Abstract
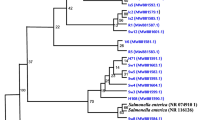
Canastra Minas Artisanal Cheese is produced in the Brazilian State of Minas Gerais using raw milk, rennet, and pingo, a natural endogenous starter culture (fermented whey) collected from the previous day’s production. Due to the use of raw milk, the product can carry microorganisms that may cause foodborne diseases (FBD), including Staphylococcus aureus. Genomic characterization of S. aureus is an important tool to assess diversity, virulence, antimicrobial resistance, and the potential for causing food poisoning due to enterotoxin production. This study is aimed at exploring the genomic features of S. aureus strains isolated from Canastra Minas Artisanal Cheeses. Multilocus sequence typing (MLST) classified these strains as ST1, ST5, and a new profile ST7849 (assigned to the clonal complex CC97). These strains belonged to four spa types: t008, t127, t359, and t992. We identified antimicrobial resistance genes with phenotypic correlation against methicillin (MRSA) and tetracycline. Virulome analysis revealed genes associated with iron uptake, immune evasion, and potential capacity for adherence and biofilm formation. The toxigenic potential included cyto- and exotoxins genes, and all strains presented the genes that encode for Panton-Valentine toxin and hemolysin, and two strains encoded 4 and 8 Staphylococcal enterotoxin (SE) genes. The results revealed the pathogenic potential of the evaluated S. aureus strains circulating in the Canastra region, representing a potential risk to public health. This study also provides useful information to monitor and guide the application of control measures to the artisanal dairy food production chain.




Similar content being viewed by others
Data availability
The assembled genome sequences were deposited in GenBank under BioProject number PRJNA803870 and accession numbers JAKNRH000000000, JAKNRI000000000, JAKNRJ000000000, JAKNRK000000000, and JAKNRL000000000.
References
Schelin J, Wallin-Carlquist N, Cohn MT, Lindqvist R, Barker GC, Rådström P (2011) The formation of Staphylococcus aureus enterotoxin in food environments and advances in risk assessment. Virulence 2(6):580–592. https://doi.org/10.4161/viru.2.6.18122
Le Loir L, Y, Baron F, Gautier M (2003) Staphylococcus aureus and food poisoning. Genet Mol Res 2(1):63–76
Baran A, Erdogan A, Turgut T, Adıgüzel M (2017) A review on the presence of Staphylococcus aureus in cheese. Turkish Journal of Nature and Science 6(2):100–105
Pritchard L, Glover RH, Humphris S, Elphinstone JG, Toth IK (2016) Genomics and taxonomy in diagnostics for food security: soft-rotting enterobacterial plant pathogens. Anal Methods 8(1):12–24. https://doi.org/10.1039/c5ay02550h
Campos GZ, Lacorte GA, Jurkiewicz C, et al. (2021) Microbiological characteristics of Canastra cheese during manufacturing and ripening. Food Control 121. https://doi.org/10.1016/j.foodcont.2020.107598
Pineda APA, Campos GZ, Pimentel-Filho NJ, Franco BDG de M, Pinto UM (2021) Brazilian artisanal cheeses: diversity, microbiological safety, and challenges for the sector. Front Microbiol 12. https://doi.org/10.3389/fmicb.2021.666922
Brasil. Ministério da Saúde. Surtos de Doenças de Transmissão Hídrica e Alimentar no Brasil Informe 2022. [WWW Document], n.d. https://www.gov.br/saude/pt-br/assuntos/saude-de-a-a-z/d/dtha/publicacoes/surtos-de-doencas-de-transmissao-hidrica-e-alimentar-no-brasil-informe-2022/view (accessed 22.2.23).
