Abstract
Introduction
A high malignancy rate and poor prognosis are common problems with triple-negative breast cancer (TNBC). There is increasing evidence that glycolysis plays vital roles in tumorigenesis, tumor invasion, immune evasion, chemoresistance, and metastasis. However, a comprehensive analysis of the diagnostic and prognostic significance of glycolysis in TNBC is lacking.
Methods
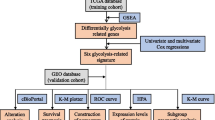
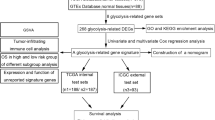
Transcriptomic and clinical data of TNBC patients were obtained from The Cancer Genome Atlas (TCGA) and Molecular Taxonomy of Breast Cancer International Consortium (METABRIC) databases, respectively. Glycolysis-related genes (GRGs) were collected from the Molecular Signatures Database (MSigDB). Differential comparative analysis was performed to obtain the differentially expressed (DE)-GRGs associated with TNBC. Based on the DE-GRGs, a glycolysis-related risk signature was established using Least Absolute Shrinkage and Selector Operation (LASSO) and multivariable Cox regression analyses. The prognostic value, tumor microenvironment, mutation status, and chemotherapy response of different risk groups were analyzed. An independent cohort from the METABRIC database was used for external validation. Furthermore, the expression patterns of five genes derived from the prognostic model were validated by quantitative real-time polymerase chain reaction (RT-qPCR).
Results
The glycolysis-related prognostic signature included five genes (IFNG, ACSS2, IRS2, GFUS, and GAL3ST1) and predicted the prognosis of TNBC patients independent of clinical factors (p < 0.05). Patients were divided into high- and low-risk groups based on the median risk score. Compared to low-risk TNBC patients, high-risk patients had significantly decreased overall survival (HR = 2.718, p < 0.001). Receiver operating characteristic and calibration curves demonstrated that the model had high performance in terms of predicting survival and risk stratification. The results remained consistent after external verification. Additionally, the tumor immune microenvironment significantly differed between the risk groups. Low-risk TNBC patients had a better immunotherapy response than high-risk patients. High-risk TNBC patients with a poor prognosis may benefit from targeted therapy.
Conclusions
This study developed a novel glycolysis and prognosis-related (GRP) signature based on GRGs to predict the prognosis of TNBC patients, and may aid clinical decision-making for these patients.










Similar content being viewed by others
References
Dent R, Trudeau M, Pritchard KI, Hanna WM, Kahn HK, Sawka CA, et al. Triple-negative breast cancer: clinical features and patterns of recurrence. Clin Cancer Res. 2007;13(15 Pt 1):4429–34.
Bianchini G, Balko JM, Mayer IA, Sanders ME, Gianni L. Triple-negative breast cancer: challenges and opportunities of a heterogeneous disease. Nat Rev Clin Oncol. 2016;13(11):674–90.
Emens LA. Breast cancer immunotherapy: facts and hopes. Clin Cancer Res. 2018;24(3):511–20.
Waks AG, Winer EP. Breast cancer treatment: a review. JAMA. 2019;321(3):288–300.
Lee JS, Yost SE, Yuan Y. Neoadjuvant treatment for triple negative breast cancer: recent progresses and challenges. Cancers. 2020;12(6):1404.
André F, Zielinski CC. Optimal strategies for the treatment of metastatic triple-negative breast cancer with currently approved agents. Ann Oncol. 2012;23(Suppl 6):vi46-51.
Lluch A, Barrios CH, Torrecillas L, Ruiz-Borrego M, Bines J, Segalla J, et al. Phase III trial of adjuvant capecitabine after standard neo-/adjuvant chemotherapy in patients with early triple-negative breast cancer (GEICAM/2003-11_CIBOMA/2004-01). J Clin Oncol. 2020;38(3):203–13.
Ganapathy-Kanniappan S. Molecular intricacies of aerobic glycolysis in cancer: current insights into the classic metabolic phenotype. Crit Rev Biochem Mol Biol. 2018;53(6):667–82.
Dias AS, Almeida CR, Helguero LA, Duarte IF. Metabolic crosstalk in the breast cancer microenvironment. Eur J Cancer (Oxford, England: 1990). 2019;121:154–71.
