Abstract
Purpose
Uveal melanoma (UM) with BAP1 inactivating mutations has a high risk of metastasis, but the mechanism behind BAP1 deficiency driving UM metastasis is unknown.
Methods
We analyzed the single-cell RNA sequencing (scRNA-Seq) data comprised primary and metastatic UM with or without BAP1 mutations (MUTs) to reveal inter- and intra-tumor heterogeneity among different groups. Then, an immune-competent mouse liver metastatic model was used to explore the role of ITGB2-ICAM1 in BAP1-associated UM metastasis.
Results
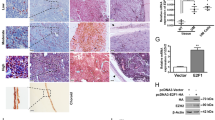
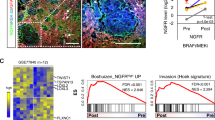
Cluster 1 tumor cells expressed high levels of genes linked to tumor metastasis, such as GDF15, ATF3, and CDKN1A, all of which are associated with poor prognosis. The strength of communication between terminally exhausted CD8+ T cells and GDF15hiATF3hiCDKN1Ahi tumor cells was enhanced in BAP1-mutated UM, with CellChat analysis predicting strong ITGB2-ICAM1 signaling between them. High expression of either ITGB2 or ICAM1 was a worse prognostic indicator. Using an immune-competent mouse liver metastatic model, we indicated that inhibiting either ICAM1 or ITGB2 prevented liver metastasis in the BAP1-mutated group in vivo. The inhibitors primarily inhibited hypoxia- and ECM-related pathways indicated by changes in the expression of genes such as ADAM8, CAV2, ENO1, PGK1, LOXL2, ITGA5, and VCAN. etc.
Conclusion
This study suggested that the ITGB2-ICAM1 axis may play a crucial role for BAP1-associated UM metastasis by preserving hypoxia- and ECM- related signatures, which provide a potential strategy for preventing UM metastasis in patients with BAP1 mutation.







Similar content being viewed by others
Data Availability
Sequence data that support the findings of this study have been deposited in NCBI database with the primary accession code PRJNA975671.
References
N. Amirouchene-Angelozzi, M. Schoumacher, M.H. Stern, N. Cassoux, L. Desjardins, S. Piperno-Neumann, O. Lantz, S. Roman-Roman, Br. J. Cancer. 113, 1249–1253 (2015). https://doi.org/10.1038/bjc.2015.269
W. Shu, X. Zhu, K. Wang, S. Cherepanoff, R.M. Conway, M.C. Madigan, H. Zhu, L. Zhu, M. Murray, F. Zhou, Cell. Oncol. (Dordr). 45, 601–619 (2022). https://doi.org/10.1007/s13402-022-00686-5
C. Pandiani, G.E. Beranger, J. Leclerc, R. Ballotti, C. Bertolotto, Genes Dev. 31, 724–743 (2017). https://doi.org/10.1101/gad.296962.117
J. Yang, D.K. Manson, B.P. Marr, R.D. Carvajal, Ther. Adv. Med. Oncol. 10, 1758834018757175 (2018). https://doi.org/10.1177/1758834018757175
M.D. Onken, L.A. Worley, J.P. Ehlers, J.W. Harbour, Cancer Res. 64, 7205–7209 (2004). https://doi.org/10.1158/0008-5472.CAN-04-1750
K.N. Smit, M.J. Jager, A. de Klein, E. Kili, Prog Retin Eye Res. 75, 100800 (2020). https://doi.org/10.1016/j.preteyeres.2019.100800
J.W. Harbour, M.D. Onken, E.D. Roberson, S. Duan, L. Cao, L.A. Worley, M.L. Council, K.A. Matatall, C. Helms, A.M. Bowcock, Science. 330, 1410–1413 (2010). https://doi.org/10.1126/science.1194472
T. Wiesner, A.C. Obenauf, R. Murali, I. Fried, K.G. Griewank, P. Ulz, C. Windpassinger, W. Wackernagel, S. Loy, I. Wolf, A. Viale, A.E. Lash, M. Pirun, N.D. Socci, A. Rutten, G. Palmedo, D. Abramson, K. Offit, A. Ott, J.C. Becker, L. Cerroni, H. Kutzner, B.C. Bastian, M.R. Speicher, Nat. Genet. 43, 1018–1021 (2011). https://doi.org/10.1038/ng.910