das Dores MT, Dias RS, Arcuri EF, da Nobrega JE, de Ferreira CLLF (2013) Enterotoxigenic potential of Staphylococcus aureus isolated from Artisan Minas cheese from the Serra da Canastra – MG, Brazil. Food Sci Technol 33(2):271–275. https://doi.org/10.1590/S0101-20612013005000033
Johler S, Giannini P, Jermini M, Hummerjohann J, Baumgartner A, Stephan R (2015) Further evidence for staphylococcal food poisoning outbreaks caused by egc-encoded enterotoxins. Toxins (Basel) 7(3):997–1004. https://doi.org/10.3390/toxins7030997
da Silva Cândido TJ, da Silva AC, de Matos LG, et al. (2020) Enterotoxigenic potential and molecular typing of Staphylococcus sp. isolated from organic and conventional fresh minas cheese in the state of São Paulo, Brazil. Int Dairy J 102. https://doi.org/10.1016/j.idairyj.2019.104605
Macori G, Bellio A, Bianchi DM, et al. (2020) Genome-wide profiling of enterotoxigenic Staphylococcus aureus strains used for the production of naturally contaminated cheeses. Genes (Basel) 11(1). https://doi.org/10.3390/genes11010033
Chacón RD, Ramírez M, Rodríguez-Cueva CL, et al. (2023) Genomic characterization and genetic profiles of Salmonella gallinarum strains isolated from layers with fowl typhoid in Colombia. Genes (Basel) 14(4). https://doi.org/10.3390/genes14040823
Igbinosa EO, Beshiru A, Akporehe LU, Oviasogie FE, Igbinosa OO (2023) Prevalence of methicillin-resistant Staphylococcus aureus and other Staphylococcus species in raw meat samples intended for human consumption in Benin City, Nigeria: implications for public health. Int J Environ Res Public Health 13(10). https://doi.org/10.3390/ijerph13100949
Clinical and Laboratory Standards Institute (CLSI) (2022) M100—performance standards for antimicrobial susceptibility testing, 32nd edn. Clinical and Laboratory Standards Institute, Wayne, PA
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30(15):2114–2120. https://doi.org/10.1093/bioinformatics/btu170
Gurevich A, Saveliev V, Vyahhi N, Tesler G (2013) QUAST: quality assessment tool for genome assemblies. Bioinformatics 29(8):1072–1075. https://doi.org/10.1093/bioinformatics/btt086
Simão FA, Waterhouse RM, Ioannidis P, Kriventseva EV, Zdobnov EM (2015) BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31(19):3210–3212. https://doi.org/10.1093/bioinformatics/btv351
Jolley KA, Bray JE, Maiden MCJ (2018) Open-access bacterial population genomics: BIGSdb software, the PubMLST.org website and their applications [version 1; referees: 2 approved]. Wellcome Open Res 3. https://doi.org/10.12688/wellcomeopenres.14826.1
Jia B, Raphenya AR, Alcock B et al (2017) CARD 2017: expansion and model-centric curation of the comprehensive antibiotic resistance database. Nucleic Acids Res 45(D1):D566–D573. https://doi.org/10.1093/nar/gkw1004
Liu B, Zheng D, Jin Q, Chen L, Yang J (2019) VFDB 2019: a comparative pathogenomic platform with an interactive web interface. Nucleic Acids Res 47(D1):D687–D692. https://doi.org/10.1093/nar/gky1080
Xie Z, Tang H (2017) ISEScan: automated identification of insertion sequence elements in prokaryotic genomes. Bioinformatics 33(21):3340–3347. https://doi.org/10.1093/bioinformatics/btx433
Alikhan NF, Petty NK, Ben Zakour NL, Beatson SA (2011) BLAST Ring Image Generator (BRIG): simple prokaryote genome comparisons. BMC Genomics 12. https://doi.org/10.1186/1471-2164-12-402
Silva NCC, Guimarães FF, Manzi MP et al (2013) Molecular characterization and clonal diversity of methicillin-susceptible Staphylococcus aureus in milk of cows with mastitis in Brazil. J Dairy Sci 96(11):6856–6862. https://doi.org/10.3168/jds.2013-6719
Bonsaglia ECR, Silva NCC, Rossi BF et al (2018) Molecular epidemiology of methicillin-susceptible Staphylococcus aureus (MSSA) isolated from milk of cows with subclinical mastitis. Microb Pathog 124:130–135. https://doi.org/10.1016/j.micpath.2018.08.031
Mama OM, Aspiroz C, Lozano C et al (2021) Penicillin susceptibility among invasive MSSA infections: a multicentre study in 16 Spanish hospitals. J Antimicrob Chemother 76(10):2519–2527. https://doi.org/10.1093/jac/dkab208
