Abstract
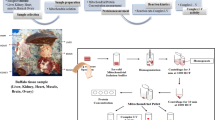
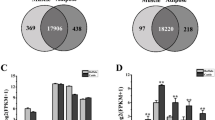
The physiological well-being of buffaloes, encompassing phenotypic traits, reproductive health, and productivity, depends on their energy status. Mitochondria, the architects of energy production, orchestrate a nuanced interplay between nuclear and mitochondrial domains. Oxidative phosphorylation complexes and associated proteins wield significant influence over metabolic functions, energy synthesis, and organelle dynamics, often linked to tissue-specific pathologies. The unexplored role of ATP synthase in buffalo tissues prompted a hypothesis: in-depth exploration of nuclear-derived mitochondrial genes, notably ATP synthase, reveals distinctive tissue-specific diversity. RNA extraction and sequencing of buffalo tissues (kidney, heart, brain, and ovary) enabled precise quantification of nuclear-derived mitochondrial protein gene expression. The analysis unveiled 24 ATP synthase transcript variants, each with unique tissue-specific patterns. Kidney, brain, and heart exhibited elevated gene expression compared to ovaries, with 10, 8, and 19 up-regulated genes, respectively. The kidney showed 3 and 12 down-regulated genes compared to the brain and heart. The heart–brain comparison highlighted ten highly expressed genes in ATP synthase functions. Gene ontology and pathway analyses revealed enriched functions linked to ATP synthesis and oxidative phosphorylation, offering a comprehensive understanding of energy production in buffalo tissues. This analysis enhances understanding of tissue-specific gene expression, emphasizing the influence of energy demands. Revealing intricate links between mitochondrial gene expression and tissue specialization in buffaloes, it provides nuanced insights into tissue-specific expression of nuclear-encoded mitochondrial protein genes, notably ATP synthase, advancing the comprehension of buffalo tissue biology.







Similar content being viewed by others
Data Availability
Data generated from this study are presented in the manuscript. Further details are presented in additional files. The information not included would be shared by the corresponding author on reasonable request.
References
Wahid, H., Rosnina, Y. (2016). Buffalo: Asia. Reference module in food science B 978-0-08-100596-5.21231–6. doi: https://doi.org/10.1016/B978-0-08-100596-5.21231-6
Nolfi-Donegan, D., Braganza, A., & Shiva, S. (2020). Mitochondrial electron transport chain: Oxidative phosphorylation, oxidant production, and methods of measurement. Redox Biology, 37, 101674. https://doi.org/10.1016/j.redox.2020.101674
Nakhle, J., Rodriguez, A. M., & Vignais, M. L. (2020). Multifaceted roles of mitochondrial components and metabolites in metabolic diseases and cancer. International Journal of Molecular Sciences, 21, 4405. https://doi.org/10.3390/ijms21124405
Chandel, N. S. (2021). Mitochondria. Cold Spring Harbor Perspectives in Biology, 13, a040543. https://doi.org/10.1101/cshperspect.a040543
Al-Kafaji, G., & Golbahar, J. (2013). High glucose-induced oxidative stress increases the copy number of mitochondrial DNA in human mesangial cells. BioMed Research International, 2013, 754946.
Laubenthal, L., Hoelker, M., Frahm, J., Dänicke, S., Gerlach, K., Südekum, K. H., Sauerwein, H., & Häussler, S. (2016). Mitochondrial DNA copy number and biogenesis in different tissues of early- and late-lactating dairy cows. Journal of Dairy Science, 99, 1571–1583.
