Abstract
Background
Although isocitrate dehydrogenase 2 (IDH2) mutations have been the hotspots in recent anticancer studies, the impact of wild-type IDH2 on cancer cell growth and metabolic alterations is still elusive.
Methods
IDH2 expression in CRC tissues was evaluated by immunohistochemistry, and the correlation between the expression level and the patient’s survival rate was analyzed. Cell functional assays included CCK8 and colony formation for cell proliferation in vitro and ectopic xenograft as in vivo experimental model for tumor progression. A targeted metabolomic procedure was performed by liquid chromatography/tandem mass spectrometry to profile the metabolites from glycolysis and tricarboxylic acid (TCA) cycle. Mitochondrial function was assessed by measuring cellular oxygen consumption (OCR) and mitochondrial membrane potential (ΔΨ). Confocal microscope analysis and Western blotting were applied to detect the expression of GLUT1 and NF-κB signaling. O-GlcNAcylation and the interaction of IDH2 with OGT were confirmed by co-immunoprecipitation, followed by Western blotting analysis.
Results
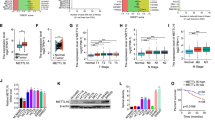
IDH2 protein was highly expressed in CRC tissues, and correlated with poor survival of CRC patients. Wild-type IDH2 promoted CRC cell growth in vitro and tumor progression in xenograft mice. Overexpression of wild-type IDH2 significantly increased glycolysis and TCA cycle metabolites, the ratios of NADH/NAD+ and ATP/ADP, OCR and mitochondrial membrane potential (ΔΨ) in CRC cells. Furthermore, α-KG activated NF-κB signaling to promote glucose uptake by upregulating GLUT1. Interesting, O-GlcNAcylation enhanced the protein half-time of IDH2 by inhibiting ubiquitin-mediated proteasome degradation. The O-GlcNAc transferase (OGT)-IDH2 axis promoted CRC progression.
Conclusion
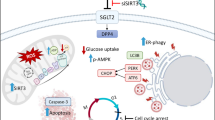
Wild-type IDH2 reprogrammed glucose metabolism and bioenergetic production via the NF-κB signaling pathway to promote CRC development and progression. O-GlcNAcylation of IDH2 elevated the stability of IDH2 protein. And the axis of OGT-IDH2 played an essential promotive role in tumor progression, suggesting a novel potential therapeutic strategy in CRC treatment.








Similar content being viewed by others
Data availability
Data and material are available in Supplementary Data.
Change history
17 December 2022
The original version of this article was revised: In this article Fig. 4d was incorrect; the figure should have appeared as shown below. The original article has been corrected.
27 December 2022
A Correction to this paper has been published: https://doi.org/10.1007/s13402-022-00764-8
Abbreviations
- CRC:
-
Colorectal cancer
- PPP:
-
Pentose phosphate pathway
- TCA:
-
Tricarboxylic acid cycle
- GLUT1:
-
Glucose transporter 1
- GLUT4:
-
Glucose transporter 4
- NF-κB:
-
Nuclear factor kappa-B
- TCA cycle:
-
Tricarboxylic acid cycle
- OXPHOS:
-
Oxidative phosphorylation
- OCR:
-
Oxygen consumption ratio
- PTMs:
-
Post-translational modifications
- IDH2:
-
Isocitrate dehydrogenase 2
- α-KG:
-
Alpha-ketoglutarate
- O-GlcNAc:
-