Vander Heiden MG, Cantley LC, Thompson CB. Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science (New York, NY). 2009;324(5930):1029–33.
Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144(5):646–74.
Sansone P, Bromberg J. Targeting the interleukin-6/Jak/stat pathway in human malignancies. J Clin Oncol. 2012;30(9):1005–14.
Yu H, Lee H, Herrmann A, Buettner R, Jove R. Revisiting STAT3 signalling in cancer: new and unexpected biological functions. Nat Rev Cancer. 2014;14(11):736–46.
Meric-Bernstam F, Gonzalez-Angulo AM. Targeting the mTOR signaling network for cancer therapy. J Clin Oncol. 2009;27(13):2278–87.
Comprehensive molecular portraits of human breast tumours. Nature. 2012;490(7418):61–70.
Alam H, Tang M, Maitituoheti M, Dhar SS, Kumar M, Han CY, et al. KMT2D deficiency impairs super-enhancers to confer a glycolytic vulnerability in lung cancer. Cancer Cell. 2020;37(4):599-617.e7.
Kim HS, Xiao C, Wang RH, Lahusen T, Xu X, Vassilopoulos A, et al. Hepatic-specific disruption of SIRT6 in mice results in fatty liver formation due to enhanced glycolysis and triglyceride synthesis. Cell Metab. 2010;12(3):224–36.
Warburg O. On the origin of cancer cells. Science (New York, NY). 1956;123(3191):309–14.
Gatenby RA, Gillies RJ. Why do cancers have high aerobic glycolysis? Nat Rev Cancer. 2004;4(11):891–9.
Fong MY, Zhou W, Liu L, Alontaga AY, Chandra M, Ashby J, et al. Breast-cancer-secreted miR-122 reprograms glucose metabolism in premetastatic niche to promote metastasis. Nat Cell Biol. 2015;17(2):183–94.
Jin J, Qiu S, Wang P, Liang X, Huang F, Wu H, et al. Cardamonin inhibits breast cancer growth by repressing HIF-1α-dependent metabolic reprogramming. J Exp Clin Cancer Res. 2019;38(1):377.
Schug ZT, Peck B, Jones DT, Zhang Q, Grosskurth S, Alam IS, et al. Acetyl-CoA synthetase 2 promotes acetate utilization and maintains cancer cell growth under metabolic stress. Cancer Cell. 2015;27(1):57–71.
Bulusu V, Tumanov S, Michalopoulou E, van den Broek NJ, MacKay G, Nixon C, et al. Acetate recapturing by nuclear acetyl-CoA synthetase 2 prevents loss of histone acetylation during oxygen and serum limitation. Cell Rep. 2017;18(3):647–58.
Miller KD, Pniewski K, Perry CE, Papp SB, Shaffer JD, Velasco-Silva JN, et al. Targeting ACSS2 with a transition-state mimetic inhibits triple-negative breast cancer growth. Can Res. 2021;81(5):1252–64.
Alderton GK. Metabolism: acetate nourishes stressed tumour cells. Nat Rev Cancer. 2015;15(2):67.
Jackson JG, Zhang X, Yoneda T, Yee D. Regulation of breast cancer cell motility by insulin receptor substrate-2 (IRS-2) in metastatic variants of human breast cancer cell lines. Oncogene. 2001;20(50):7318–25.
Cui X, Kim HJ, Kuiatse I, Kim H, Brown PH, Lee AV. Epidermal growth factor induces insulin receptor substrate-2 in breast cancer cells via c-Jun NH(2)-terminal kinase/activator protein-1 signaling to regulate cell migration. Can Res. 2006;66(10):5304–13.
Pankratz SL, Tan EY, Fine Y, Mercurio AM, Shaw LM. Insulin receptor substrate-2 regulates aerobic glycolysis in mouse mammary tumor cells via glucose transporter 1. J Biol Chem. 2009;284(4):2031–7.
Tonetti M, Sturla L, Bisso A, Benatti U, De Flora A. Synthesis of GDP-L-fucose by the human FX protein. J Biol Chem. 1996;271(44):27274–9.
Sun J, Thurin J, Cooper HS, Wang P, Mackiewicz M, Steplewski Z, et al. Elevated expression of H type GDP-l-fucose: β-d-galactoside α-2-l-fucosyltransferase is associated with human colon adenocarcinoma progression. Proc Natl Acad Sci USA. 1995;92(12):5724–8.
