Abstract
Purpose
Melanoma is a malignant skin tumor, and its incidence is rising. To explore the specific differences in benign and malignant melanoma at the genetic level, we performed a series of bioinformatics analyses, including differential gene analysis, co-expression analysis, enrichment analysis, and regulatory prediction.
Methods
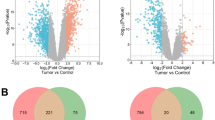
The microarray data of benign and malignant melanocytes were downloaded from GEO, and 1917 differential genes were obtained by differential analysis (p < 0.05). Weighted gene co-expression network analysis obtained three functional barrier modules. The essential genes of each module are SMARTA4, HECA, and C1R.
Results
The results of the enrichment analysis showed that the dysfunctional module gene was mainly associated with RNA splicing and Adherens junction. Through the pivotal analysis of ncRNA, it was found that miR-448, miR-152-3p, and miR-302b-3p essentially regulate three modules, which we consider to be critical regulators. In the pivot analysis of TF, more control modules include ARID3A, E2F1, E2F3, and E2F8.
Conclusions
We believe that the regulator (miR-448, miR-152-3p, miR-302b-3p) regulates the expression of the core gene SMARCA4, which in turn affects the signal transduction of the Adherens junction. It eventually leads to the deterioration of benign skin spasms into melanoma.






Similar content being viewed by others
References
Situm M, Buljan M, Kolic M, Vucic M. Melanoma–clinical, dermatoscopical, and histopathological morphological characteristics. Acta Dermatovenerol Croat. 2014;22:1–12.
Berger AC. Melanoma. Surg Oncol Clin N Am. 2015;24:xv–xvi.
Rastrelli M, Tropea S, Rossi CR, Alaibac M. Melanoma: epidemiology, risk factors, pathogenesis, diagnosis and classification. Vivo. 2014;28:1005–111.
Read J, Wadt KA, Hayward NK. Melanoma genetics. J Med Genet. 2016;53:1–14.
Shannan B, Perego M, Somasundaram R, Herlyn M. Heterogeneity in melanoma. Cancer Treat Res. 2016;167:1–15.
Fischer GM, Vashisht Gopal YN, McQuade JL, Peng W, DeBerardinis RJ, Davies MA. Metabolic strategies of melanoma cells: mechanisms, interactions with the tumor microenvironment, and therapeutic implications. Pigment Cell Melanoma Res. 2018;31:11–30.
Leachman SA, Lucero OM, Sampson JE, Cassidy P, Bruno W, Queirolo P, Ghiorzo P. Identification, genetic testing, and management of hereditary melanoma. Cancer Metastasis Rev. 2017;36:77–90.
Abbas O, Miller DD, Bhawan J. Cutaneous malignant melanoma: update on diagnostic and prognostic biomarkers. Am J Dermatopathol. 2014;36:363–79.
Chen X, Gao G, Liu S, Yu L, Yan D, Yao X, Sun W, Han D, Dong H. Long noncoding RNA PVT1 as a novel diagnostic biomarker and therapeutic target for melanoma. Biomed Res Int. 2017;2017:7038579.
Torabian S, Kashani-Sabet M. Biomarkers for melanoma. Curr Opin Oncol. 2005;17:167–71.
Kunz M, Holzel M. The impact of melanoma genetics on treatment response and resistance in clinical and experimental studies. Cancer Metastasis Rev. 2017;36:53–755.
Leonardi GC, Falzone L, Salemi R, Zanghi A, Spandidos DA, McCubrey JA, Candido S, Libra M. Cutaneous melanoma: from pathogenesis to therapy (Review). Int J Oncol. 2018;52:1071–80.
Bombelli FB, Webster CA, Moncrieff M, Sherwood V. The scope of nanoparticle therapies for future metastatic melanoma treatment. Lancet Oncol. 2014;15:e22–32.
Tang T, Eldabaje R, Yang L. Current status of biological therapies for the treatment of metastatic melanoma. Anticancer Res. 2016;36:3229–411.
Talantov D, Mazumder A, Yu JX, Briggs T, Jiang Y, Backus J, Atkins D, Wang Y. Novel genes associated with malignant melanoma but not benign melanocytic lesions. Clin Cancer Res. 2005;11:7234–42.
Barrett T, Wilhite SE, Ledoux P, Evangelista C, Kim IF, Tomashevsky M, Marshall KA, Phillippy KH, Sherman PM, Holko M, Yefanov A, Lee H, Zhang N, Robertson CL, Serova N, Davis S, Soboleva A. NCBI GEO: archive for functional genomics data sets–update. Nucleic Acids Res. 2013;41:D991–D99595.
