Abstract
Background
Although Pasteurella multocida is highly prevalent pathogen in animals and plays an important role in swine respiratory diseases, only a few studies on the use of bacteriophages specific to Pasteurella multocida disease have been reported.
Objective
The object of this study was to investigate the therapeutic effect of specific P. multocida bacteriophages and to identify genes related to bacteriophage signaling utilizing RNA microarrays in swine nasal turbinate cells.
Methods
Pas-MUP-1 phages were applied 24 h prior to P. multocida infection (1 × 107 cfu/ml) at several concentrations of bacterial infection. Cells were incubated to detect cytokines and 24 h to detect mucin production. And real-time quantitative PCR was performed to examine related genes expression. To determine the change of total gene expression based on P. multocida and Pas-MUP-1 treatment, we performed RNA sequencing experiments.
Results
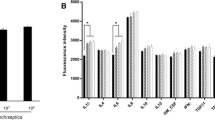
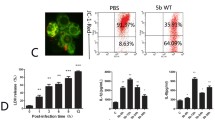
We found that P. multocida-infected PT-K75 cells show increased gene expression of IL-1β, IL-6, and Muc1 in a dose-dependent manner. Interestingly, these genes resulted in decreased expression in P. multocida pretreated with the P. multocida-specific Pas-MUP-1 bacteriophage. RNA sequencing analysis revealed that bacteriophage administration regulated genes associated with immune and inflammatory responses, and the regulated genes were dramatically concentrated in the cytokine/chemokine-based signaling pathways. Pas-MUP-1 treatment was shown to regulate P. multocida induced gene expression in the bacteria.
Conclusion
These results suggest the specific bacteriophage has therapeutic potential as an alternative to antibiotic treatment to defend against P. multocida infection by altering inflammatory gene expression profiles.




Similar content being viewed by others
References
Bartel DP (2009) MicroRNAs: target recognition and regulatory functions. Cell 136:215–233
Blondeau JM, Fitch SD (2019) Mutant prevention and minimum inhibitory concentration drug values for enrofloxacin, ceftiofur, florfenicol, tilmicosin and tulathromycin tested against swine pathogens Actinobacillus pleuropneumoniae, Pasteurella multocida and Streptococcus suis. PLoS ONE 14:e0210154
Brockmeier SL, Register KB, Magyar T, Lax AJ, Pullinger GD, Kunkle RA (2002) Role of the dermonecrotic toxin of Bordetella bronchiseptica in the pathogenesis of respiratory disease in swine. Infect Immun 70:481–490
Brüssow H (2012) What is needed for phage therapy to become a reality in Western medicine? Virology 434:138–142
Cao P, Guo D, Liu J, Jiang Q, Xu Z, Qu L (2017) Genome-wide analyses reveal genes subject to positive selection in Pasteurella multocida. Front Microbiol 8:961
Carrillo MP, Martinez NM, Patiño Mdel P, Iregui CA (2015) Inhibition of Pasteurella multocida adhesion to rabbit respiratory epithelium using lectins. Vet Med Int 2015:365428
Chen Y, Sun E, Song J, Yang L, Wu B (2018) Complete genome sequence of a novel T7-like bacteriophage from a Pasteurella multocida capsular type A isolate. Curr Microbiol 75:574–579
Chitarra CS, Oliveira Filho JX, Morés N, Silva MIVD, Cândido SL, Cezarino PG, Nakazato L, Dutra V (2018) Identification of Pasteurella multocida transcribed genes in porcine lungs through RNAseq. Microb Pathog 22:180–183
Davies RL, MacCorquodale R, Baillie S, Caffrey B (2003) Characterization and comparison of Pasteurella multocida strains associated with porcine pneumonia and atrophic rhinitis. J Med Microbiol 52:59–67
Dharmaraj N, Wang P, Carson DD (2010) Cytokine and progesterone receptor interplay in the regulation of MUC1 gene expression. Mol Endocrinol 24:2253–2266
Dorey L, Lees P (2017) Impact of growth matrix on pharmacodynamics of antimicrobial drugs for pig pneumonia pathogens. BMC Vet Res 13:192
Enss ML, Cornberg M, Wagner S, Gebert A, Henrichs M, Eisenblätter R, Beil W, Kownatzki R, Hedrich HJ (2000) Proinflammatory cytokines trigger MUC gene expression and mucin release in the intestinal cancer cell line LS180. Inflamm Res 49:162–169
