Abstract
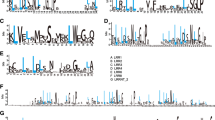
Coding sequence (CDS) architecture affects gene expression levels in organisms. Codon optimization can increase the gene expression level. Therefore, understanding codon usage patterns has important implications for research on genetic engineering and exogenous gene expression. To date, the codon usage patterns of many model plants have been analyzed. However, the relationship between CDS architecture and gene expression in Arachis duranensis remains poorly understood. According to the results of genome sequencing, A. duranensis has many resistant genes that can be used to improve the cultivated peanut. In this study, bioinformatic approaches were used to estimate A. duranensis CDS architectures, including frequency of the optimal codon (Fop), polypeptide length and GC contents at the first (GC1), second (GC2) and third (GC3) codon positions. In addition, Arachis RNA-seq datasets were downloaded from PeanutBase. The relationships between gene expression and CDS architecture were assessed both under normal growth as well as nematode and drought stress conditions. A total of 26 codons with high frequency were identified, which preferentially ended with A or T in A. duranensis CDSs under the above-mentioned three conditions. A similar CDS architecture was found in differentially expressed genes (DEGs) under nematode and drought stresses. The GC1 content differed between DEGs and non-differentially expressed genes (NDEGs) under both drought and nematode stresses. The expression levels of DEGs were affected by different CDS architectures compared with NDEGs under drought stress. In addition, no correlation was found between differential gene expression and CDS architecture neither under nematode nor under drought stress. These results aid the understanding of gene expression in A. duranensis.


Similar content being viewed by others
References
Akashi H (1994) Synonymous codon usage in Drosophila melanogaster: nature selection and translation accuracy. Genetics 136(3):927–935
Akashi H (2001) Gene expression and molecular evolution. Curr Opin Genet Dev 11:660–666
Anders S, Huber W (2010) Differential expression analysis for sequence count data. Genome Biol 11(10):106–110
Arava Y (2003) Isolation of polysomal RNA for microarray analysis. Methods Mol Bio 224:79–87
Arunkumar R, Josephs EB, Williamson RJ, Wright SI (2013) Pollen-specific, but not sperm-specific, genes show stronger purifying selection and higher rates of positive selection than sporophytic genes in Capsella grandiflora. Mol Bio Evol 30(11):2475–2486
Begley U, Dyavaiah M, Patil A, Rooney JP, DiRenzo D, Young CM, Conklin DS, Zitomer RS, Begley TJ (2007) Trm9-catalyzed tRNA modifications link translation to the DNA damage response. Mol Cell 28:860–870
Behura SK, Severson DW (2012) Comparative analysis of codon usage bias and codon context patterns between dipteran and hymenopteran sequenced genomes. PLoS ONE 7(8):e43111
Bertioli DJ, Seijo G, Freitas FO, Valls JFM, Leal-Bertioli SCM, Moretzsohn MC (2011) An overview of peanut and its wild relatives. Plant Genet Resour Characteriz Utiliz 9(1):134–149
Bertioli DJ, Cannon SB, Froenicke L, Huang G, Farmer AD, Cannon EKS, Liu X, Gao D, Clevenger J, Dash S, Ren L, Moretzsohn MC, Shirasawa K, Huang W, Vidigal B, Abernathy B, Chu Y, Burow MD, Varshney RK, Wang X, Zhang X, Barkley N, Guimarães PM, Isobe S, Guo B, Liao B, Stalker HT, Schmitz RJ, Scheffler BE, Leal-Bertioli SCM, Xun X, Jackson SA, Michelmore R, Ozias-Akins P (2016) The genome sequences of Arachis duranensis and Arachis ipaensis, the diploid ancestors of cultivated peanut. Nat Genet 48(4):438–446
