Abstract
Background
Tibetan hulless barley (Hordeum vulgare var. nudum), adjusting to the harsh environment on Qinghai–Tibet Plateau, is a good subject for analyzing drought tolerance mechanism. Several unannotated differentially expressed genes (DEGs) were identified through our previous RNA-Seq study using two hulless barley accessions with contrasting drought tolerance. One of these DEGs, HVU010048.2, showed up-regulated pattern under dehydration stress in both drought tolerant (DT) and drought susceptible (DS) accessions, while its function in drought resistance remains unknown. This new gene was named as HvLRX (light responsive X), because its expression was induced under high light intensity while suppressed under dark.
Objective
To provide preliminary bioinformatics prediction, expression pattern, and drought resistance function of this new gene.
Methods
Bioinformatics analysis of HvLRX were conducted by MEGA, PlantCARE, ProtParam, CELLO et al. The expression pattern of HvLRX under different light intensity, dehydration shock, gradual drought stress, NaCl stress, polyethylene glycol (PEG) 6000 stress and abscisic acid (ABA) treatment was investigated by quantitative reverse transcription-polymerase chain reaction (RT-qPCR). The function of HvLRX was analyzed by virus induced gene silencing (VIGS) in hulless barley and by transgenic method in tobacco.
Results
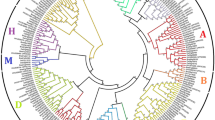
Full cDNAs of HvLRX were cloned and compared in three hulless barley accessions. Homologues of HvLRX protein in other plants were excavated and their phylogenetic relationship was analyzed. Several light responsive elements (ATC-motif, Box 4, G-box, Sp1, and chs-CMA1a) were identified in its promoter region. Its expression can be promoted under high light intensity, dehydration shock, gradual drought stress, PEG 6000, and NaCl stress, but was almost unchanged in ABA treatment. HvLRX-silenced plants had a higher leaf water loss rate (WLR) and a lower survival rate (SR) compared with controls under dehydration stress. The infected leaves of HvLRX-silenced plants lost their water content quickly and became withered at 10 dpi. The SR of HvLRX overexpressed transgenic tobacco plants was significantly higher than that of wild-type plants. These results indicated HvLRX play a role in drought resistance. Besides, retarded vegetative growth was detected in HvLRX-silenced hulless barley plants, which suggested that this gene is important for plant development.
Conclusions
This study provided data of bioinformatics, expression pattern, and function of HvLRX. To our knowledge, this is the first report of this new dehydration and light responsive gene.








Similar content being viewed by others
Abbreviations
- ABA:
-
Abscisic acid
- ANOVA:
-
Analysis of variance
- BPs:
-
Bootstrap probabilities
- BSMV:
-
Barley stripe mosaic virus
- DEGs:
-
Differentially expressed genes
- DPI:
-
Days post inoculation
- DT:
-
Drought tolerant
- DS:
-
Drought susceptible
- DW:
-
Dry weight
- EF:
-
Elongation factor
- FW:
-
Fresh weight
- GFP:
-
Green fluorescent protein
- LSD:
-
Least significant differences
- MS:
-
Murashige and Skoog
- NCBI:
-
National Center for Biotechnology Information
- PEG:
-
Polyethylene glycol
- PPDK:
-
Pyruvate phosphate dikinase
- RT-qPCR:
-
Quantitative reverse transcription-polymerase chain reaction