Gaemers IC, Vos HL, Volders HH, van der Valk SW, Hilkens J (2001) A stat-responsive element in the promoter of the episialin/MUC1 gene is involved in its overexpression in carcinoma cells. J Biol Chem 276:6191–6199
Gentleman RC, Carey VJ, Bates DM, Bolstad B, Dettling M, Dudoit S, Ellis B, Gautier L, Ge Y, Gentry J, Hornik K, Hothorn T, Huber W, Iacus S, Irizarry R, Leisch F, Li C, Maechler M, Rossini AJ, Sawitzki G, Smith C, Smyth G, Tierney L, Yang JY, Zhang J (2004) Bioconductor: open software development for computational biology and bioinformatics. Genome Biol 5:R80
Harper M, Boyce JD, Adler B (2006) Pasteurella multocida pathogenesis: 125 years after Pasteur. FEMS Microbiol Lett 265:1–10
Kato K, Lillehoj EP, Lu W, Kim KC (2017) MUC1: the first respiratory mucin with an anti-inflammatory function. J Clin Med 6:E110
Langmead B, Salzberg SL (2012) Fast gapped-read alignment with Bowtie 2. Nat Methods 9:357–359
Nakai T, Kume K, Yoshikawa H, Oyamada T, Yoshikawa T (1988) Adherence of Pasteurella multocida or Bordetella bronchiseptica to the swine nasal epithelial cell in vitro. Infect Immun 56:234–240
Oh YH, Moon DC, Lee YJ, Hyun BH, Lim SK (2019) Genetic and phenotypic characterization of tetracycline-resistant Pasteurella multocida isolated from pigs. Vet Microbiol 233:159–163
Park GY, Lee HM, Yu HJ, Son JS, Park SJ, Song KS (2018) Bordetella bronchiseptica bateriophage suppresses B. bronchiseptica-induced inflammation in swine nasal turbinate cells. Genes Genomics 40:1383–1388
Praveena PE, Periasamy S, Kumar AA, Singh N (2010) Cytokine profiles, apoptosis and pathology of experimental Pasteurella multocida serotype A1 infection in mice. Res Vet Sci 89:332–339
Priya GB, Nagaleekar VK, Milton AAP, Saminathan M, Kumar A, Sahoo AR, Wani SA, Kumar A, Gupta SK, Sahoo AP, Tiwari AK, Agarwal RK, Gandham RK (2017) Genome wide host gene expression analysis in mice experimentally infected with Pasteurella multocida. PLoS ONE 12:e0179420
Son JS, Kim EB, Lee SJ, Jun SY, Yoon SJ, Kang SH, Choi YJ (2010) Characterization of Staphylococcus aureus derived from bovine mastitis and isolation of two lytic bacteriophages. J Gen Appl Microbiol 56:347–353
Sulakvelidze A, Alavidze Z, Morris JG Jr (2001) Bacteriophage therapy. Antimicrob Agents Chemother 45:649–659
Tocqueville V, Kempf I, Paboeuf F, Marois-Créhan C (2017) Quantification of Pasteurella multocida in experimentally infected pigs using a real-time PCR assay. Res Vet Sci 112:177–184
Turni C, Singh R, Blackall PJ (2018) Genotypic diversity of Pasteurella multocida isolates from pigs and poultry in Australia. Aust Vet J 96:390–394
Ujvári B, Makrai L, Magyar T (2019) Virulence gene profiling and ompA sequence analysis of Pasteurella multocida and their correlation with host species. Vet Microbiol 233:190–195
Wilson BA, Ho M (2013) Pasteurella multocida: from zoonosis to cellular microbiology. Clin Microbiol Rev 26:631–655
Yeh JC, Lo DY, Chang SK, Chou CC, Kuo HC (2017) Antimicrobial susceptibility, serotypes and genotypes of Pasteurella multocida isolates associated with swine pneumonia in Taiwan. Vet Rec 181:323
Acknowledgements
This work was supported by a grant from the Korea Institute of Planning and Evaluation for Technology in Food, Agriculture, Forestry (IPET) through Technology Commercialization Support Program, funded by Ministry of Agriculture, Food and Rural Affairs (MAFRA) (817029-03-2-SB010).
Author information
Authors and Affiliations
Contributions
GYP and KSS: Designed the study. GYP, HJY, and JSS: Performed experiments. JSS, SJP, HJC, and KSS: Analyzed data. GYP: Wrote the manuscript.
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Park, G.Y., Yu, H.J., Son, J.S. et al. Pasteurella multocida specific bacteriophage suppresses P. multocida-induced inflammation: identification of genes related to bacteriophage signaling by Pasteurella multocida-infected swine nasal turbinate cells. Genes Genom 42, 235–243 (2020). https://doi.org/10.1007/s13258-019-00898-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-019-00898-4