Sun Y, Liu X, Zhang Q, Mao X, Feng L, Su P, et al. Oncogenic potential of TSTA3 in breast cancer and its regulation by the tumor suppressors miR-125a-5p and miR-125b. Tumour Biol. 2016;37(4):4963–72.
Owczarek TB, Suchanski J, Pula B, Kmiecik AM, Chadalski M, Jethon A, et al. Galactosylceramide affects tumorigenic and metastatic properties of breast cancer cells as an anti-apoptotic molecule. PLoS One. 2013;8(12): e84191.
Gresser I. Biologic effects of interferons. J Invest Dermatol. 1990;95(6 Suppl):66s–71s.
Castro F, Cardoso AP, Gonçalves RM, Serre K, Oliveira MJ. Interferon-gamma at the crossroads of tumor immune surveillance or evasion. Front Immunol. 2018;9:847.
Mojic M, Takeda K, Hayakawa Y. The dark side of IFN-γ: its role in promoting cancer immunoevasion. Int J Mol Sci. 2017;19(1):89.
Siska PJ, Rathmell JC. Metabolic signaling drives IFN-γ. Cell Metab. 2016;24(5):651–2.
Jiang B. Aerobic glycolysis and high level of lactate in cancer metabolism and microenvironment. Genes Dis. 2017;4(1):25–7.
Zhang Y, Yu G, Chu H, Wang X, Xiong L, Cai G, et al. Macrophage-associated PGK1 phosphorylation promotes aerobic glycolysis and tumorigenesis. Mol Cell. 2018;71(2):201-15.e7.
Jiang Z, Liu Z, Li M, Chen C, Wang X. Increased glycolysis correlates with elevated immune activity in tumor immune microenvironment. EBioMedicine. 2019;42:431–42.
Deberardinis RJ, Sayed N, Ditsworth D, Thompson CB. Brick by brick: metabolism and tumor cell growth. Curr Opin Genet Dev. 2008;18(1):54–61.
Ohashi T, Akazawa T, Aoki M, Kuze B, Mizuta K, Ito Y, et al. Dichloroacetate improves immune dysfunction caused by tumor-secreted lactic acid and increases antitumor immunoreactivity. Int J Cancer. 2013;133(5):1107–18.
Chang CH, Qiu J, O’Sullivan D, Buck MD, Noguchi T, Curtis JD, et al. Metabolic competition in the tumor microenvironment is a driver of cancer progression. Cell. 2015;162(6):1229–41.
Vasaikar S, Huang C, Wang X, Petyuk VA, Savage SR, Wen B, et al. Proteogenomic analysis of human colon cancer reveals new therapeutic opportunities. Cell. 2019;177(4):1035-49.e19.
Cascone T, McKenzie JA, Mbofung RM, Punt S, Wang Z, Xu C, et al. Increased tumor glycolysis characterizes immune resistance to adoptive T cell therapy. Cell Metab. 2018;27(5):977-87.e4.
Rooney MS, Shukla SA, Wu CJ, Getz G, Hacohen N. Molecular and genetic properties of tumors associated with local immune cytolytic activity. Cell. 2015;160(1–2):48–61.
Moossavi M, Parsamanesh N, Bahrami A, Atkin SL, Sahebkar A. Role of the NLRP3 inflammasome in cancer. Mol Cancer. 2018;17(1):158.
Pardoll DM. The blockade of immune checkpoints in cancer immunotherapy. Nat Rev Cancer. 2012;12(4):252–64.
Tumeh PC, Harview CL, Yearley JH, Shintaku IP, Taylor EJ, Robert L, et al. PD-1 blockade induces responses by inhibiting adaptive immune resistance. Nature. 2014;515(7528):568–71.
Karn T, Jiang T, Hatzis C, Sänger N, El-Balat A, Rody A, et al. Association between genomic metrics and immune infiltration in triple-negative breast cancer. JAMA Oncol. 2017;3(12):1707–11.
Kon E, Benhar I. Immune checkpoint inhibitor combinations: current efforts and important aspects for success. Drug Resist Updates. 2019;45:13–29.