Sadeesh, E. M., Singla, N., Lahamge, M. S., Kumari, S., Ampadi, A. N., & Malik, A. (2023). Tissue heterogeneity of mitochondrial activity, biogenesis and mitochondrial protein gene expression in buffalo. Mol Bio Rep, 50, 5255–5266. https://doi.org/10.1007/s11033-023-08416-2
San-Millán, I. (2023). The key role of mitochondrial function in health and disease. Antioxidants, 12, 782. https://doi.org/10.3390/antiox12040782
Fernández-Vizarra, E., Enríquez, J. A., Pérez-Martos, A., Montoya, J., & Fernández-Silva, P. (2011). Tissue-specific differences in mitochondrial activity and biogenesis. Mitochondrion, 11, 207–213. https://doi.org/10.1016/j.mito.2010.09.011
Mossman, J. A., Biancani, L. M., & Rand, D. M. (2019). Mitochondrial genomic variation drives differential nuclear gene expression in discrete regions of Drosophila gene and protein interaction networks. BMC Genom, 20, 691. https://doi.org/10.1186/s12864-019-6061-y
Bujan, N., Morén, C., García-García, F. J., Blázquez, A., Carnicer, C., Cortés, A. B., González, C., López-Gallardo, E., Lozano, E., Moliner, S., Gort, L., Tobías, E., Delmiro, A., Martin, M. Á., Fernández-Moreno, M. Á., Ruiz-Pesini, E., Garcia-Arumí, E., Rodríguez-Aguilera, J. C., & Garrabou, G. (2022). Multicentric standardization of protocols for the diagnosis of human mitochondrial respiratory chain defects. Antioxidants, 11, 741.
Saki, M., & Prakash, A. (2017). DNA damage related crosstalk between the nucleus and mitochondria. Free rad boil med, 107, 216–227.
Osellame, L. D., Blacker, T. S., & Duchen, M. R. (2012). Cellular and molecular mechanisms of mitochondrial function. Best practice & research. Journal of Clinical Endocrinology and Metabolism, 26, 711–723. https://doi.org/10.1016/j.beem.2012.05.003
Bock, F. J., & Tait, S. W. G. (2020). Mitochondria as multifaceted regulators of cell death. Nature Reviews. Molecular and Cellular Biology, 21, 85–100. https://doi.org/10.1038/s41580-019-0173-8
Rotig, A. (2014). Genetics of mitochondrial respiratory chain deficiencies. Revista de Neurologia, 170, 309–322.
Fox, T. D. (2012). Mitochondrial protein synthesis, import, and assembly. Genetics, 192, 1203–1234.
Weiss, S. L., Cvijanovich, N. Z., Allen, G. L., Thomas, N. J., Freishtat, R. J., Anas, N., Meyer, K., Checchia, P. A., ShanleyTP, B. M. T., Fitzgerald, J., Banschbach, S., Beckman, E., Howard, K., Frank, E., Harmon, K., & Wong, H. R. (2014). Differential expression of the nuclear-encoded mitochondrial transcriptome in pediatric septicshock. Critical Care (London, England), 18, 623. https://doi.org/10.1186/s13054-014-0623-914
Jadiya, P., & Tomar, D. (2020). Mitochondrial protein quality control mechanisms. Genes, 11, 563.
Niemi, N. M., & Pagliarini, D. J. (2021). The extensive and functionally uncharacterized mitochondrial phosphoproteome. Journal of Biological Chemistry, 297, 7. https://doi.org/10.1016/j.jbc.2021.100880
Patel, A. P., Tirosh, I., Trombetta, J. J., Shalek, A. K., Gillespie, S. M., Wakimoto, H., Cahill, D. P., Nahed, B. V., Curry, W. T., Martuza, R. L., Louis, D. N., Rozenblatt-Rosen, O., Suvà, M. L., Regev, A., & Bernstein, B. E. (2014). Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science, 344, 1396–1401. https://doi.org/10.1126/science.1254257
Kolitsida, P., Zhou, J., Rackiewicz, M., Nolic, V., Dengjel, J., & Abeliovich, H. (2019). Phosphorylation of mitochondrial matrix proteins regulates their selective mitophagic degradation. Proceedings of the National academy of Sciences of the United States of America, 116, 20517–20527.