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, Smyth GK. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015;43:e47.
Law CW, Chen Y, Shi W, Smyth GK. Voom: precision weights unlock linear model analysis tools for RNA-seq read counts. Genome Biol. 2014;15:R29.
Smyth GK (2004) Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat Appl Genet Mol Biol. 3: Article3.
Langfelder P, Horvath S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics. 2008;9:559.
Liu W, Li L, Ye H, Tu W. Weighted gene co-expression network analysis in biomedicine research. Sheng Wu Gong Cheng Xue Bao. 2017;33:1791–801.
Yu G, Wang LG, Han Y, He QY. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS. 2012;16:284–7.
Xu MJ, Dorsey JF, Amaravadi R, Karakousis G, Simone CB 2nd, Xu X, Xu W, Carpenter EL, Schuchter L, Kao GD. Circulating tumor cells, DNA, and mRNA: potential for clinical utility in patients with melanoma. Oncologist. 2016;21:84–94.
Kappelmann M, Bosserhoff A, Kuphal S. AP-1/c-Jun transcription factors: regulation and function in malignant melanoma. Eur J Cell Biol. 2014;93:76–81.
Pei G, Chen L, Zhang W. WGCNA application to proteomic and metabolomic data analysis. Methods Enzymol. 2017;585:135–58.
Wang LX, Li Y, Chen GZ. Network-based co-expression analysis for exploring the potential diagnostic biomarkers of metastatic melanoma. PLoS ONE. 2018;13:e0190447.
Journe F, Id Boufker H, Van Kempen L, Galibert MD, Wiedig M, Sales F, Theunis A, Nonclercq D, Frau A, Laurent G, Awada A, Ghanem G. TYRP1 mRNA expression in melanoma metastases correlates with clinical outcome. Br J Cancer. 2011;105:1726–32.
El Hajj P, Gilot D, Migault M, Theunis A, van Kempen LC, Sales F, Fayyad-Kazan H, Badran B, Larsimont D, Awada A, Bachelot L, Galibert MD, Ghanem G, Journe F. SNPs at miR-155 binding sites of TYRP1 explain discrepancy between mRNA and protein and refine TYRP1 prognostic value in melanoma. Br J Cancer. 2015;113:91–8.
Sheppard HM, Feisst V, Chen J, Print C, Dunbar PR. AHNAK is downregulated in melanoma, predicts poor outcome, and may be required for the expression of functional cadherin-1. Melanoma Res. 2016;26:108–16.
Wan X, Liu R, Li Z. The prognostic value of HRAS mRNA expression in cutaneous melanoma. Biomed Res Int. 2017;2017:5356737.
Hind CK, Carter MJ, Harris CL, Chan HT, James S, Cragg MS. Role of the pro-survival molecule Bfl-1 in melanoma. Int J Biochem Cell Biol. 2015;59:94–102.
Craig EA, Weber JD, Spiegelman VS. Involvement of the mRNA binding protein CRD-BP in the regulation of metastatic melanoma cell proliferation and invasion by hypoxia. J Cell Sci. 2012;125:5950–4.
Park EJ, Lee YM, Oh TI, Kim BM, Lim BO, Lim JH. Vanillin suppresses cell motility by inhibiting STAT3-mediated HIF-1alpha mRNA expression in malignant melanoma cells. Int J Mol Sci. 2017;18:532.
Saenger Y, Magidson J, Liaw B, de Moll E, Harcharik S, Fu Y, Wassmann K, Fisher D, Kirkwood J, Oh WK, Friedlander P. Blood mRNA expression profiling predicts survival in patients treated with tremelimumab. Clin Cancer Res. 2014;20:3310–8.
Author information
Authors and Affiliations
Contributions
SL and XY are responsible for the conception or design of the work. SL, LQ, ZZ and YJ contribute the acquisition, analysis, or interpretation of data for the work. XY and LQ provide the tissue samples. ZZ helps in the follow-up of the patients. SL helps in reviewing the histopathology slides. All authors finally approved the manuscript version to be published. YJ is the guarantor of the article.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
The study was approved by Ethical Committee of The Second Hospital of Jilin University and conducted in accordance with the ethical standards.
Informed consent
Yes.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Liu, SR., Yang, X., Qi, L. et al. SMARCA4 promotes benign skin malignant transformation into melanoma through Adherens junction signal transduction. Clin Transl Oncol 23, 591–600 (2021). https://doi.org/10.1007/s12094-020-02453-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12094-020-02453-0