G.M. Shankar, M. Abedalthagafi, R.A. Vaubel, P.H. Merrill, N. Nayyar, C.M. Gill, R. Brewster, W.L. Bi, P.K. Agarwalla, A.R. Thorner, D.A. Reardon, O. Al-Mefty, P.Y. Wen, B.M. Alexander, P. van Hummelen, T.T. Batchelor, K.L. Ligon, A.H. Ligon, M. Meyerson, I.F. Dunn, R. Beroukhim, D.N. Louis, A. Perry, S.L. Carter, C. Giannini, W.T. Jr. Curry, D.P. Cahill, F.G. 2 Barker nd, P.K. Brastianos, S. Santagata, Neuro Oncol. 19, 535–545 (2017). https://doi.org/10.1093/neuonc/now235
J.R. Testa, M. Cheung, J. Pei, J.E. Below, Y. Tan, E. Sementino, N.J. Cox, A.U. Dogan, H.I. Pass, S. Trusa, M. Hesdorffer, M. Nasu, A. Powers, Z. Rivera, S. Comertpay, M. Tanji, G. Gaudino, H. Yang, M. Carbone, Nat. Genet. 43, 1022–1025 (2011). https://doi.org/10.1038/ng.912
M. Bott, M. Brevet, B.S. Taylor, S. Shimizu, T. Ito, L. Wang, J. Creaney, R.A. Lake, M.F. Zakowski, B. Reva, C. Sander, R. Delsite, S. Powell, Q. Zhou, R. Shen, A. Olshen, V. Rusch, M. Ladanyi, Nat. Genet. 43, 668–672 (2011). https://doi.org/10.1038/ng.855
Y. Jiao, T.M. Pawlik, R.A. Anders, F.M. Selaru, M.M. Streppel, D.J. Lucas, N. Niknafs, V.B. Guthrie, A. Maitra, P. Argani, G.J.A. Offerhaus, J.C. Roa, L.R. Roberts, G.J. Gores, I. Popescu, S.T. Alexandrescu, S. Dima, M. Fassan, M. Simbolo, A. Mafficini, P. Capelli, R.T. Lawlor, A. Ruzzenente, A. Guglielmi, G. Tortora, F. de Braud, A. Scarpa, W. Jarnagin, D. Klimstra, R. Karchin, V.E. Velculescu, R.H. Hruban, B. Vogelstein, K.W. Kinzler, N. Papadopoulos, L.D. Wood, Nat. Genet. 45, 1470–1473 (2013). https://doi.org/10.1038/ng.2813
T. Popova, L. Hebert, V. Jacquemin, S. Gad, V. Caux-Moncoutier, C. Dubois-d’Enghien, B. Richaudeau, X. Renaudin, J. Sellers, A. Nicolas, X. Sastre-Garau, L. Desjardins, G. Gyapay, V. Raynal, O.M. Sinilnikova, N. Andrieu, E. Manie, A. de Pauw, P. Gesta, V. Bonadona, C.M. Maugard, C. Penet, M.F. Avril, E. Barillot, O. Cabaret, O. Delattre, S. Richard, O. Caron, M. Benfodda, H.H. Hu, N. Soufir, B. Bressac-de, D. Paillerets, Stoppa-Lyonnet, M.H. Stern, Am. J. Hum. Genet. 92, 974–980 (2013). https://doi.org/10.1016/j.ajhg.2013.04.012
S. Pena-Llopis, S. Vega-Rubin-de-Celis, A. Liao, N. Leng, A. Pavia-Jimenez, S. Wang, T. Yamasaki, L. Zhrebker, S. Sivanand, P. Spence, L. Kinch, T. Hambuch, S. Jain, Y. Lotan, V. Margulis, A.I. Sagalowsky, P.B. Summerour, W. Kabbani, S.W. Wong, N. Grishin, M. Laurent, X.J. Xie, C.D. Haudenschild, M.T. Ross, D.R. Bentley, P. Kapur, J. Brugarolas, Nat. Genet. 44, 751–759 (2012). https://doi.org/10.1038/ng.2323
A. Bononi, C. Giorgi, S. Patergnani, D. Larson, K. Verbruggen, M. Tanji, L. Pellegrini, V. Signorato, F. Olivetto, S. Pastorino, M. Nasu, A. Napolitano, G. Gaudino, P. Morris, G. Sakamoto, L.K. Ferris, A. Danese, A. Raimondi, C. Tacchetti, S. Kuchay, H.I. Pass, E.B. Affar, H. Yang, P. Pinton, M. Carbone, Nature. 546, 549–553 (2017). https://doi.org/10.1038/nature22798
Y. Zhang, J. Shi, X. Liu, L. Feng, Z. Gong, P. Koppula, K. Sirohi, X. Li, Y. Wei, H. Lee, L. Zhuang, G. Chen, Z.D. Xiao, M.C. Hung, J. Chen, P. Huang, W. Li, B. Gan, Nat. Cell. Biol. 20, 1181–1192 (2018). https://doi.org/10.1038/s41556-018-0178-0