Suomalainen, A., & Battersby, B. J. (2018). Mitochondrial diseases: The contribution of organelle stress responses to pathology. Nature reviews. Mole Cell Biol, 19, 77–92. https://doi.org/10.1038/nrm.2017.66
Sadeesh, E.M., Lahamge, M.S., Malik, A., & Ampadi, A.N. (2023). Nuclear genome-encoded mitochondrial OXPHOS complex I genes in Buffalo show tissue-specific differences. https://doi.org/10.21203/rs.3.rs-3053067/v1
Faccenda, D., & Campanella, M. (2012). Molecular regulation of the mitochondrial F(1) F(o)-ATPsynthase: Physiological and pathological significance of the inhibitory factor 1 (IF(1)). Int J Cell Biol, 2012, 367934. https://doi.org/10.1155/2012/367934
Colina-Tenorio, L., Dautant, A., Miranda-Astudillo, H., Giraud, M. F., & González-Halphen, D. (2018). The peripheral stalk of rotary ATPases. Frontiers in Physiology, 9, 1243.
Bonora, M., Bononi, A., De Marchi, E., Giorgi, C., Lebiedzinska, M., Marchi, S., Patergnani, S., Rimessi, A., Suski, J. M., Wojtala, A., Wieckowski, M. R., Kroemer, G., Galluzzi, L., & Pinton, P. (2013). Role of the c subunit of the FO ATP synthase in mitochondrial permeability transition. Cell Cycle (Georgetown, Tex.), 12, 674–683. https://doi.org/10.4161/cc.23599
Jonckheere, A. I., Smeitink, J. A., & Rodenburg, R. J. (2012). Mitochondrial ATP synthase: Architecture, function and pathology. Journal of Inherited Metabolic Disease, 35, 211–225. https://doi.org/10.1007/s10545-011-9382-9
Garone, C., Pietra, A., & Nesci, S. (2022). From the structural and (Dys) function of ATP synthase to deficiency in age-related diseases. Life (Basel, Switzerland), 12, 401.
Zhou, H., Ren, J., Toan, S., & Mui, D. (2021). Role of mitochondrial quality surveillance in myocardial infarction: From bench to bedside. Ageing Research Reviews, 66, 101250.
Pavlovic, K., Krako Jakovljevic, N., Isakovic, A. M., Ivanovic, T., Markovic, I., & Lalic, N. M. (2022). Therapeutic vs. Suprapharmacological metformin concentrations: Different effects on energy metabolism and mitochondrial function in skeletal muscle cells in vitro. Frontiers in Pharmacology, 13, 930308.
He, J., Ford, H. C., Carroll, J., Ding, S., Fearnley, I. M., & Walker, J. E. (2017). Persistence of the mitochondrial permeability transition in the absence of subunit c of human ATP synthase. Proceedings of the National academy of Sciences of the United States of America, 114, 3409–3414.
Sánchez-Aragó, M., Formentini, L., Martínez-Reyes, I., García-Bermudez, J., Santacatterina, F., Sánchez-Cenizo, L., & Cuezva, J. M. (2013). Expression, regulation and clinical relevance of the ATPase inhibitory factor 1 in human cancers. Oncogenesis, 2, e46–e46.
Vazquez-Martin, A., Corominas-Faja, B., Cufi, S., Vellon, L., Oliveras-Ferraros, C., Menendez, O. J., Joven, J., Lupu, R., & Menendez, J. A. (2013). The mitochondrial H(+)-ATP synthase and the lipogenic switch: New core components of metabolic reprogramming in induced pluripotent stem (iPS) cells. Cell Cycle (Georgetown, Tex.), 12, 207–218.
Davoli, R., Vegni, J., Cesarani, A., Dimauro, C., Zappaterra, M., & Zambonelli, P. (2022). Identification of differentially expressed genes in early-postmortem Semimembranosus muscle of Italian large white heavy pigs divergent for glycolytic potential. Meat Science, 187, 108754.
Sawyer, E. M., Brunner, E. C., Hwang, Y., Ivey, L. E., Brown, O., Bannon, M., Akrobetu, D., Sheaffer, K. E., Morgan, O., Field, C. O., & Suresh, N. (2017). Testis-specific ATP synthase peripheral stalk subunits required for tissue-specific mitochondrial morphogenesis in Drosophila. BMC Cell Biology, 18, 1–15.
Dorji, J., VanderJagt, C. J., Gamer, J. B., Marett, L. C., Mason, B. A., Reich, C. M., Xiang, R., Clark, E. L., Cocks, B. G., Al, C., MacLeod, I. M., & Daetwyler, H. D. (2020). Expression of mitochondrial protein genes encoded by nuclear and mitochondrial genomes correlate with energy metabolism in dairy cattle. BMC Genom, 21, 720.