O-Linked β-N-acetylglucosamine
- OGT:
-
O-GlcNAc transferase
- OGA:
-
O-GlcNAcase
- ROS:
-
Reactive oxygen species
References
J. Ferlay, I. Soerjomataram, R. Dikshit, S. Eser, C. Mathers, M. Rebelo, D.M. Parkin, D. Forman, F. Bray, Int. J. Cancer 136, E359–386 (2015). https://doi.org/10.1002/ijc.29210
L.A. Torre, F. Bray, R.L. Siegel, J. Ferlay, J. Lortet-Tieulent, A. Jemal, CA Cancer J. Clin. 65, 87–108 (2015). https://doi.org/10.3322/caac.21262
R.L. Siegel, K.D. Miller, A. Goding Sauer, S.A. Fedewa, L.F. Butterly, J.C. Anderson, A. Cercek, R.A. Smith, A. Jemal, CA Cancer J. Clin. 70, 145–164 (2020). https://doi.org/10.3322/caac.21601
S.N. Hong, Intest. Res. 16, 327–337 (2018). https://doi.org/10.5217/ir.2018.16.3.327
S. La Vecchia, C. Sebastian, Semin. Cell Dev. Biol. (2019). https://doi.org/10.1016/j.semcdb.2019.05.018
H. Wang, L. Wang, H. Zhang, P. Deng, J. Chen, B. Zhou, J. Hu, J. Zou, W. Lu, P. Xiang, T. Wu, X. Shao, Y. Li, Z. Zhou, Y.L. Zhao, Mol. Cancer 12, 121 (2013). https://doi.org/10.1186/1476-4598-12-121
V. Chekulayev, K. Mado, I. Shevchuk, A. Koit, A. Kaldma, A. Klepinin, N. Timohhina, K. Tepp, M. Kandashvili, L. Ounpuu, K. Heck, L. Truu, A. Planken, V. Valvere, T. Kaambre, Biochem. Biophys. Rep. 4, 111–125 (2015). https://doi.org/10.1016/j.bbrep.2015.08.020
J. Ding, Q. Gou, X. Jia, Q. Liu, J. Jin, J. Shi, Y. Hou, J. Biol. Chem. 297, >100954 (2021). https://doi.org/10.1016/j.jbc.2021.100954
M. Moretti, J. Bennett, L. Tornatore, A.K. Thotakura, G. Franzoso, Int. J. Biochem. Cell Biol. 44, 2238–2243 (2012). https://doi.org/10.1016/j.biocel.2012.08.002
P.S. Ward, C.B. Thompson, Cancer Cell 21, 297–308 (2012). https://doi.org/10.1016/j.ccr.2012.02.014
T. Fujii, M.R. Khawaja, C.D. DiNardo, J.T. Atkins, F. Janku, Discov. Med. 21, 373–380 (2016)
K. Smolkova, P. Jezek, Int. J. Cell. Biol. 2012, 273947 (2012). https://doi.org/10.1155/2012/273947
W. Xiao, R.S. Wang, D.E. Handy, J. Loscalzo, Antioxid. Redox Signal 28, 251–272 (2018). https://doi.org/10.1089/ars.2017.7216
H.Q. Ju, J.F. Lin, T. Tian, D. Xie, R.H. Xu, Signal Transduct. Target Ther. 5, 231 (2020). https://doi.org/10.1038/s41392-020-00326-0
N.R. Rose, M.A. McDonough, O.N. King, A. Kawamura, C.J. Schofield, Chem. Soc. Rev. 40, 4364–4397 (2011). https://doi.org/10.1039/c0cs00203h
L. Jin, J. Chun, C. Pan, A. Kumar, G. Zhang, Y. Ha, D. Li, G.N. Alesi, Y. Kang, L. Zhou, W.M. Yu, K.R. Magliocca, F.R. Khuri, C.K. Qu, C. Metallo, T.K. Owonikoko and S. Kang, Mol. Cell 69, 87–99 e87 (2018). https://doi.org/10.1016/j.molcel.2017.11.025
X. Wang, R. Liu, X. Qu, H. Yu, H. Chu, Y. Zhang, W. Zhu, X. Wu, H. Gao, B. Tao, W. Li, J. Liang, G. Li, W. Yang, Mol. Cell 76, 148–162 e147 (2019). https://doi.org/10.1016/j.molcel.2019.07.007
T.Q. Tran, E.A. Hanse, A.N. Habowski, H. Li, M.B.I. Gabra, Y. Yang, X.H. Lowman, A.M. Ooi, S.Y. Liao, R.A. Edwards, M.L. Waterman, M. Kong, Nat. Cancer 1, 345–358 (2020). https://doi.org/10.1038/s43018-020-0035-5
Z.J. Reitman, H. Yan, Jnci-J Natl. Cancer I 102, 932–941 (2010). https://doi.org/10.1093/jnci/djq187
H. Yang, D. Ye, K.L. Guan, Y. Xiong, Clin. Cancer Res. 18, 5562–5571 (2012). https://doi.org/10.1158/1078-0432.CCR-12-1773