- SD:
-
Standard deviation
- SR:
-
Survival rate
- RT-PCR:
-
Reverse transcription-polymerase chain reaction
- TDW:
-
Total dry weight
- VIGS:
-
Virus induced gene silencing
- WC:
-
Water content
- WLR:
-
Water loss rate
- WT:
-
Wild type
References
Ahmed F, Senthil-Kumar M, Dai X, Ramu VS, Lee S, Mysore KS, Zhao PX (2020) pssRNAit: a web server for designing effective and specific plant siRNAs with genome-wide off-target assessment. Plant Physiol 184(1):65–81
Almagro Armenteros JJ, Tsirigos KD, Sønderby CK, Petersen TN, Winther O, Brunak S, von Heijne G, Nielsen H (2019) SignalP 5.0 improves signal peptide predictions using deep neural networks. Nat Biotechnol 37(4):420–423
Aprile A, Mastrangelo AM, De Leonardis AM, Galiba G, Roncaglia E, Ferrari F, De Bellis L, Turchi L, Giuliano G, Cattivelli L (2009) Transcriptional profiling in response to terminal drought stress reveals differential responses along the wheat genome. BMC Genomics 10:279
Bedada G, Westerbergh A, Müller T, Galkin E, Bdolach E, Moshelion M, Fridman E, Schmid KJ (2014) Transcriptome sequencing of two wild barley (Hordeum spontaneum L.) ecotypes differentially adapted to drought stress reveals ecotypespecific transcripts. BMC Genomics 15(1):995
Boyer JS (1982) Plant productivity and environment. Science 218(4571):443–448
Dobson L, Reményi I, Tusnády GE (2015) CCTOP: a consensus constrained TOPology prediction web server. Nucleic Acids Res 43(W1):W408–W412
Garnier J, Gibrat JF, Robson B (1996) GOR method for predicting protein secondary structure from amino acid sequence. Methods Enzymol 266:540–553
Gasteiger E, Hoogland C, Gattiker A, Duvaud S, Wilkins MR, Appel RD, Bairoch A (2005) Protein identification and analysis tools on the ExPASy server. In: JohnM W (ed) The proteomics protocols handbook. Humana Press, Totowa, New Jersey, pp 571–607
Haupt S, Duncan GH, Holzberg S, Oparka KJ (2001) Evidence for symplastic phloem unloading in sink leaves of barley. Plant Physiol 125(1):209–218
He L, Bian J, Xu J, Yang K (2019) Novel maize NAC transcriptional repressor ZmNAC071 confers enhanced sensitivity to ABA and osmotic stress by downregulating stress-responsive genes in transgenic Arabidopsis. J Agric Food Chem 67(32):8905–8918
Huang L, Yan H, Jiang X, Yin G, Zhang X, Qi X, Zhang Y, Yan Y, Ma X, Peng Y (2014) Identification of candidate reference genes in perennial ryegrass for quantitative RT-PCR under various abiotic stress conditions. PLoS ONE 9(4):e93724
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33(7):1870–1874
Kumazaki A, Suzuki N (2019) Enhanced tolerance to a combination of heat stress and drought in Arabidopsis plants deficient in ICS1 is associated with modulation of photosynthetic reaction center proteins. Physiol Plant 165(2):232–246
Kundu A, Patel A, Pal A (2013) Defining reference genes for qPCR normalization to study biotic and abiotic stress responses in Vigna mungo. Plant Cell Rep 32(10):1647–1658
Lescot M, Déhais P, Thijs G, Marchal K, Moreau Y, Van de Peer Y, Rouzé P, Rombauts S (2002) PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res 30(1):325–327
Lesk C, Rowhani P, Ramankutty N (2016) Influence of extreme weather disasters on global crop production. Nature 529(7584):84–87
Li H, Li M, Wei X, Zhang X, Xue R, Zhao Y, Zhao H (2017) Transcriptome analysis of drought-responsive genes regulated by hydrogen sulfide in wheat (Triticum aestivum L.) leaves. Mol Genet Genomics 292(5):1091–1110