Hansen, F. M., Kremer, L. S., Karayel, O., Kühl, I., Bludau, I., Larsson, N. G., & Mann, M. (2022). Mitochondrial phosphoproteomes are functionally specialized across tissues. bioRxiv. https://doi.org/10.1101/2022.03.23.485457
Fagerberg, L., Hallström, B. M., Oksvold, P., Kampf, C., Djureinovic, D., Odeberg, J., Habuka, M., Tahmasebpoor, S., Danielsson, A., Edlund, K., & Asplund, A. (2014). Analysis of the human tissue-specific expression by genome-wide integration of transcriptomics and antibody-based proteomics. Molecular & Cellular Proteomics, 13, 397–406.
Oláhová, M., Yoon, W. H., Thompson, K., Jangam, S., Fernandez, L., Davidson, J. M., Kyle, J. E., Grove, M. E., Fisk, D. G., Kohler, J. N., & Holmes, M. (2018). Biallelic mutations in ATP5F1D, which encodes a subunit of ATP synthase, cause a metabolic disorder. American Journal of Human Genetics, 102, 494–504.
Hirono, K., Ichida, F., Nishio, N., Ogawa-Tominaga, M., Fushimi, T., Mayr, J. A., Kohda, M., Kishita, Y., Okazaki, Y., Ohtake, A., & Murayama, K. (2019). Mitochondrial complex deficiency by novel compound heterozygous TMEM70 variants and correlation with developmental delay, undescended testicle, and left ventricular noncompaction in a Japanese patient: A case report. Clinical Case Reports, 7, 553.
Hahn, A., Parey, K., Bublitz, M., Mills, D. J., Zickermann, V., Vonck, J., Kühlbrandt, W., & Meier, T. (2016). Structure of a complete ATP synthase dimer reveals the molecular basis of inner mitochondrial membrane morphology. Molecular Cell, 63, 445–456.
Limongelli, G., Masarone, D., & Pacileo, G. (2017). Mitochondrial disease and the heart. Heart, 103, 390–398.
Marković, A., Tauchmannová, K., Šimáková, M., Mlejnek, P., Kaplanová, V., Pecina, P., Pecinová, A., Papoušek, F., Liška, F., Šilhavý, J., & Mikešová, J. (2022). Genetic complementation of ATP synthase deficiency due to dysfunction of TMEM70 assembly factor in rat. Biomedicines, 10, 276.
Kovalčíkova, J., Vrbacký, M., Pecina, P., Tauchmannová, K., Nůsková, H., Kaplanova, V., Brazdova, A., Alán, L., Eliáš, J., Čunátová, K., & Kořínek, V. (2019). TMEM70 facilitates biogenesis of mammalian ATP synthase by promoting subunit c incorporation into the rotor structure of the enzyme. The FASEB Journal, 33, 14103–14117.
Sánchez-Caballero, L., Elurbe, D. M., Baertling, F., Guerrero-Castillo, S., van den Brand, M., van Strien, J., van Dam, T. J. P., Rodenburg, R., Brandt, U., Huynen, M. A., & Nijtmans, L. G. J. (2020). TMEM70 functions in the assembly of complexes I and V. Biochimica et Biophysica Acta, Bioenergetics, 1861, 148202. https://doi.org/10.1016/j.bbabio.2020.148202
Carroll, J., He, J., Ding, S., Fearnley, I. M., & Walker, J. E. (2021). TMEM70 and TMEM242 help to assemble the rotor ring of human ATP synthase and interact with assembly factors for complex I. Proceedings of the National Academy of Sciences of the United States of America, 118, e2100558118. https://doi.org/10.1073/pnas.2100558118
Bahri, H., Buratto, J., Rojo, M., Dompierre, J. P., Salin, B., Blancard, C., Cuvellier, S., Rose, M., Ben Ammar Elgaaied, A., Tetaud, E., di Rago, J. P., Devin, A., & Duvezin-Caubet, S. (2021). TMEM70 forms oligomeric scaffolds within mitochondrial cristae promoting in situ assembly of mammalian ATP synthase proton channel. Biochimica et Biophysica Acta, Molecular Cell Research, 1868, 118942. https://doi.org/10.1016/j.bbamcr.2020.118942
Vrbacký, M., Kovalčíková, J., Chawengsaksophak, K., Beck, I. M., Mráček, T., Nůsková, H., Sedmera, D., Papoušek, F., Kolář, F., Sobol, M., Hozák, P., Sedlacek, R., & Houštěk, J. (2016). Knockout of Tmem70 alters biogenesis of ATP synthase and leads to embryonal lethality in mice. Human Molecular Genetics, 25, 4674–4685. https://doi.org/10.1093/hmg/ddw295
Galber, C., Fabbian, S., Gatto, C., Grandi, M., Carissimi, S., Acosta, M. J., et al. (2023). The mitochondrial inhibitor IF1 binds to the ATP synthase OSCP subunit and protects cancer cells from apoptosis. Cell Death & Disease, 14, 54.