C.J. Pirozzi, H. Yan, Nat. Rev. Clin. Oncol. 18, 645–661 (2021). https://doi.org/10.1038/s41571-021-00521-0
J.S. Becker, A.T. Fathi, Curr. Cancer Drug Targets 20, 490–500 (2020). https://doi.org/10.2174/1568009620666200424145622
M.R. Kang, M.S. Kim, J.E. Oh, Y.R. Kim, S.Y. Song, S.I. Seo, J.Y. Lee, N.J. Yon, S.H. Lee, Int. J. Cancer 125, 353–355 (2009). https://doi.org/10.1002/ijc.24379
W.L. Li, M.S. Xiao, D.F. Zhang, D. Yu, R.X. Yang, X.Y. Li, Y.G. Yao, Gene 546, 263–270 (2014). https://doi.org/10.1016/j.gene.2014.05.070
D. Tougeron, K. Guilloteau, L. Karayan-Tapon, Dig. Liver Dis. 48, 681–683 (2016). https://doi.org/10.1016/j.dld.2016.02.019
S. Qiao, W. Lu, C. Glorieux, J. Li, P. Zeng, N. Meng, H. Zhang, S. Wen, P. Huang, Oncogene 40, 5880–5892 (2021). https://doi.org/10.1038/s41388-021-01968-2
E. Bergaggio, R. Piva, Cancers (Basel) 11, (2019). https://doi.org/10.3390/cancers11040563
J. Ma, C. Wu, G.W. Hart, Chem. Rev. 121, 1513–1581 (2021). https://doi.org/10.1021/acs.chemrev.0c00884
J.C. Chatham, J. Zhang, A.R. Wende, Physiol. Rev. 101, 427–493 (2021). https://doi.org/10.1152/physrev.00043.2019
P. Jozwiak, E. Forma, M. Brys, A. Krzeslak, Front. Endocrinol. (Lausanne) 5, 145 (2014). https://doi.org/10.3389/fendo.2014.00145
J.P. Singh, K. Zhang, J. Wu, X. Yang, Cancer Lett. 356, 244–250 (2015). https://doi.org/10.1016/j.canlet.2014.04.014
Y. Fardini, V. Dehennaut, T. Lefebvre, T. Issad, Front. Endocrinol. (Lausanne) 4, 99 (2013). https://doi.org/10.3389/fendo.2013.00099
H.B. Ruan, J.P. Singh, M.D. Li, J. Wu, X. Yang, Trends Endocrinol. Metab. 24, 301–309 (2013). https://doi.org/10.1016/j.tem.2013.02.002
M. Jiang, B. Xu, X. Li, Y. Shang, Y. Chu, W. Wang, D. Chen, N. Wu, S. Hu, S. Zhang, M. Li, K. Wu, X. Yang, J. Liang, Y. Nie, D. Fan, Oncogene 38, 301–316 (2019). https://doi.org/10.1038/s41388-018-0435-5
D. Xu, W. Wang, T. Bian, W. Yang, M. Shao, H. Yang, Int. J. Clin. Exp. Pathol. 12, 1305–1314 (2019)
X. Lu, P. Yang, X. Zhao, M. Jiang, S. Hu, Y. Ouyang, L. Zeng, J. Wu, Exp. Cell Res. 382, 111483 (2019). https://doi.org/10.1016/j.yexcr.2019.06.028
X. Lu, N. Wu, W. Yang, J. Sun, K. Yan, J. Wu, Onco. Targets Ther. 12, 7489–7500 (2019). https://doi.org/10.2147/OTT.S208848
J. Wu, J. Liu, K. Lapenta, R. Desrouleaux, M.D. Li, X. Yang, J. Mol. Cell Biol. (2022). https://doi.org/10.1093/jmcb/mjac016
R.J. Molenaar, J.P. Maciejewski, J.W. Wilmink, C.J.F. van Noorden, Oncogene 37, 1949–1960 (2018). https://doi.org/10.1038/s41388-017-0077-z
N. Wu, M. Jiang, Y. Han, H. Liu, Y. Chu, H. Liu, J. Cao, Q. Hou, Y. Zhao, B. Xu, X. Xie, J. Cell Mol. Med. 23, 1354–1362 (2019)
H.B. Ruan, Y. Nie, X. Yang, Mol. Cell. Proteomics 12, 3489–3497 (2013). https://doi.org/10.1074/mcp.R113.029751
J.N. Moloney, T.G. Cotter, Semin. Cell Dev. Biol. 80, 50–64 (2018). https://doi.org/10.1016/j.semcdb.2017.05.023
M. Jiang, B. Xu, X. Li, Y. Shang, Y. Chu, W. Wang, D. Chen, N. Wu, S. Hu, S. Zhang, M. Li, Oncogene 38, 301–316 (2019)
J. Huang, L.H. Tseng, V. Parini, P.M. Lokhandwala, A. Pallavajjala, E. Rodriguez, R. Xian, L. Chen, C.D. Gocke, J.R. Eshleman, M.T. Lin, Am. J. Clin. Pathol. 156, 777–786 (2021). https://doi.org/10.1093/ajcp/aqab023