Francisco LM, Salinas VH, Brown KE, Vanguri VK, Freeman GJ, Kuchroo VK, et al. PD-L1 regulates the development, maintenance, and function of induced regulatory T cells. J Exp Med. 2009;206(13):3015–29.
Bhattacharya B, Mohd Omar MF, Soong R. The Warburg effect and drug resistance. Br J Pharmacol. 2016;173(6):970–9.
He M, Jin Q, Chen C, Liu Y, Ye X, Jiang Y, et al. The miR-186-3p/EREG axis orchestrates tamoxifen resistance and aerobic glycolysis in breast cancer cells. Oncogene. 2019;38(28):5551–65.
Metzger-Filho O, Tutt A, de Azambuja E, Saini KS, Viale G, Loi S, et al. Dissecting the heterogeneity of triple-negative breast cancer. J Clin Oncol. 2012;30(15):1879–87.
Asghar US, Barr AR, Cutts R, Beaney M, Babina I, Sampath D, et al. Single-cell dynamics determines response to CDK4/6 inhibition in triple-negative breast cancer. Clin Cancer Res. 2017;23(18):5561–72.
Teo ZL, Versaci S, Dushyanthen S, Caramia F, Savas P, Mintoff CP, et al. Combined CDK4/6 and PI3Kα inhibition is synergistic and immunogenic in triple-negative breast cancer. Cancer Res. 2017;77(22):6340–52.
Mao Z, Zhang W. Role of mTOR in glucose and lipid metabolism. Int J Mol Sci. 2018;19(7):2043.
Yamamoto T, Kanaya N, Somlo G, Chen S. Synergistic anti-cancer activity of CDK4/6 inhibitor palbociclib and dual mTOR kinase inhibitor MLN0128 in pRb-expressing ER-negative breast cancer. Breast Cancer Res Treat. 2019;174(3):615–25.
Lanceta L, Lypova N, O’Neill C, Li X, Rouchka E, Chesney J, et al. Differential gene expression analysis of palbociclib-resistant TNBC via RNA-seq. Breast Cancer Res Treat. 2021;186(3):677–86.
Acknowledgements
Funding
This project was financially supported by the National Science Foundation of China (grant number: 81871295, 82001740). Ji-min Cao funded the journal’s Rapid Service fees.
Editorial Assistance
We gratefully acknowledge contributions from the GEO, TCGA, and METABRIC databases. The English in this document has been checked by at least two professional editors of Textcheck (http://www.textcheck.com/) both native speakers of English.
Author Contributions
All authors were involved in drafting the article or revising it critically for important content. Jian Zheng and Ji-Min Cao developed the methodology and acquired the related data. Conception and design of the study: Jian Zheng and Ji-Min Cao. Acquisition of data: Yi-Fan Zhang, Guo-Hui Han, Meng-Ying Fan, Ming-Hui Du, Guo-Chen Zhang and Yue Feng. Analysis and interpretation of data: Run-Qi Chen, Jun Qiao, Jian Zheng, Bo Zhang, Yi-Fan Zhang and Sheng-Xiao Zhang. Jian Zheng collected specimens and carried out in vitro assays. Drafting the article: Jian Zheng, Yi-Fan Zhang and Sheng-Xiao Zhang. Revising the article critically: Ji-Min Cao, Sheng-Xiao Zhang, and Guo-Hui Han. All authors contributed to the article and approved the submitted version.
Disclosures
All named authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Compliance with Ethics Guidelines
The study was approved by the Ethics Review Board at Shanxi Cancer Hospital. All patients provided written informed consent for the experimental analysis of their excised tissues.
Data Availability
The datasets analyzed for this study can be found in the [TCGA database] [https://cancergenome.nih.gov], [METABRIC database] [http://molonc.bccrc.ca/aparicio-lab/research/metabric/] and [GTEx database] [https://gtexportal.org/]. All data generated or analyzed during this study are included as supplementary information files.
Author information
Authors and Affiliations
Corresponding author
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zheng, J., Zhang, YF., Han, GH. et al. Identification and Validation of a Novel Glycolysis-Related Gene Signature for Predicting the Prognosis and Therapeutic Response in Triple-Negative Breast Cancer. Adv Ther 40, 310–330 (2023). https://doi.org/10.1007/s12325-022-02330-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12325-022-02330-y