Liang JJ, Deng GB, Long H, Pan ZF, Wang CP, Cai P, Xu DL, Nima ZX, Yu MQ (2012) Virus-induced silencing of genes encoding LEA protein in Tibetan hulless barley (Hordeum vulgare ssp. vulgare) and their relationship to drought tolerance. Mol Breed 30(1):441–451
Liang JJ, Chen X, Deng GB, Pan ZF, Zhang HL, Li Q, Yang KJ, Long H, Yu MQ (2017) Dehydration induced transcriptomic responses in two Tibetan hulless barley (Hordeum vulgare var. nudum) accessions distinguished by drought tolerance. BMC Genomics 18(1):775
Lim CW, Baek W, Lee SC (2018) Roles of pepper bZIP protein CaDILZ1 and its interacting partner RING-type E3 ligase CaDSR1 in modulation of drought tolerance. Plant J 96(2):452–467
Liu Z, Xin M, Qin J, Peng H, Ni Z, Yao Y, Sun Q (2015) Temporal transcriptome profiling reveals expression partitioning of homeologous genes contributing to heat and drought acclimation in wheat (Triticum aestivum L.). BMC Plant Biol 15:152
Liu J, Fan Y, Zou J, Fang Y, Wang L, Wang M, Jiang X, Liu Y, Gao J, Zhang C (2017) A RhABF2/Ferritin module affects rose (Rosa hybrida) petal dehydration tolerance and senescence by modulating iron levels. Plant J 92(6):1157–1169
Liu J, Chen X, Wang S, Wang Y, Ouyang Y, Yao Y, Li R, Fu S, Hu X, Guo J (2019) MeABL5, an ABA insensitive 5-like basic leucine zipper transcription factor, positively regulates MeCWINV3 in Cassava (Manihot esculenta Crantz). Front Plant Sci 10:772
Lu L, Dong C, Liu R, Zhou B, Wang C, Shou H (2018) Roles of soybean plasma membrane intrinsic protein GmPIP2;9 in drought tolerance and seed development. Front Plant Sci 9:530
Ma J, Li R, Wang H, Li D, Wang X, Zhang Y, Zhen W, Duan H, Yan G, Li Y (2017) Transcriptomics analyses reveal wheat responses to drought stress during reproductive stages under field conditions. Front Plant Sci 8:592
Marcos FCC, Silveira NM, Mokochinski JB, Sawaya ACHF, Marchiori PER, Machado EC, Souza GM, Landell MGA, Ribeiro RV (2018) Drought tolerance of sugarcane is improved by previous exposure to water deficit. J Plant Physiol 223:9–18
Matsumoto T, Tanaka T, Sakai H, Amano N, Kanamori H, Kurita K, Kikuta A, Kamiya K, Yamamoto M, Ikawa H, Fujii N, Hori K, Itoh T, Sato K (2011) Comprehensive sequence analysis of 24,783 barley full-length cDNAs derived from 12 clone libraries. Plant Physiol 156(1):20–28
Munné-Bosch S, Müller M (2013) Hormonal cross-talk in plant development and stress responses. Front Plant Sci 4:529
Pan ZF, Deng GB, Zhai XG, Wu F, Yu MQ (2007) Genetic Diversity of acid-PAGE monomeric prolamins in cultivated hulless barley (Hordeum vulgare L.) from Qinghai-Tibet plateau in China. Genet Resour Crop Evol 54(8):1691–1699
Qian G, Han ZX, Zhao T, Deng GB, Pan ZF, Yu MQ (2007) Genotypic variability in sequence and expression of HVA1 gene in Tibetan hulless barley, Hordeum vulgare ssp. vulgare, associated with resistance to water deficit. Aust J Agric Res 58(5):425–431
Ristic Z, Jenks MA (2002) Leaf cuticle and water loss in maize lines differing in dehydration avoidance. J Plant Physiol 159(6):645–651
Rosche E, Westhoff P (1995) Genomic structure and expression of the pyruvate, orthophosphate dikinase gene of the dicotyledonous C4 plant Flaveria trinervia (Asteraceae). Plant Mol Biol 29(4):663–678
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4(4):406–425