Pires Da Silva, J., Wargny, M., Raffin, J., Croyal, M., Duparc, T., Combes, G., et al. (2023). Plasma level of ATPase inhibitory factor 1 (IF1) is associated with type 2 diabetes risk in humans: A prospective cohort study. Diabetes & Metabolism, 49, 101391.
Wyant, G. A., Yu, W., Doulamis, I. P., Nomoto, R. S., Saeed, M. Y., Duignan, T., et al. (2022). Mitochondrial remodeling and ischemic protection by G protein-coupled receptor 35 agonists. Science, 377, 621–629.
Zhou, B., Caudal, A., Tang, X., Chavez, J. D., McMillen, T. S., Keller, A., et al. (2022). Upregulation of mitochondrial ATPase inhibitory factor 1 (ATPIF1) mediates increased glycolysis in mouse hearts. The Journal of Clinical Investigation, 132, e155333.
Domínguez-Zorita, S., Romero-Carramiñana, I., Santacatterina, F., et al. (2023). IF1 ablation prevents ATP synthase oligomerization, enhances mitochondrial ATP turnover and promotes an adenosine-mediated pro-inflammatory phenotype. Cell Death & Disease, 14, 413. https://doi.org/10.1038/s41419-023-05957-z
Esparza-Moltó, P. B., Nuevo-Tapioles, C., Chamorro, M., Näjera, L., Torresano, L., Santacatterina, F., & Cuezva, J. M. (2019). Tissue-specific expression and post-transcriptional regulation of the ATPase inhibitory factor 1 (IF1) in human and mouse tissues. The FASEB Journal, 33, 1836–1851.
Viscomi, C., & Zeviani, M. (2020). Strategies for fighting mitochondrial diseases. Journal of Internal Medicine, 287, 665–684.
Acknowledgements
We extend our appreciation to the Science and Engineering Research Board, a part of the Department of Science and Technology under the Government of India, for financially supporting this endeavour. The authors wish to convey their thanks to the Director of ICAR-National Dairy Research Institute for enabling the essential resources essential to carry out this study.
Funding
This study was financially supported by the Science and Engineering Research Board, Department of Science and Technology, Government of India.
Author information
Authors and Affiliations
Contributions
The project’s conception and experimental design were spearheaded by SEM. SEM executed the experiments, conducted data analysis, and initiated manuscript drafting alongside MSL and AM. AAN contributed to the preparation of tables and figures. All authors critically reviewed and granted their approval for the final manuscript version.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethical Approval
No endorsement from research ethics committees was deemed necessary to achieve the objectives of this study, as the experimental procedures were carried out using post-slaughter tissues of buffalo specimens.
Informed Consent
Not applicable.
Consent to Participate
This article does not contain any individual person’s data in any form.
Consent for Publication
All authors agreed to publish the research in this journal.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sadeesh, E.M., Lahamge, M.S., Malik, A. et al. Differential Expression of Nuclear-Encoded Mitochondrial Protein Genes of ATP Synthase Across Different Tissues of Female Buffalo. Mol Biotechnol (2024). https://doi.org/10.1007/s12033-024-01085-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12033-024-01085-x