D. Shen, J. Zhang, K. Yuan, J. Zhao, Z. Zhao, L. Cui, Y. Zhang, G. Wang, S. Cai, Y. Bai, W. Li, X. Huang, Mol. Genet. Genomic. Med. 9 e1697 (2021). https://doi.org/10.1002/mgg3.1697
T. Sjoblom, S. Jones, L.D. Wood, D.W. Parsons, J. Lin, T.D. Barber, D. Mandelker, R.J. Leary, J. Ptak, N. Silliman, S. Szabo, P. Buckhaults, C. Farrell, P. Meeh, S.D. Markowitz, J. Willis, D. Dawson, J.K. Willson, A.F. Gazdar, J. Hartigan, L. Wu, C. Liu, G. Parmigiani, B.H. Park, K.E. Bachman, N. Papadopoulos, B. Vogelstein, K.W. Kinzler, V.E. Velculescu, Science 314, 268–274 (2006). https://doi.org/10.1126/science.1133427
H.M. Yusof, S. Ab-Rahim, L.S. Suddin, M.S.A. Saman, M. Mazlan, Malays. J. Med. Sci. 25, 16–34 (2018). https://doi.org/10.21315/mjms2018.25.5.3
K. Satoh, S. Yachida, M. Sugimoto, M. Oshima, T. Nakagawa, S. Akamoto, S. Tabata, K. Saitoh, K. Kato, S. Sato, K. Igarashi, Y. Aizawa, R. Kajino-Sakamoto, Y. Kojima, T. Fujishita, A. Enomoto, A. Hirayama, T. Ishikawa, M.M. Taketo, Y. Kushida, R. Haba, K. Okano, M. Tomita, Y. Suzuki, S. Fukuda, M. Aoki, T. Soga, Proc. Natl. Acad. Sci USA 114, E7697–E7706 (2017). https://doi.org/10.1073/pnas.1710366114
R.E. Brown, S.P. Short, C.S. Williams, Curr. Colorectal Cancer Rep. 14, 226–241 (2018). https://doi.org/10.1007/s11888-018-0420-y
L. Xue, X. Liu, Q. Wang, C.Q. Liu, Y. Chen, W. Jia, R. Hsie, Y. Chen, F. Luh, S. Zheng, Y. Yen, Am. J. Transl. Res. 10, 3635–3649 (2018)
A.I. Aljohani, M.S. Toss, S. Kurozumi, C. Joseph, M.A. Aleskandarany, I.M. Miligy, R.E. Ansari, N.P. Mongan, I.O. Ellis, A.R. Green, E.A. Rakha, Breast Cancer Res. Treat. 179, 79–90 (2020). https://doi.org/10.1007/s10549-019-05459-7
A. Kaldma, A. Klepinin, V. Chekulayev, K. Mado, I. Shevchuk, N. Timohhina, K. Tepp, M. Kandashvili, M. Varikmaa, A. Koit, M. Planken, K. Heck, L. Truu, A. Planken, V. Valvere, E. Rebane, T. Kaambre, Int. J. Biochem. Cell Biol. 55, 171–186 (2014). https://doi.org/10.1016/j.biocel.2014.09.004
I. Martinez-Reyes, N.S. Chandel, Nat. Commun. 11, 102 (2020). https://doi.org/10.1038/s41467-019-13668-3
T.Q. Tran, E.A. Hanse, A.N. Habowski, H. Li, M.B. Ishak Gabra, Y. Yang, X.H. Lowman, A.M. Ooi, S.Y. Liao, R.A. Edwards, M.L. Waterman and M. Kong, Nat. Cancer 1, 345–358 (2020). https://doi.org/10.1038/s43018-020-0035-5
L. Yang, S. Venneti, D. Nagrath, Annu. Rev. Biomed. Eng. 19, 163–194 (2017). https://doi.org/10.1146/annurev-bioeng-071516-044546
W. Yu, K.E. Dittenhafer-Reed, J.M. Denu, J. Biol. Chem. 287, 14078–14086 (2012). https://doi.org/10.1074/jbc.M112.355206
L. Zhou, F. Wang, R. Sun, X. Chen, M. Zhang, Q. Xu, Y. Wang, S. Wang, Y. Xiong, K.L. Guan, P. Yang, H. Yu, D. Ye, EMBO Rep. 17, 811–822 (2016). https://doi.org/10.15252/embr.201541643
Y. Xu, L. Liu, A. Nakamura, S. Someya, T. Miyakawa, M. Tanokura, Sci. Rep. 7, 9785 (2017). https://doi.org/10.1038/s41598-017-10337-7
Q. Ong, W. Han, X. Yang, Front. Endocrinol. (Lausanne) 9, 599 (2018). https://doi.org/10.3389/fendo.2018.00599
Y. Hu, J. Suarez, E. Fricovsky, H. Wang, B.T. Scott, S.A. Trauger, W. Han, Y. Hu, M.O. Oyeleye, W.H. Dillmann, J. Biol. Chem. 284, 547–555 (2009). https://doi.org/10.1074/jbc.M808518200