K.N. Smit, M.J. Jager, A. de Klein, E. Kiliҫ, Prog Retin Eye Res. 75, 100800 (2020). https://doi.org/10.1016/j.preteyeres.2019.100800
C.G. Ioannides, T.L. Whiteside, Clin. Immunol. Immunopathol. 66, 91–106 (1993). https://doi.org/10.1006/clin.1993.1012
F.R. Greten, S.I. Grivennikov, Immunity. 51, 27–41 (2019). https://doi.org/10.1016/j.immuni.2019.06.025
D.F. Quail, J.A. Joyce, Nat. Med. 19, 1423–1437 (2013). https://doi.org/10.1038/nm.3394
J.M. Pitt, A. Marabelle, A. Eggermont, J.C. Soria, G. Kroemer, L. Zitvogel, Ann. Oncol. 27, 1482–1492 (2016). https://doi.org/10.1093/annonc/mdw168
O. Tredan, C.M. Galmarini, K. Patel, I.F. Tannock, J. Natl. Cancer Inst. 99, 1441–1454 (2007). https://doi.org/10.1093/jnci/djm135
M. Li, P. Jiang, S. Wei, J. Wang, C. Li, Front. Immunol. 14, 1113312 (2023). https://doi.org/10.3389/fimmu.2023.1113312
C. Pandiani, T. Strub, N. Nottet, Y. Cheli, G. Gambi, K. Bille, C. Husser, M. Dalmasso, G. Beranger, S. Lassalle, V. Magnone, F. Pedeutour, M. Irondelle, C. Maschi, S. Nahon-Esteve, A. Martel, J.P. Caujolle, P. Hofman, K. LeBrigand, I. Davidson, S. Baillif, P. Barbry, R. Ballotti, C. Bertolotto, Cell. Death Differ. 28, 1990–2000 (2021). https://doi.org/10.1038/s41418-020-00730-7
M.A. Durante, D.A. Rodriguez, S. Kurtenbach, J.N. Kuznetsov, M.I. Sanchez, C.L. Decatur, H. Snyder, L.G. Feun, A.S. Livingstone, J.W. Harbour, Nat. Commun. 11, 496 (2020). https://doi.org/10.1038/s41467-019-14256-1
W. Lin, A.B. Beasley, N.M. Ardakani, E. Denisenko, L. Calapre, M. Jones, B.A. Wood, L. Warburton, A.R.R. Forrest, E.S. Gray, Cold Spring Harb Mol Case Stud. 7 (2021). https://doi.org/10.1101/mcs.a006111
Z. Souri, A.P.A. Wierenga, C. van Weeghel, P.A. van der Velden, W.G.M. Kroes, G.P.M. Luyten, S.H. van der Burg, A.G. Jochemsen, M.J. Jager, Cancers (Basel). 11 (2019). https://doi.org/10.3390/cancers11081102
C. Ericsson, S. Seregard, A. Bartolazzi, E. Levitskaya, S. Ferrone, R. Kiessling, O. Larsson, Invest. Ophthalmol. Vis. Sci. 42, 2153–2156 (2001)
M.J. Jager, H.M. Hurks, J. Levitskaya, R. Kiessling, Hum. Immunol. 63, 444–451 (2002). https://doi.org/10.1016/s0198-8859(02)00389-0
F. De Sanctis, S. Ugel, J. Facciponte, A. Facciabene, Semin Immunol. 35, 35–47 (2018). https://doi.org/10.1016/j.smim.2018.02.002
S. Soleimani, M. Shamsi, M.A. Ghazani, H.P. Modarres, K.P. Valente, M. Saghafian, M.M. Ashani, M. Akbari, A. Sanati-Nezhad, Biotechnol. Adv. 36, 880–893 (2018). https://doi.org/10.1016/j.biotechadv.2018.01.013
B.C. Miller, D.R. Sen, R. Al Abosy, K. Bi, Y.V. Virkud, M.W. LaFleur, K.B. Yates, A. Lako, K. Felt, G.S. Naik, M. Manos, E. Gjini, J.R. Kuchroo, J.J. Ishizuka, J.L. Collier, G.K. Griffin, S. Maleri, D.E. Comstock, S.A. Weiss, F.D. Brown, A. Panda, M.D. Zimmer, R.T. Manguso, F.S. Hodi, S.J. Rodig, A.H. Sharpe, W.N. Haining, Nat. Immunol. 20, 326–336 (2019). https://doi.org/10.1038/s41590-019-0312-6
L. Ny, H. Jespersen, J. Karlsson, S. Alsen, S. Filges, C. All-Eriksson, B. Andersson, A. Carneiro, H. Helgadottir, M. Levin, I. Ljuslinder, R. Olofsson Bagge, V.R. Sah, U. Stierner, A. Stahlberg, G. Ullenhag, L.M. Nilsson, J.A. Nilsson, Nat. Commun. 12, 5155 (2021). https://doi.org/10.1038/s41467-021-25332-w