Bertioli DJ, Jenkins J, Clevenger J, Dudchenko O, Gao D, Seijo G, Leal-Bertioli SCM, Ren L, Farmer AD, Pandey MK, Samoluk SS, Abernathy B, Agarwal G, Ballé-Taborda C, Cameron C, Campbell J, Chavarro C, Chitikineni A, Chu Y, Dash S, Eiaidouri M, Guo B, Huang W, Kim KD, Korani W, Lanciano S, Lui CG, Mirouze M, Moretzsohn MC, Pham M, Shin JH, Shirasawa K, Sinharoy S, Sreedasyam A, Weeks NT, Zhang X, Zheng Z, Sun Z, Froenicke L, Aiden EL, Michelmore R, Varshney RK, Holbrook CC, Cannon EKS, Scheffler BE, Grimwood J, Ozias-Akins P, Cannon SB, Jackson SA, Schmutz J (2019) The genome sequence of segmental allotetraploid peanut Arachis hypogaea. Nat Genet 51:877–884
Bertioli DJ, Abernathy B, Seijo G, Clevenger J, Cannon SB (2020) Evaluating two different models of peanut’s origin. Nat Genet 52:557–559
Brandis G, Hughes D (2016) The selective advantages of synonymous codon usage bias in Salmonella. PLoS Genet 12(3):e1005926
Brasileiro ACM, Morgante CV, Araujo ACG, Leal-Bertioli SCM, Silva AK, Martins ACQ, Vinson CC, Santos CMR, Bonfim O, Togawa RC, Saraiva MAP, Bertioli DJ, Guimaraes PM (2015) Transcriptome profiling of wild Arachis from water-limited environments uncovers drought tolerance candidate genes. Plant Mol Biol Rep 33:1876–1892
Camiolo S, Melito S, Porceddu A (2015) New insights into the interplay between codon bias determinants in plants. DNA Res 5:1–9
Castillo-Davis C, Mekhedov S, Hartl D, Koonin E, Kondrashov F (2002) Selection for short introns in highly expressed genes. Nat Genet 31:415–418
Chan CT, Pang YL, Deng W, Babu IR, Dyavaiah M, Begley TJ, Dedon PC (2012) Reprogramming of tRNA modifications controls the oxidative stress response by codon-biased translation of proteins. Nat Commun 3:937
Chaney J, Clark PL (2015) Roles for synonymous codon usage in protein biogenesis. Ann Rev Biophys 44:143–166
Chu D, Kazana E, Bellanger N, Singh T, Tuite MF, von der Haar T (2014) Translation elongation can control translation initiation on eukaryotic mRNA. EMBO J 33:21–34
Clevenger J, Chu Y, Scheffler B, Ozias-Akins P (2016) A developmental transcriptome map for allotetraploid Arachis hypogaea. Front Plant Sci 7:1446
Dangl JL, Jones JD (2001) Plant pathogens and integrated defence responses to infection. Nature 411(6839):826–833
Dash S, Cannon EKS, Kalberer SR, Farmer AD, Cannon SB (2016) PeanutBase and other bioinformatic resources for peanut. In: Stalker HT, Wilson RF (eds) Peanuts Genetics Processing and Utilization. AOCS Press, USA, pp 241–252
De La Torre AR, Lin YC, Van de Peer Y, Ingvarsson PK (2015) Genome-wide analysis reveals diverged pattern of codon bias, gene expression and rates of sequence evolution in Picea gene families. Genome Biol Evol 7(4):1002–1015
Duret L (2000) tRNA gene number and codon usage in the C. elegans genome are co-adapted for optimal translation of highly expressed genes. Trends Genet 16:287–289
Duret L, Mouchiroud D (1999) Expression pattern and surprisingly, gene length shape codon usage in Caenorhabdits, Drosophila and Arabidopsis. Proc Natl Acad Sci USA 96(8):4482–4487
Elhaik E, Pellegrini M, Tatarinova TV (2014) Gene expression and nucleotide composition are associated with genic methylation level in Oryza sativa. BMC Bioinformatics 15:23
Ellegren H, Parsch J (2007) The evolution of sex-biased genes and sex-biased gene expression. Nat Rev Genet 8:689–698
Glémin S, Clément Y, David J, Ressayre A (2014) GC content evolution in coding regions of angiosperm genomes: a unifying hypothesis. Trends Genet 30(7):263–270
Guimarães PM, Guimaraes LA, Morgante CV, Silva OB Jr, Araujo ACG, Martins ACQ, Saraiva MAP, Oliveira TN, Togawa RC, Leal-Bertioli SCM, Bertioli DJ, Brasileiro ACM (2015) Root transcriptome analysis of wild peanut reveals candidate genes for nematode resistance. PLoS ONE 10(10):e0140937
Gustafsson C, Govindarajan S, Minshull J (2004) Codon bias and heterologous protein expression. Trends Biochem Sci 22:346–353
