Abstract
Purpose
Methylenetetrahydrofolate dehydrogenase (MTHFD1), a key enzyme on the folate pathway, has been implicated in the tumor development of distinct types of cancers. The single nucleotide polymorphism (SNP) of 1958G > A mutation in the coding region of MTHFD1 (arginine 653 is mutated into glutamine) has been detected in a significant proportion of clinical samples of hepatocellular carcinoma (HCC).
Methods
Hepatoma cell lines, 97H and Hep3B were used. The expression of MTHFD1 and SNP mutation protein was determined by immunoblotting analysis. The protein ubiquitination of MTHFD1 was detected by immunoprecipitation analysis. The post-translational modification sites and interacting proteins of MTHFD1 in the presence of G1958A SNP were identified by mass spectrometry. Metabolic flux analysis was used to detect the synthesis of relevant metabolites sourced from serine isotope.
Results
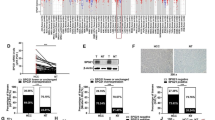
The present study showed G1958A SNP of MTHFD1, encoding MTHFD1 R653Q, was associated with the attenuated protein stability caused by ubiquitination-mediated protein degradation. Mechanistically, MTHFD1 R653Q displayed an enhanced binding to the E3 ligase TRIM21, which was responsible for the augmented ubiquitination, and MTHFD1 K504 was identified to be the primary ubiquitination site. The subsequent metabolite analysis revealed MTHFD1 R653Q resulted in the repressed flux of serine-derived methyl group into metabolite precursors for purine synthesis, and the compromised purine synthesis was demonstrated to be responsible for the impeded growth capability in MTHFD1 R653Q-expressing cells. Moreover, the suppressive effect of MTHFD1 R653Q expression in tumorigenesis was verified by xenograft analysis, and the relationship between MTHFD1 G1958A SNP and its protein levels was revealed in clinical human liver cancer specimens.
Conclusion
Our results uncovered an unidentified mechanism underlying of the impact of G1958A SNP on MTHFD1 protein stability and tumor metabolism in HCC. which provides a molecular basis for the according clinical management when considering MTHFD1 as a therapeutic target.






Similar content being viewed by others
Data availability
The original contributions presented in the study are included in the article. Further inquiries can be directed to the corresponding authors.
References
R.L. Siegel, K.D. Miller, H.E. Fuchs, A. Jemal, CA Cancer J. Clin 72 (2022). https://doi.org/10.3322/caac.21708
J.M. Llovet, F. Castet, M. Heikenwalder, M.K. Maini, V. Mazzaferro, D.J. Pinato, E. Pikarsky, A.X. Zhu, R.S. Finn, Nat. Rev. Clin. Oncol 19, 151–172 (2022). https://doi.org/10.1038/s41571-021-00573-2
J.M. Llovet, T. De Baere, L. Kulik, P.K. Haber, T.F. Greten, T. Meyer, R. Lencioni, Nat. Rev. Gastroenterol. Hepatol 18, 293–313 (2021). https://doi.org/10.1038/s41575-020-00395-0
H. Zhu, Y. Shan, K. Ge, J. Lu, W. Kong, C. Jia, Cell. Oncol. (Dordr) 43, 1203–1214 (2020). https://doi.org/10.1007/s13402-020-00552-2
R. Nilsson, M. Jain, N. Madhusudhan, N.G. Sheppard, L. Strittmatter, C. Kampf, J. Huang, A. Asplund, V.K. Mootha, Nat. Commun 5, 3128 (2014). https://doi.org/10.1038/ncomms4128
M. Shang, H. Yang, R. Yang, T. Chen, Y. Fu, Y. Li, X. Fang, K. Zhang, J. Zhang, H. Li, X. Cao, J. Gu, J. Xiao, Q. Zhang, X. Liu, Q. Yu, T. Wang, Nat. Commun 12, 1940 (2021). https://doi.org/10.1038/s41467-021-22173-5
P. Xie, J.-L. Mo, J.-H. Liu, X. Li, L.-M. Tan, W. Zhang, H.-H. Zhou, Z.-Q. Liu, Cell. Oncol. (Dordr) 43 (2020). https://doi.org/10.1007/s13402-020-00529-1
