Abstract
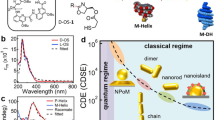
Plasmonic circular dichroism(CD) has been emerged as a promising signal for building biosensors due to its high sensitivity and specificity. In the past years, DNA nanotechnology enabled diverse chiral plasmonic devices, which can response biomolecules and then generate dynamic plasmonic CD signals at the visible range. Although some of them have been successfully employed as biosensors, the detection sensitivity is still relatively low. Herein we report a chiral plasmonic sensor with an improved detection sensitivity by integrating catalytic hairpin assembly circuits into DNA origami structures. We tested two kinds of tumor marker RNA sequences as detection targets and it turns out that the detection limit is below 10 pmol/L, improving one order of magnitude compared to previous work. The chiral plasmonic sensor with internal signal amplification circuits can stimulate a variety of smart nano-sensors for biological detection and offer a promising strategy for pathogenic RNA detection with plasmonic CD output.
Similar content being viewed by others
References
Fan Z., Govorov A. O., Nano Lett., 2010, 10, 2580
Hu Z., Meng D., Lin F., Zhu X., Fang Z. Wu X., Adv. Opt. Mater., 2019, 7, 1801590
Wang X., Tang Z., Small, 2017, 13, 1601115
Wu X. L., Hao C. L., Kumar J., Kuang H., Kotov N. A., Liz-Marzan L. M., Xu C. L., Chem. Soc. Rev., 2018, 47, 4677
Mejía-Salazar J. R., Oliveira O. N., Chem. Rev., 2018, 118, 10617
Ma W., Kuang H., Xu L. G., Ding L., Xu C. L., Wang L. B., Kotov N. A., Nat. Commun., 2013, 4, 2689
Lee H. E., Ahn H. Y., Mun J., Lee Y. Y., Kim M., Cho N. H., Chang K., Kim W. S., Rho J., Nam K. T., Nature, 2018, 556, 360
Wu X., Xu L., Ma W., Liu L., Kuang H., Yan W., Wang L., Xu C., Adv. Funct. Mater., 2015, 25, 850
Lan X., Zhou X., McCarthy L. A., Govorov A. O., Liu Y., Link S., J. Am. Chem. Soc., 2019, 141, 19336
Ma W., Xu L. G., de Moura A. F., Wu X. L., Kuang H., Xu C. L., Kotov N. A., Chem. Rev., 2017, 117, 8041
Liu N., Liedl T., Chem. Rev., 2018, 118, 3032
Kuzyk A., Schreiber R., Fan Z. Y., Pardatscher G., Roller E. M., Hogele A., Simmel F. C., Govorov A. O., Liedl T., Nature, 2012, 483, 311
Shen X. B., Song C., Wang J. Y., Shi D. W., Wang Z. A., Liu N., Ding B. Q., J. Am. Chem. Soc., 2012, 134, 146
Shen X. B., Zhan P. F., Kuzyk A., Liu Q., Asenjo-Garcia A., Zhang H., de Abajo F. J. G., Govorov A., Ding B. Q., Liu N., Nanoscale, 2014, 6, 2077
Lan X., Chen Z., Dai G., Lu X., Ni W., Wang Q., J. Am. Chem. Soc., 2013, 135, 11441
Lan X., Lu X., Shen C., Ke Y., Ni W., Wang Q., J. Am. Chem. Soc., 2015, 137, 457
Shen C., Lan X., Zhu C., Zhang W., Wang L., Wang Q., Adv. Mater., 2017, 29, 1606533
Wang M., Dong J., Zhou C., Xie H., Ni W., Wang S., Jin H., Wang Q., ACS Nano, 2019, 13, 13702
Zhou C., Duan X., Liu, N., Acc. Chem. Res., 2017, 50, 2906
Lan X., Wang Q. B., Adv. Mater., 2016, 28, 10499
Ren S., Wang J., Song C., Li Q., Yang Y., Teng N., Su S., Zhu D., Huang W., Chao J., Wang L., Fan C., Research, 2019, 2019, 7403580
Niu R., Song C., Gao F., Fang W., Jiang X., Ren S., Zhu D., Su S., Chao J., Chen S., Fan C., Wang L., Angew. Chem. Int. Ed., 2021, doi: https://doi.org/10.1002/anie.202016014
Jiang Q., Liu Q., Shi Y. F., Wang Z. G., Zhan P. F., Liu J. B., Liu C., Wang H., Shi X. H., Zhang L., Sun J. S., Ding B. Q. Liu M. H., Nano Lett., 2017, 17, 7125
Kuzyk A., Schreiber R., Zhang H., Govorov A. O., Liedl T., Liu N., Nat. Mater., 2014, 13, 862
Xin L., Zhou C., Duan X., Liu N., Nat. Commun., 2019, 10, 5394
Zhou C., Duan X. Y., Liu N., Nat. Commun., 2015, 6, 8102
Zhou C., Xin L., Duan X., Urban M. J., Liu N., Nano Lett., 2018, 18, 7395
Kuzyk A., Yang Y. Y., Duan X., Stoll S., Govorov A. O., Sugiyama H., Endo M., Liu N., Nat. Commun., 2016, 7, 10591
Kuzyk A., Urban M. J., Idili A., Ricci F., Liu N., Sci. Adv., 2017, 3, eaax6023
Dong J., Wang M., Zhou Y., Zhou C., Wang Q., Angew. Chem. Int. Ed., 2020, 59, 15038
Funck T., Nicoli F., Kuzyk A., Liedl T., Angew. Chem. Int. Ed., 2018, 57, 13495
Yin P., Choi H. M. T., Calvert C. R., Pierce N. A., Nature, 2008, 451, 318
Li B. L., Ellington A. D., Chen X., Nucleic Acids Res., 2011, 39, e110
Rothemund P. W., Nature, 2006, 440, 297
Douglas S. M., Dietz H., Liedl T., Hogberg B., Graf F., Shih W. M., Nature, 2009, 459, 414
Liu J., Zhang Y., Xie H., Zhao L., Zheng L., Ye H., Small, 2019, 15, 1902989
Wang H. M., Li C. X., Liu X. Q., Zhou X., Wang F., Chem. Sci., 2018, 9, 5842
Wei Q. M., Huang J., Li J., Wang J. L., Yang X. H., Liu J. B., Wang K. M., Chem. Sci., 2018, 9, 7802
Zang Y., Lei J., Ling P., Ju H., Anal. Chem., 2015, 87, 5430
Tang Y., Zhu Z., Lu B., Li B., Chem. Commun., 2016, 52, 13043
Qing Z. H., Hu J. L., Xu J. Y., Zou Z., Lei Y. L., Qing T. P., Yang R. H., Chem. Sci., 2020, 11, 1985
Yin J., Wang J. K., Niu R. J., Ren S. K., Wang D. X., Chao J., Chem. Res. Chinese Universities, 2020, 36(2), 219
Acknowledgements
This work was supported by the National Natural Science Foundation of China(No.21977112), the Natural Science Foundation of Jiangsu Province, China(No.BK20190227), and the Strategic Priority Research Program of Chinese Academy of Sciences(No.XDB36000000).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
The authors declare no conflicts of interest.
Supporting Information
Rights and permissions
About this article
Cite this article
Liu, Z., Dong, J., Pan, J. et al. Catalytic DNA Origami-based Chiral Plasmonic Biosensor. Chem. Res. Chin. Univ. 37, 914–918 (2021). https://doi.org/10.1007/s40242-021-1115-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40242-021-1115-x