agriculture Press
Nicholson P, Mon-on N, Jaemwimol P, Tattiyapong P, Surachetpong W (2020) Coinfection of tilapia lake virus and Aeromonas hydrophila synergistically increased mortality and worsened the disease severity in tilapia (Oreochromis spp.). Aquaculture 520. https://doi.org/10.1016/j.aquaculture.2019.734746
Persson S, Al-Shuweli S, Yapici S, Jensen JN, Olsen KE (2015) Identification of clinical aeromonas species by rpoB and gyrB sequencing and development of a multiplex PCR method for detection of Aeromonas hydrophila, A. caviae, A. veronii, and A. media. J Clin Microbiol 53:653–656
Martino ME, Fasolato L, Montemurro F, Rosteghin M, Manfrin A, Patarnello T, Novelli E, Cardazzo B (2011) Determination of microbial diversity of Aeromonas strains on the basis of multilocus sequence typing, phenotype, and presence of putative virulence genes. Appl Environ Microbiol 77:4986–5000
Church DL, Cerutti L, Gurtler A, Griener T, Zelazny A, Emler S (2020) Performance and application of 16S rRNA gene cycle sequencing for routine identification of bacteria in the clinical microbiology laboratory. Clin Microbiol Rev 33:e00053-e119
Fernandez-Bravo A, Figueras MJ (2020) An update on the genus Aeromonas: taxonomy, epidemiology, and pathogenicity. Microorganisms 8:129
Sautour M, Mary P, Chihib NE, Hornez JP (2003) The effects of temperature, water activity and pH on the growth of Aeromonas hydrophila and on its subsequent survival in microcosm water. J Appl Microbiol 95:807–813
Maiden MC, Jansen van Rensburg MJ, Bray JE, Earle SG, Ford SA, Jolley KA, McCarthy ND (2013) MLST revisited: the gene-by-gene approach to bacterial genomics. Nat Rev Microbiol 11:728–736
Awan F, Dong Y, Liu J, Wang N, Mushtaq MH, Lu C, Liu Y (2018) Comparative genome analysis provides deep insights into Aeromonas hydrophila taxonomy and virulence-related factors. BMC Genomics 19:712
Zhang X, Yang W, Wu H, Gong X, Li A (2014) Multilocus sequence typing revealed a clonal lineage of Aeromonas hydrophila caused motile Aeromonas septicemia outbreaks in pond-cultured cyprinid fish in an epidemic area in central China. Aquaculture 432:1–6
Pang M, Jiang J, Xie X, Wu Y, Dong Y, Kwok AH, Zhang W, Yao H, Lu C, Leung FC, Liu Y (2015) Novel insights into the pathogenicity of epidemic Aeromonas hydrophila ST251 clones from comparative genomics. Sci Rep 5:9833
Rasmussen-Ivey CR, Hossain MJ, Odom SE, Terhune JS, Hemstreet WG, Shoemaker CA, Zhang D, Xu DH, Griffin MJ, Liu YJ, Figueras MJ, Santos SR, Newton JC, Liles MR (2016) Classification of a hypervirulent Aeromonas hydrophila pathotype responsible for epidemic outbreaks in warm-water fishes. Front Microbiol 7:1615
Nikiforov I, Goldman J, Cheriyath P, Vyas A, Nookala V (2014) Aeromonas hydrophila sepsis associated with consumption of raw oysters. Case Rep Infect Dis 2014:163040
Saleh AER, Younis G (2021) Virulent and multiple antimicrobial resistance Aeromonas hydrophila isolated from diseased Nile tilapia fish (Oreochromis niloticus) in Egypt with sequencing of some virulence-associated genes. Biocontrol Sci 26:167–176
Fikri F, Wardhana DK, Purnomo A, Khairani S, Chhetri S, Purnama MTE (2022) Aerolysin gene characterization and antimicrobial resistance profile of Aeromonas hydrophila isolated from milkfish (Chanos chanos) in Gresik, Indonesia. Vet World 15:1759–1764
Schubert RH, Matzinou D (1990) Temperature as an environmental factor influencing the pathogenicity of Aeromonas hydrophila. Zentralbl Bakteriol 273:327–331
Gong Z, Li J, Luo H, Zhan D, Liu X, Gao C, Huang J, Qian Y, Song Y, Quan W, An S, Tian Y, Hu Z, Sun J, Yuan H, Jiang R (2020) Low-temperature laminar flow ward for the treatment of multidrug resistance Acinetobacter baumannii pneumonia. Nat Public Health Emergency Collection 39:877–887
Medina E, Pieper DH (2016) Tackling threats and future problems of multidrug-resistant bacteria. Curr Top Microbiol Immunol 398:3–33