Aires-de-Sousa M, Parente CESR, Vieira-da-Motta O, Bonna ICF, Silva DA, De Lencastre H (2007) Characterization of Staphylococcus aureus isolates from buffalo, bovine, ovine, and caprine milk samples collected in Rio de Janeiro State, Brazil. Appl Environ Microbiol 73(12):3845–3849. https://doi.org/10.1128/AEM.00019-07
Pereira-Franchi EPL, Barreira MRN, de Costa NSLM et al (2019) Molecular epidemiology of methicillin-resistant Staphylococcus aureus in the Brazilian primary health care system. Trop Med Int Health 24(3):339–347. https://doi.org/10.1111/tmi.13192
Dabul ANG, Camargo ILBC (2014) Clonal complexes of Staphylococcus aureus: all mixed and together. FEMS Microbiol Lett 351(1):7–8. https://doi.org/10.1111/1574-6968.12358
Kaatz GW, McAleese F, Seo SM (2005) Multidrug resistance in Staphylococcus aureus due to overexpression of a novel multidrug and toxin extrusion (MATE) transport protein. Antimicrob Agents Chemother 49(5):1857–1864. https://doi.org/10.1128/AAC.49.5.1857-1864.2005
Mores CR, Montelongo C, Putonti C, Wolfe AJ, Abouelfetouh A (2021) Investigation of plasmids among clinical Staphylococcus aureus and Staphylococcus haemolyticus isolates from Egypt. Front Microbiol 12. https://doi.org/10.3389/fmicb.2021.659116
Asante J, Hetsa BA, Amoako DG, Abia ALK, Bester LA, Essack SY (2021) Genomic analysis of antibiotic-resistant Staphylococcus epidermidis isolates from clinical sources in the Kwazulu-Natal Province, South Africa. Front Microbiol 12. https://doi.org/10.3389/fmicb.2021.656306
Phillips-Jones MK, Harding SE (2018) Antimicrobial resistance (AMR) nanomachines—mechanisms for fluoroquinolone and glycopeptide recognition, efflux and/or deactivation. Biophys Rev 10(2):347–362. https://doi.org/10.1007/s12551-018-0404-9
Fournier B, Klier A, Rapoport G (2001) The two-component system ArlS-ArlR is a regulator of virulence gene expression in Staphylococcus aureus. Mol Microbiol 41(1):247–261. https://doi.org/10.1046/j.1365-2958.2001.02515.x
Lekshmi M, Ammini P, Adjei J et al (2018) Modulation of antimicrobial efflux pumps of the major facilitator superfamily in <em>Staphylococcus aureus</em>. AIMS Microbiol 4(1):1–18. https://doi.org/10.3934/microbiol.2018.1.1
Fu Z, Liu Y, Chen C, et al. (2016) Characterization of fosfomycin resistance gene, fosB, in methicillin-resistant Staphylococcus aureus isolates. PLoS One 11(5). https://doi.org/10.1371/journal.pone.0154829
Fowler PW, Cole K, Gordon NC et al (2018) Robust prediction of resistance to trimethoprim in Staphylococcus aureus. Cell Chem Biol 25(3):339-349.e4. https://doi.org/10.1016/j.chembiol.2017.12.009
Ito T, Hiramatsu K (1998) Acquisition of methicillin resistance and progression of multiantibiotic resistance in methicillin-resistant Staphylococcus aureus. Yonsei Med J 39:526–533. https://doi.org/10.3349/ymj.1998.39.6.526
da Silva Abreu AC, Matos LG, da Silva Cândido TJ, et al. (2021) Antimicrobial resistance of Staphylococcus spp. isolated from organic and conventional Minas Frescal cheese producers in São Paulo, Brazil. J Dairy Sci 104(4). https://doi.org/10.3168/jds.2020-19338
Kateete DP, Bwanga F, Seni J, et al. (2019) CA-MRSA and HA-MRSA coexist in community and hospital settings in Uganda. Antimicrob Resist Infect Control 8(1). https://doi.org/10.1186/s13756-019-0551-1
Foster TJ, Höök M (1998) Surface protein adhesins of Staphylococcus aureus. Trends Microbiol 6(12):484–488. https://doi.org/10.1016/s0966-842x(98)01400-0
Harraghy N, Hussain M, Haggar A et al (2003) The adhesive and immunomodulating properties of the multifunctional Staphylococcus aureus protein Eap. Microbiology (N Y) 149(10):2701–2707. https://doi.org/10.1099/mic.0.26465-0
Soltani E, Farrokhi E, Zamanzad B, et al. (2019) Prevalence and distribution of adhesins and the expression of fibronectin-binding protein (FnbA and FnbB) among Staphylococcus aureus isolates from Shahrekord Hospitals 06 Biological Sciences 0604 Genetics. BMC Res Notes 12(1). https://doi.org/10.1186/s13104-019-4055-0