Sasaki K, Ida Y, Kitajima S, Kawazu T, Hibino T, Hanba YT (2019) Overexpressing the HD-Zip class II transcription factor EcHB1 from Eucalyptus camaldulensis increased the leaf photosynthesis and drought tolerance of Eucalyptus. Sci Rep 9(1):14121
Sato H, Takasaki H, Takahashi F, Suzuki T, Iuchi S, Mitsuda N, Ohme-Takagi M, Ikeda M, Seo M, Yamaguchi-Shinozaki K, Shinozaki K (2018) Arabidopsis thaliana NGATHA1 transcription factor induces ABA biosynthesis by activating NCED3 gene during dehydration stress. PNAS 115(47):E11178–E11187
Shivhare R, Lata C (2016) Selection of suitable reference genes for assessing gene expression in pearl millet under different abiotic stresses and their combinations. Sci Rep 6:23036
Szklarczyk D, Gable AL, Lyon D, Junge A, Wyder S, Huerta-Cepas J, Simonovic M, Doncheva NT, Morris JH, Bork P, Jensen LJ, Mering CV (2019) STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res 47(D1):D607–D613
Tester M, Langridge P (2010) Breeding technologies to increase crop production in a changing world. Science 327(5967):818–822
Wang J, Abbas M, Wen Y, Niu D, Wang L, Sun Y, Li Y (2018) Selection and validation of reference genes for quantitative gene expression analyses in black locust (Robinia pseudoacacia L.) using real-time quantitative PCR. PLoS ONE 13(3):0193076
Wang HY, Liu C, Ma PA, Lu C, Li KM, Wang WQ (2018) Functional characterization of cytosolic pyruvate phosphate dikinase gene (MecyPPDK) and Promoter (MecyPPDKP) of cassava in response to abiotic stress in transgenic tobacco. Crop Sci 58(5):2002–2009
Wang J, Liu S, Liu H, Chen K, Zhang P (2019) PnSAG1, an E3 ubiquitin ligase of the Antarctic moss Pohlia nutans, enhanced sensitivity to salt stress and ABA. Plant Physiol Biochem 141:343–352
Xie T, Gu W, Zhang L, Li L, Qu D, Li C, Meng Y, Li J, Wei S, Li W (2018) Modulating the antioxidant system by exogenous 2-(3,4-dichlorophenoxy) triethylamine in maize seedlings exposed to polyethylene glycol-simulated drought stress. PLoS ONE 13(9):e0203626
Yadav S, Mishra A (2020) Ectopic expression of C4 photosynthetic pathway genes improves carbon assimilation and alleviate stress tolerance for future climate change. Physiol Mol Biol Plants 26(2):195–209
Yang Z, Dai Z, Lu R, Wu B, Tang Q, Xu Y, Cheng C, Su J (2017) Transcriptome analysis of two species of jute in response to polyethylene glycol (PEG)-induced drought stress. Sci Rep 7(1):16565
Yu CS, Chen YC, Lu CH, Hwang JK (2006) Prediction of protein subcellular localization. Proteins Struct Funct Bioinform 64(3):643–651
Zeng X, Long H, Wang Z, Zhao S, Tang Y, Huang Z, Wang Y, Xu Q, Mao L, Deng G et al (2015) The draft genome of Tibetan hulless barley reveals adaptive patterns to the high stressful Tibetan Plateau. PNAS 112(4):1095–1100
Zeng XQ, Bai LJ, Wei ZX, Yuan HJ, Wang YL, Xu QJ, Tang YW, Tashi N (2016) Transcriptome analysis revealed the drought-responsive genes in Tibetan hulless barley. BMC Genomics 17:386
Zhang S, Zeng Y, Yi X, Zhang Y (2016) Selection of suitable reference genes for quantitative RT-PCR normalization in the halophyte Halostachys caspica under salt and drought stress. Sci Rep 6:30363
Zhang X, Pu P, Tang Y, Zhang LX, Lv JY (2019) C4 photosynthetic enzymes play a key role in wheat spike bracts primary carbon metabolism response under water deficit. Plant Physiol Biochem 142:163–172