S.T. Pike, R. Rajendra, K. Artzt, D.R. Appling, J. Biol. Chem 285, 4612–4620 (2010). https://doi.org/10.1074/jbc.M109.079855
G.S. Ducker, L. Chen, R.J. Morscher, J.M. Ghergurovich, M. Esposito, X. Teng, Y. Kang, J.D. Rabinowitz, Cell. Metab 23, 1140–1153 (2016). https://doi.org/10.1016/j.cmet.2016.04.016
A.J. MacFarlane, C.A. Perry, H.H. Girnary, D. Gao, R.H. Allen, S.P. Stabler, B. Shane, P.J. Stover, J. Biol. Chem 284, 1533–1539 (2009). https://doi.org/10.1074/jbc.M808281200
N. Lévesque, K.E. Christensen, L. Van Der Kraak, A.F. Best, L. Deng, D. Caldwell, A.J. MacFarlane, N. Beauchemin, R. Rozen, Mol. Carcinog 56, 1030–1040 (2017). https://doi.org/10.1002/mc.22568
H. Yu, H. Wang, H.-R. Xu, Y.-C. Zhang, X.-B. Yu, M.-C. Wu, G.-Z. Jin, W.-M. Cong, Future Oncol 15, 1771–1780 (2019). https://doi.org/10.2217/fon-2018-0606
A. Parle-McDermott, P.N. Kirke, J.L. Mills, A.M. Molloy, C. Cox, V.B. O’Leary, F. Pangilinan, M. Conley, L. Cleary, L.C. Brody, J.M. Scott, Eur. J. Hum. Genet 14, 768–772 (2006)
K.E. Christensen, C.V. Rohlicek, G.U. Andelfinger, J. Michaud, J.-L. Bigras, A. Richter, R.E. Mackenzie, R. Rozen, Hum. Mutat 30, 212–220 (2009). https://doi.org/10.1002/humu.20830
S. Moruzzi, P. Guarini, S. Udali, A. Ruzzenente, A. Guglielmi, S. Conci, P. Pattini, N. Martinelli, O. Olivieri, S.A. Tammen, S.-W. Choi, S. Friso, PLoS ONE 12, e0185792 (2017). https://doi.org/10.1371/journal.pone.0185792
Y. Jiang, X. Qian, J. Shen, Y. Wang, X. Li, R. Liu, Y. Xia, Q. Chen, G. Peng, S.-Y. Lin, Z. Lu, Nat. Cell. Biol 17, 1158–1168 (2015). https://doi.org/10.1038/ncb3209
A.V. Kuznetsova, J. Meller, P.O. Schnell, J.A. Nash, M.L. Ignacak, Y. Sanchez, J.W. Conaway, R.C. Conaway, M.F. Czyzyk-Krzeska, Proc. Natl. Acad. Sci. U S A 100, 2706–2711 (2003)
P.J. Thul, C. Lindskog, Protein Sci 27, 233–244 (2018). https://doi.org/10.1002/pro.3307
Acknowledgements
We thank Core Facility of Basic Medical Sciences, Shanghai Jiao Tong University School of Medical for the technical assistance.
Funding
This work was supported by the Shanghai Yangfan Program (21YF1424900); Shanghai Organ Transplantation Research Center (2022ZZ01016); National Natural Science Foundation of China (81972205 and 92059205).
Author information
Authors and Affiliations
Contributions
This study was conceived by Q.X. and Y.J., K.R. and K.Z. designed the study. K.R. and K.Z. performed experiments. Q.Z., G.H., N.S., W.W., M.Y. B.Z., Y.Z. and Y.J. provided technical and material support. K.R. and J.H wrote the paper with comments from all authors.
Corresponding authors
Ethics declarations
Ethical approval
The animal study was reviewed and approved by School of Medicine, Shanghai Jiao Tong University. This study related to patients has been approved by the Institutional Review Board of Renji Hospital affiliated to Shanghai Jiao Tong University School of Medicine. This study related to patients has been approved by the Institutional Review Board of Renji Hospital affiliated to Shanghai Jiao Tong University School of Medicine and was performed in accordance with the ethical standards as laid down in the 1964 Declaration of Helsinki and its later amendments or comparable ethical standards.
Conflict of interest
The authors have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Rao, K., Zheng, K., Zhao, Q. et al. The negative effect of G1958A polymorphism on MTHFD1 protein stability and HCC growth. Cell Oncol. 46, 735–744 (2023). https://doi.org/10.1007/s13402-023-00780-2
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13402-023-00780-2