He B, Dong H, Jiang C, Cao F, Tao S, Xu L (2016) Analysis of codon usage patterns in Ginkgo biloba reveals codon usage tendency from A/U-ending to G/C ending. Sci Rep 6:35927
Hershberg R, Petrov DA (2008) Selection on codon bias. Annu Rev Genet 42:287–299
Hershberg R, Petrov DA (2009) General rules for optimal codon choice. PLoS Genet 5(7):e1000556
Ingvarsson PK (2007) Gene expression and protein length influence codon usage and rates of sequence evolution in Populus tremula. Mol Bio Evol 24(3):836–844
Jia X, Liu S, Zheng H, Li B, Qi Q, Wei L, Zhao T, He J, Sun J (2015) Non-uniqueness of factors constraint on the codon usage in Bombyx mori. BMC Genomics 16:356
Jiang Y, Deng F, Wang H, Hu Z (2008) An extensive analysis on the global codon usage pattern of baculoviruses. Arch Virol 153:2273–2282
Kawabe A, Miyashita NT (2003) Patterns of codon usage bias in three dicot and four monocot plant species. Genes Genet Syst 78(5):343–352
Kochert G, Stalker H, Gimenes M, Galgaro M, Lopes C, Moore K (1996) RFLP and cytogenetic evidence on the origin and evolution of allotetraploid domesticated peanut, Arachis hypogaea (Leguminosae). Am J Bot 83(10):1282–1291
Lavner Y, Kotlar D (2005) Codon bias as a factor in regulating expression via translation rate in the human genome. Gene 345:127–138
Leal-Bertioli SCM, Bertioli DJ, Guimarães PM, Pereira TD, Galhardo I, Silva JP, Brasileiro ACM, Oliveira RS, Silva PÍT, Vadez V, Araujo ACG (2012) The effect tetraploidization of wild Arachis on leaf morphology and other drought-related traits. Environ Exp Bot 84:17–24
Li B, Dewey CN (2011) RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinfo 12:323
Li N, Li Y, Zheng C, Huang J, Zhang S (2016a) Genome-wide comparative analysis of the codon usage patterns in plants. Genes Genom 38(8):723–731
Li X, Song H, Kuang Y, Chen S, Tian P, Li C, Nan Z (2016b) Genome-wide analysis of codon usage bias in Epichloë festucae. Int J Mol Sci 17(7):E1138
Liu Q, Hu H, Wang H (2015) Mutational bias is the driving force for shaping the synonymous codon usage pattern of alternatively spliced in rice (Oryza sativa L). Mol Genet Genomics 290(2):649–660
Maurer-Alcalá XX, Katz LA (2016) Nuclear architecture and patterns of molecular evolution are correlated in the Chilodonella uncinata. Genome Biol Evol 8(6):1634–1642
Mitra S, Ray SK, Banerjee R (2016) Synonymous codon influencing gene expression. Resear Rep Biochem 6:57–65
Mohasses FC, Solouki M, Ghareyazie B, Fahmideh L, Mohsenpour M (2020) Correlation between gene expression levels under drought stress and synonymous codon usage in rice plant by in-silico study. PLoS ONE 15(8):e0237334
Morton BR, Wright SI (2007) Selective constraints on codon usage of nuclear genes from Arabidopsis thaliana. Mol Bio Evol 24:122–129
Nepal MP, Andersen EJ, Neupane S, Benson BV (2017) Comparative genomics of non-TNL disease resistance genes from six plant species. Genes 8:249
Qiu S, Bergero R, Zeng K, Charlesworth D (2011) Patterns of codon usage bias in Silene latifolia. Mol Bio Evol 28(1):771–780
Quax TEF, Claassens NJ, Söll D, van der Oost J (2015) Codon bias as a means to fine-tune gene expression. Mol Cell 59:149–161
Ramos M, Fleming G, Chu Y, Akiyama Y, Gallo M, Ozias-Akins P (2006) Chromosomal and phylogenetic context for conglutin genes in Arachis based on genomic sequence. Mol Genet Genomics 275(6):578–592
RoyChoudhury S, Mukherjee D (2010) A detailed comparative analysis on the overall codon usage pattern in herpesviruses. Virus Res 148(1–2):31–43
Seijo J, Lavia G, Fernandez A, Krapovickas A, Ducasse D, Moscone E (2004) Physical mapping of the 5S and 18S–25S rRNA genes by FISH as evidence that Arachis duranensis and A ipaënsis are the wild diploid progenitors of A hypogaea (Leguminosae). Am J Bot 91(9):1294–1303