Zhao MJ, Yin LJ, Ma J, Zheng JC, Wang YX, Lan JH, Fu JD, Chen M, Xu ZS, Ma YZ (2019) The roles of GmERF135 in improving salt tolerance and decreasing ABA sensitivity in soybean. Front Plant Sci 10:940
Zhou HB, Li SF, Deng ZY, Wang XP, Chen T, Zhang JS, Chen SY, Ling HQ, Zhang AM, Wang DW, Zhang XQ (2007) Molecular analysis of three new receptor-like kinase genes from hexaploid wheat and evidence for their participation in wheat hypersensitive response to stripe rust fungus infection. Plant J 52(3):420–434
Zhu JK (2002) Salt and drought stress signal transduction in plants. Annu Rev Plant Biol 53:247–273
Zhu JK (2016) Abiotic stress signaling and responses in plants. Cell 167(2):313–324
Acknowledgements
Prof. Daowen Wang of Institute of Genetics and Developmental Biology, Chinese Academy of Sciences, Beijing, are kindly acknowledged for providing BSMV vectors.
Funding
This work was financially supported by Applied Basic Research Program of Sichuan Province (2019YJ0011); Science and Technology Service Network Initiative (KFJ-STS-QYZD-2021–22-001) by Chinese Academy of Sciences; Major Tibet Science and Technology Projects (XZ2021NA01) by Science and Technology Department of Tibet; Innovation Team of Triticeae Crops of Sichuan Province by Sichuan Provincial Department of Agriculture and Rural Affairs. All the funding bodies did not participate in the design of the study, collection, analysis, interpretation of data, or in writing the manuscript.
Author information
Authors and Affiliations
Contributions
JL and GD conceived and designed the experiments; JL, HZ and LY conducted the experiments; JL and HZ analyzed the data; JL and HL wrote the manuscript; MY and YT revised the manuscript. All authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
13258_2021_1147_MOESM1_ESM.xlsx
Fig. S1: Map of vector pJG081. Fig. S2: Nucleotide sequences alignment of HvLRX and AK353794.1. Fig. S3: Amino acid sequences alignment of HvLRX and AK353794.1. Fig. S4: Nucleotide sequences alignment of HvLRX, TaLRX-2A, TaLRX-2B, and TaLRX-2D. Fig. S5: Protein domain of HvLRX analyzed by Pfam. Fig. S6: Details for all protein motifs in Fig. 2. Fig. S7: Sequence analysis of HvLRX promoter by PlantCARE. Fig. S8: Secondary structure prediction of HvLRX and AK353794.1 using GOR 4. Fig. S9: Subcellular localization predication of HvLRX (Z772) using CELLO v.2.5. Fig. S10: PCR amplification analysis for transgenic lines confirmation. Table S1: Primers for gene cloning, vector construction, and RT-qPCR. Table S2: Nucleotide sequences of HvLRX in Z013, Z033, and Z772. Table S3: The deduced amino acid sequence of HvLRX protein in Z013, Z033, and Z772. Table S4: Nucleotide sequences of TaLRX-2A, TaLRX-2B, and TaLRX-2D. Table S5: Motifs found in the promoter of HvLRX (2000 bp before ATG) detected using PlantCare algorithm. Table S6: Protein properties prediction of HvLRX and AK353794.1 using ProtParam. Table S7: Secondary structure prediction of HvLRX and AK353794.1 using GOR 4. (XLSX 16405 KB)
Rights and permissions
About this article
Cite this article
Liang, J., Zhang, H., Yi, L. et al. Identification of HvLRX, a new dehydration and light responsive gene in Tibetan hulless barley (Hordeum vulgare var. nudum). Genes Genom 43, 1445–1461 (2021). https://doi.org/10.1007/s13258-021-01147-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-021-01147-3