Cassat JE, Dunman PM, McAleese F, Murphy E, Projan SJ, Smeltzer MS (2005) Comparative genomics of Staphylococcus aureus musculoskeletal isolates. J Bacteriol 187(2):576–592. https://doi.org/10.1128/JB.187.2.576-592.2005
Ajayi C, Åberg E, Askarian F, Sollid JUE, Johannessen M, Hanssen AM (2018) Genetic variability in the sdrD gene in Staphylococcus aureus from healthy nasal carriers. BMC Microbiol 18(1). https://doi.org/10.1186/s12866-018-1179-7
Götz F (2002) Staphylococcus and biofilms. Mol Microbiol 43(6):1367–1378. https://doi.org/10.1046/j.1365-2958.2002.02827.x
Cao Z, Casabona MG, Kneuper H, Chalmers JD, Palmer T (2016) The type VII secretion system of Staphylococcus aureus secretes a nuclease toxin that targets competitor bacteria. Nat Microbiol 2(1). https://doi.org/10.1038/nmicrobiol.2016.183
Batool N, Shamim A, Chaurasia AK, Kim KK (2021) Genome-wide analysis of Staphylococcus aureus sequence type 72 isolates provides insights into resistance against antimicrobial agents and virulence potential. Front Microbiol 11:613800. https://doi.org/10.3389/fmicb.2020.613800
Haupt K, Reuter M, Van Den Elsen J, et al. (2008) The Staphylococcus aureus protein Sbi acts as a complement inhibitor and forms a tripartite complex with host complement factor H and C3b. PLoS Pathog 4(12). https://doi.org/10.1371/journal.ppat.1000250
Chan YGY, Kim HK, Schneewind O, Missiakas D (2014) The capsular polysaccharide of Staphylococcus aureus is attached to peptidoglycan by the LytR-CpsA-Psr (LCP) family of enzymes. J Biol Chem 289(22):15680–15690. https://doi.org/10.1074/jbc.M114.567669
Rooijakkers SHM, Ruyken M, Roos A et al (2005) Immune evasion by a staphylococcal complement inhibitor that acts on C3 convertases. Nat Immunol 6(9):920–927. https://doi.org/10.1038/ni1235
De Haas CJC, Veldkamp KE, Peschel A et al (2004) Chemotaxis inhibitory protein of Staphylococcus aureus, a bacterial antiinflammatory agent. J Exp Med 199(5):687–695. https://doi.org/10.1084/jem.20031636
Jonsson IM, Mazmanian SK, Schneewind O, Bremell T, Tarkowski A (2003) The role of Staphylococcus aureus sortase A and sortase B in murine arthritis. Microbes Infect 5(9):775–780. https://doi.org/10.1016/S1286-4579(03)00143-6
Johler S, Giannini P, Jermini M, Hummerjohann J, Baumgartner A, Stephan R (2015) Further evidence for staphylococcal food poisoning outbreaks caused by egc-encoded enterotoxins. Toxins (Basel) 7(3):997–1004. https://doi.org/10.3390/toxins7030997
Tong SYC, Davis JS, Eichenberger E, Holland TL, Fowler VG (2015) Staphylococcus aureus infections: epidemiology, pathophysiology, clinical manifestations, and management. Clin Microbiol Rev 28(3):603–661. https://doi.org/10.1128/CMR.00134-14
Fraser JD, Proft T (2008) The bacterial superantigen and superantigen-like proteins. Immunol Rev 225(1):226–243. https://doi.org/10.1111/j.1600-065X.2008.00681.x
Hall JW, Yang J, Guo H, Ji Y (2015) The AirSR two-component system contributes to Staphylococcus aureus survival in human blood and transcriptionally regulates sspABC operon. Front Microbiol 6:682. https://doi.org/10.3389/fmicb.2015.00682
Nguyen LT, Vogel HJ. (2016) Staphylokinase has distinct modes of interaction with antimicrobial peptides, modulating its plasminogen-activation properties. Sci Rep 6. https://doi.org/10.1038/srep31817
Paharik AE, Salgado-Pabon W, Meyerholz DK, White MJ, Schlievert PM, Horswill AR (2016) The Spl serine proteases modulate Staphylococcus aureus protein production and virulence in a rabbit model of pneumonia. mSphere 1(5). https://doi.org/10.1128/msphere.00208-16
Hennekinne JA, De Buyser ML, Dragacci S (2012) Staphylococcus aureus and its food poisoning toxins: characterization and outbreak investigation. FEMS Microbiol Rev 36(4):815–836. https://doi.org/10.1111/j.1574-6976.2011.00311.x
Filipello V, Bonometti E, Campagnani M et al (2020) Investigation and follow-up of a staphylococcal food poisoning outbreak linked to the consumption of traditional hand-crafted alm cheese. Pathogens 9(12):1–6. https://doi.org/10.3390/pathogens9121064