Seijo G, Lavia GI, Fernandez A, Krapovickas A, Ducasse DA, Bertioli DJ, Moscone EA (2007) Genomic relationships between the cultivated peanut (Arachis hypogaea, Leguminosae) and its close relatives revealed by double GISH. Am J Bot 94(12):1963–1971
Sharp PM, Li WH (1986) An evolutionary perspective on synonymous codon usage in unicellular organisms. J Mol Evol 24:28–38
Sharp PM, Li WH (1987) The codon adaption index - a measure of directional synonymous codon usage bias, and its potential applications. Nucleic Acids Res 15:1281–1295
Sharp PM, Tuohy TM, Mosurski KR (1986) Codon usage in yeast: cluster analysis clearly differentiates highly and lowly expressed genes. Nucleic Acids Res 14:5125–5143
Sidorenko LV, Lee T, Woosley A, Moskal WA, Bevan SA, Ann Owens Merlo P, Walsh TA, Wang X, Weaver S, Glancy TP, Wang P, Yang X, Sriram S, Meyers BC (2017) GC-rich coding sequences reduce transposon-like, small RNA-mediated transgene silencing. Nat Plants 3:875–884
Singh R, Ming R, Yu Q (2016) Comparative analysis of GC content variations in plant genomes. Trop Plant Biol 9(3):136–149
Song H, Wang P, Hou L, Zhao S, Zhao C, Xia H, Li P, Zhang Y, Bian X, Wang X (2016) Global analysis of WRKY genes and their response to dehydration and salt stress in soybean. Front Plant Sci 7:9
Song H, Gao H, Liu J, Tian P, Nan Z (2017a) Comprehensive analysis of correlations among codon usage bias, gene expression, and substitution rate in Arachis duranensis and Arachis ipaënsis orthologs. Sci Rep 7:14853
Song H, Liu J, Song Q, Zhang Q, Tian P, Nan Z (2017b) Comprehensive analysis of codon usage bias in seven Epichloë species and their peramine-coding genes. Front Microbiol 8:1419
Song H, Zhang Q, Tian P, Nan Z (2017c) Differential evolutionary patterns and expression levels between sex-specific and somatic tissue-specific genes in peanut. Sci Rep 7:9016
Song H, Guo Z, Chen T, Sun J, Yang G (2018a) Genome-wide identification of LRR-containing sequences and the response of these sequences to nematode infection in Arachis duranensis. BMC Plant Biol 18:279
Song H, Liu J, Chen T, Nan Z (2018b) Synonymous codon usage pattern in model legume Medicago truncatula. J Integr Agr 17(9):2074–2081
Song H, Sun J, Yang G (2018c) Comparative analysis of selection mode reveals different evolutionary rate and expression pattern in Arachis duranensis and Arachis ipaënsis duplicated genes. Plant Mol Biol 98(4–5):349–361
Whittle CA, Extavour CG (2015) Codon and amino acid usage are shaped by selection across divergent model organisms of the Pancrustacea. G3 Gene Genom Genet 5:2307–2321
Williford A, Demuth JP (2012) Gene expression levels are correlated with synonymous codon usage, amino acid composition, and gene architecture in the red flour beetle. Tribolium Castaneum Mol Bio Evol 29(12):3755–3766
Xu Y, Ma P, Shah P, Rokas A, Liu Y, Johnson CH (2013) Non-optimal codon usage is a mechanism to achieve circadian clock conditionality. Nature 495:116–120
Yang H (2009) In plants, expression breadth and expression level distinctly and non-linearly correlate with gene structure. Biology Direct 4:45
Acknowledgements
The work was supported by the First Class Grassland Science Discipline Program of Shandong Province, China, the Natural Science Foundation of Shandong Province, China (ZR2019QC017, ZR2020MC172 and ZR2020QC185), Key Research and Development Program in Shandong Province (2019GNC106077), and Start-up Foundation for High Talents of Qingdao Agricultural University (No. 665/1120012, No. 663/1119038 and No. 663/1119040).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
There is no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Dong, S., Zhang, L., Pang, W. et al. Comprehensive analysis of coding sequence architecture features and gene expression in Arachis duranensis. Physiol Mol Biol Plants 27, 213–222 (2021). https://doi.org/10.1007/s12298-021-00938-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12298-021-00938-y