R. Folberg, Z. Arbieva, J. Moses, A. Hayee, T. Sandal, S. Kadkol, A.Y. Lin, K. Valyi-Nagy, S. Setty, L. Leach, P. Chevez-Barrios, P. Larsen, D. Majumdar, J. Pe’er and, A.J. Maniotis, Am J Pathol 69, 1376–1389 (2006) https://doi.org/10.2353/ajpath.2006.060223
Z. Lin, D. Susskind, Cancers (Basel). 13 (2021). https://doi.org/10.3390/cancers13102464
R.D. Carvajal, J.J. Sacco, M.J. Jager, D.J. Eschelman, R. Olofsson Bagge, J.W. Harbour, N.D. Chieng, S.P. Patel, A.M. Joshua, S. Piperno-Neumann, Nat. Rev. Clin. Oncol. 20, 99–115 (2023). https://doi.org/10.1038/s41571-022-00714-1
W. Dai, S. Liu, S. Wang, L. Zhao, X. Yang, J. Zhou, Y. Wang, J. Zhang, P. Zhang, K. Ding, Y. Li, J. Pan, Signal. Transduct. Target. Ther. 6, 176 (2021). https://doi.org/10.1038/s41392-021-00563-x
J.A. Eble, S. Niland, Clin. Exp. Metastasis. 36, 171–198 (2019). https://doi.org/10.1007/s10585-019-09966-1
F. Gattazzo, A. Urciuolo, P. Bonaldo, Biochim. Biophys. Acta. 1840, 2506–2519 (2014). https://doi.org/10.1016/j.bbagen.2014.01.010
M. Berube, M. Talbot, C. Collin, C. Paquet-Bouchard, L. Germain, S.L. Guerin, E. Petitclerc, Int. J. Oncol. 26, 405–413 (2005)
J. Zhao, Q. Yi, K. Li, L. Chen, L. Dai, J. Feng, Y. Li, M. Zhou, J. Sun, Comput. Struct. Biotechnol. J. 20, 3182–3194 (2022). https://doi.org/10.1016/j.csbj.2022.06.034
J. Liu, J. Lu, W. Li, Front. Immunol. 12, 738068 (2021). https://doi.org/10.3389/fimmu.2021.738068
Acknowledgements
We are grateful to Prof. Hong Tu and Dr. Qian Li for their assistance in generating the liver metastatic mouse model. The work was supported by grants from National Natural Science Foundation of China (32170924), Science and Technology Commission of Shanghai Municipality (21140901600, 23ZR1438400), Shanghai Jiaotong University School of Medicine (02.101005.001.29.38 A), State Key Laboratory of Oncogenes and Related Genes (SB21-22, ZZ-21-13rcpy, ZZ-GCYJ-23-02, ZZ-94-2313), China Postdoctoral Science Foundation (2022M722115).
Author information
Authors and Affiliations
Contributions
J.L., S.S., Y.Z., A.Z. and D.X. designed and performed experiments; J.L., D.C., L.J., and Y.Z. analyzed the data; J.L. and D.X. wrote and edited the manuscript; D.X. supervised the study, obtained funding, and provided critical review. All authors approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Ethics approval
Animal experiments were performed in accordance with the recommendations in the Guide for the Care and Use of Laboratory Animals and relevant Chinese laws and regulations. The protocol was approved by the Institutional Animal Care and Use Committee (IACUC) of Shanghai Jiao Tong University, the Animal Protocol number is RJ2022-1010. Ethical approval was obtained from the Research Ethics Committee of Renji Hospital, School of Medicine, Shanghai Jiao Tong University.
Competing interests
The authors Dongxi Xiang, Jiaoduan Li, Lixin Jiang have submitted a patent (Application NO: 202310685260.6) related to this work. Dongxi Xiang is an editorial board member. Siyuan Shao is an employee of Shanghai OneTar Biomedicine.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, J., Cao, D., Jiang, L. et al. ITGB2-ICAM1 axis promotes liver metastasis in BAP1-mutated uveal melanoma with retained hypoxia and ECM signatures. Cell Oncol. (2023). https://doi.org/10.1007/s13402-023-00908-4
Accepted:
Published:
DOI: https://doi.org/10.1007/s13402-023-00908-4