Nhinh DT, Le DV, Van KV, Huong Giang NT, Dang LT, Hoai TD (2021) Prevalence, virulence gene distribution and alarming the multidrug resistance of Aeromonas hydrophila associated with disease outbreaks in freshwater aquaculture. Antibiotics (Basel) 10:532
Woo SJ, Kim MS, Jeong MG, Do MY, Hwang SD, Kim WJ (2022) Establishment of epidemiological cut-off values and the distribution of resistance genes in Aeromonas hydrophila and Aeromonas veronii isolated from aquatic animals. Antibiotics (Basel) 11:343
Guo Y, Zeng C, Ma C, Cai H, Jiang X, Zhai S, Xu X, Lin M (2022) Comparative genomics analysis of the multidrug-resistant Aeromonas hydrophila MX16A providing insights into antibiotic resistance genes. Front Cell Infect Microbiol 12:1042350
Qiuli He GM, Liang M (2019) Analysis of drug resistance of Aeromonas parenterinal to β-lactam drugs in a hospital in Shaoxing. Int J Epidemiol Epidemiol 46:5
Dong X, Shi P, Liu W, Bai J, Bian L (2022) Metallo-beta-lactamase CphA evolving into more efficient hydrolases through gene mutation is a novel pathway for the resistance of super bacteria. Appl Microbiol Biotechnol 106:2471–2480
McMahon MA, Xu J, Moore JE, Blair IS, McDowell DA (2007) Environmental stress and antibiotic resistance in food-related pathogens. Appl Environ Microbiol 73:211–217
Kolář M, Urbánek K, Látal T (2001) Antibiotic selective pressure and development of bacterial resistance. Int J Antimicrob Agents 5:357–531
Voigt AM, Faerber HA, Wilbring G, Skutlarek D, Felder C, Mahn R, Wolf D, Brossart P, Hornung T, Engelhart S, Exner M, Schmithausen RM (2019) The occurrence of antimicrobial substances in toilet, sink and shower drainpipes of clinical units: a neglected source of antibiotic residues. Int J Hyg Environ Health 222:455–467
Boy-Roura M, Mas-Pla J, Petrovic M, Gros M, Soler D, Brusi D, Mencio A (2018) Towards the understanding of antibiotic occurrence and transport in groundwater: findings from the Baix Fluvia alluvial aquifer (NE Catalonia, Spain). Sci Total Environ 612:1387–1406
Apreja M, Sharma A, Balda S, Kataria K, Capalash N, Sharma P (2022) Antibiotic residues in environment: antimicrobial resistance development, ecological risks, and bioremediation. Environ Sci Pollut Res 29:3355–3371
Acknowledgements
We thank all authors who kindly provided their hard work in the program.
Funding
This research was supported by the Sichuan Innovation Team Project of Agricultural Industry Technology System (SCCXTD-15) and Fish Diseases Prevention and Control Project of JPDC (No.23XB0027).
Author information
Authors and Affiliations
Contributions
Kun Peng: writing, methodology, investigation. Mengzhu Chen: investigation, methodology, formal analysis. Yilin Wang: methodology, investigation. Ziqi Tian: investigation, formal analysis. Longjun Deng: investigation, data curation. Tiancai Li: data curation, formal analysis. Yang Feng: methodology, data curation. Ping Ouyang: project administration, resources. Xiaoli Huang: conceptualization, methodology. Defang Chen: writing — review and editing. Yi Geng: funding acquisition, supervision. All authors contributed to manuscript revision, read, and approved the submitted version. Kun Peng and Mengzhu Chen contributed equally to this work.
Corresponding author
Ethics declarations
Ethics approval
All animal procedures were conducted by the Animal Experiment General Requirement in China (record number GB/T 35823–2018) and were approved by the Institutional Animal Care and Use Committee (IACUC) of Sichuan Agricultural University.
Conflict of interest
The authors declare no competing interests.
Additional information
Responsible Editor: Maria Aparecida Scatamburlo Moreira
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Peng, K., Chen, M., Wang, Y. et al. Genotype diversity and antibiotic resistance risk in Aeromonas hydrophila in Sichuan, China. Braz J Microbiol 55, 901–910 (2024). https://doi.org/10.1007/s42770-023-01187-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42770-023-01187-9