Gonzalez AGM, Marques LMP, Da Silva Amorim Gomes M, et al. (2020) Methicillin-resistant Staphylococcus aureus in Minas Frescal cheese: evaluation of classic enterotoxin genes, antimicrobial resistance and clonal diversity. FEMS Microbiol Lett 364(23). https://doi.org/10.1093/femsle/fnx232
Stephens C, Cho PJY, de Araujo VA, et al. (2015) Draft genome sequence of a community-associated methicillin-resistant Panton-Valentine leukocidin-positive Staphylococcus aureus sequence type 30 isolate from a pediatric patient with a lung infection in Brazil. Genome Announc 3(4). https://doi.org/10.1128/genomeA.00907-15
Wang L, Si W, Xue H, Zhao X (2018) Characterization of a functional insertion sequence ISSau2 from Staphylococcus aureus. Mob DNA 9(1).https://doi.org/10.1186/s13100-018-0108-5
McCarthy AJ, Lindsay JA (2013) Staphylococcus aureus innate immune evasion is lineage-specific: a bioinfomatics study. Infect Genet Evol 19:7–14. https://doi.org/10.1016/j.meegid.2013.06.012
Pietrocola G, Nobile G, Rindi S, Speziale P (2017) Staphylococcus aureus manipulates innate immunity through own and host-expressed proteases. Front Cell Infect Microbiol 7:166. https://doi.org/10.3389/fcimb.2017.00166
Peacock SJ, Moore CE, Justice A et al (2002) Virulent combinations of adhesin and toxin genes in natural populations of Staphylococcus aureus. Infect Immun 70(9):4987–4996. https://doi.org/10.1128/IAI.70.9.4987-4996.2002
Davis RW, Brannen AD, Hossain MJ, et al. (2016) Complete genome of Staphylococcus aureus Tager 104 provides evidence of its relation to modern systemic hospital-acquired strains. BMC Genomics 17(1). https://doi.org/10.1186/s12864-016-2433-8
Pinchuk IV, Beswick EJ, Reyes VE (2010) Staphylococcal enterotoxins. Toxins (Basel) 2(8):2177–2197. https://doi.org/10.3390/toxins2082177
Chambers HF, DeLeo FR (2009) Waves of resistance: Staphylococcus aureus in the antibiotic era. Nat Rev Microbiol 7(9):629–641. https://doi.org/10.1038/nrmicro2200
Acknowledgements
The authors thank the Sao Paulo Research Foundation (FAPESP, Brazil) for the financial support (grant #2013/07914-8) to the Food Research Center. We thank Fabiana da Silva Lima, Loredana d’Ovidio, and Luciano Queiroz for technical assistance during library preparation for WGS. We also thank Christian Hoffmann for the helpful discussions and critical review of the manuscript. We extend our special gratitude to each cheese producer who actively participated in this study, demonstrating eagerness to contribute not only to this research but also to the various other studies we undertake.
Funding
This work was supported by “Fundação de Amparo a Pesquisa de Sao Paulo (FAPESP-Brazil-grant #2013/07914–8).
Author information
Authors and Affiliations
Contributions
Conceptualization: Gustavo Lacorte and Uelinton M. Pinto; methodology: Ana P.A. Pineda, Carmen L.R. Cueva, Ruy D. Chacón, Manuel Ramírez, Débora P. Oliveira, and Gustavo Lacorte; formal analysis and investigation: Ana P.A. Pineda, Carmen L.R. Cueva, Ruy D. Chácon, Manuel Ramírez, Otávio G.G. Almeida, Débora P. Oliveira, Gustavo Lacorte, and Nathalia C.S. Silva; writing—original draft preparation: Ana P.A. Pineda, Carmen L.R. Cueva, and Ruy D. Chácon; writing—review and editing: Manuel Ramírez, Otávio G.G. Almeida, Bernadette D.G.M. Franco, Nathalia C.S. Silva, and Uelinton M. Pinto; funding acquisition: Bernadette D.G.M. Franco, Mariza Landgraf, and Uelinton M. Pinto; resources: Bernadette D.G.M. Franco, Mariza Landgraf, and Uelinton M. Pinto; supervision: Bernadette D.G.M. Franco, Gustavo Lacorte, Mariza Landgraf, Nathalia C.S. Silva, and Uelinton M. Pinto.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Responsible Editor: Luis Augusto Nero
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Pineda, A.P.A., Cueva, C.L.R., Chacón, R.D. et al. Genomic characterization of Staphylococcus aureus from Canastra Minas Artisanal Cheeses. Braz J Microbiol 54, 2103–2116 (2023). https://doi.org/10.1007/s42770-023-01099-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42770-023-01099-8