J. Ma, T. Liu, A.C. Wei, P. Banerjee, B. O’Rourke, G.W. Hart, J. Biol. Chem. 290, 29141–29153 (2015). https://doi.org/10.1074/jbc.M115.691741
R. Trapannone, D. Mariappa, A.T. Ferenbach, D.M. van Aalten, Biochem. J. 473, 1693–1702 (2016). https://doi.org/10.1042/BCJ20160092
P. Jozwiak, P. Ciesielski, P.K. Zakrzewski, K. Kozal, J. Oracz, G. Budryn, D. Zyzelewicz, S. Flament, A.S. Vercoutter-Edouart, F. Bray, T. Lefebvre and A. Krzeslak, Cancers (Basel) 13, (2021). https://doi.org/10.3390/cancers13122956
G.W. Hart, C. Slawson, G. Ramirez-Correa, O. Lagerlof, Annu. Rev. Biochem. 80, 825–858 (2011). https://doi.org/10.1146/annurev-biochem-060608-102511
Z. Ma, R.J. Chalkley, K. Vosseller, J. Biol. Chem. 292, 9150–9163 (2017). https://doi.org/10.1074/jbc.M116.766568
Y.R. Yang, D.H. Kim, Y.K. Seo, D. Park, H.J. Jang, S.Y. Choi, Y.H. Lee, G.H. Lee, K. Nakajima, N. Taniguchi, J.M. Kim, E.J. Choi, H.Y. Moon, I.S. Kim, J.H. Choi, H. Lee, S.H. Ryu, L. Cocco, P.G. Suh, Oncotarget 6, 12529–12542 (2015). https://doi.org/10.18632/oncotarget.3725
H. Nie, H. Ju, J. Fan, X. Shi, Y. Cheng, X. Cang, Z. Zheng, X. Duan, W. Yi, Nat. Commun. 11, 36 (2020). https://doi.org/10.1038/s41467-019-13601-8
L. Liu, S. Shah, J. Fan, J.O. Park, K.E. Wellen, J.D. Rabinowitz, Nat. Chem. Biol. 12, 345–352 (2016). https://doi.org/10.1038/nchembio.2047
Funding
This work was supported by the National Natural Science Foundation of China (No.81471401), the Natural Science Foundation of Shaanxi Provincial of China (No.2019JZ-41) and the Opening Foundation of State Key Laboratory of Cancer Biology, National Clinical Research Center for Digestive Diseases and Xijing Hospital of Digestive Diseases, Air Force Medical University.
Author information
Authors and Affiliations
Contributions
Jing Wu and Yongzhan Nie wrote the main manuscript text. Xiaoli He and Nan Wu prepared Figs. 1–6 and Fig. 8. Renlong Li and Haohao zhang prepared the Fig. 7 and supplementary Fig S1. Yu zhao did the MASS analysis. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Ethical Approval and Consent to participate
The CRC tissue microarrays were purchased from Shanghai Outdo Biotech Co. LTD. (Shanghai, P.R.C). The mice study received approval from the institutional review board of the Air Force Medical University.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests as defined by Springer, or other interests that might be perceived to influence the results and/or discussion reported in this paper.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original version of this article was revised: In this article Fig. 4d was incorrect; the figure should have appeared as shown below. The original article has been corrected.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
He, X., Wu, N., Li, R. et al. IDH2, a novel target of OGT, facilitates glucose uptake and cellular bioenergy production via NF-κB signaling to promote colorectal cancer progression. Cell Oncol. 46, 145–164 (2023). https://doi.org/10.1007/s13402-022-00740-2
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13402-022-00740-2