Abstract
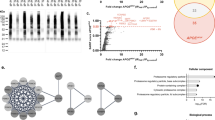
Lysine-based post-translational modification (PTM) such as acylation, acetylation, deamination, methylation, SUMOylation, and ubiquitination has proven to be a major regulator of gene expression, chromatin structure, protein stability, protein–protein interaction, protein degradation, and cellular localization. However, besides all the PTMs, ubiquitination stands as the second most common PTM after phosphorylation that is involved in the etiology of neurodegenerative diseases (NDDs) namely, Alzheimer’s disease (AD) and Parkinson’s disease (PD). NDDs are characterized by the accumulation of misfolded protein aggregates in the brain that lead to disease-related gene mutation and irregular protein homeostasis. The ubiquitin–proteasome system (UPS) is in charge of degrading these misfolded proteins, which involve an interplay of E1, E2, E3, and deubiquitinase enzymes. Impaired UPS has been commonly observed in NDDs and E3 ligases are the key members of the UPS, thus, dysfunction of the same can accelerate the neurodegeneration process. Therefore, the aim of this study is firstly, to find E3 ligases that are common in both AD and PD through data mining. Secondly, to study the impact of mutation on its structure and function. The study deciphered 74 E3 ligases that were common in both AD and PD. Later, 10 hub genes were calculated of which protein–protein interaction, pathway enrichment, lysine site prediction, domain, and motif analysis were performed. The results predicted BRCA1, PML, and TRIM33 as the top three putative lysine-modified E3 ligases involved in AD and PD pathogenesis. However, based on structural characterization, BRCA1 was taken further to study RING domain mutation that inferred K32Y, K32L, K32C, K45V, K45Y, and K45G as potential mutants that alter the structural and functional ability of BRCA1 to interact with Ube2k, E2-conjugating enzyme. The most probable mutant observed after molecular dynamics simulation of 50 ns is K32L. Therefore, our study concludes BRCA1, a potential E3 ligase common in AD and PD, and RING domain mutation at sites K32 and K45 possibly disturbs its interaction with its E2, Ube2k.
Graphical Abstract
Graphical representation of all the steps involved to study mutation in the RING domain of BRCA1 which is a common E3 ligase observed in Alzheimer’s disease and Parkinson’s disease.








Similar content being viewed by others
Data Availability
The authors confirm that the data supporting the findings of this study are available within the article.
References
Watanabe, Y., Taguchi, K., & Tanaka, M. (2020). Ubiquitin autophagy and neurodegenerative diseases. Cells, 9(9), 2022. https://doi.org/10.3390/CELLS9092022
Chen, L., Liu, S., & Tao, Y. (2020). Regulating tumor suppressor genes: Post-translational modifications. Signal Transduction and Targeted Therapy, 5(1), 1–25. https://doi.org/10.1038/s41392-020-0196-9
Salas-Lloret, D., & González-Prieto, R. (2022). Insights in Post-translational modifications: Ubiquitin and SUMO. International Journal of Molecular Sciences, 23(6), 3281. https://doi.org/10.3390/IJMS23063281
Kumar, V. S., & Vellaichamy, A. (2019). Sequence and structure-based characterization of ubiquitination sites in human and yeast proteins using Chou’s sample formulation. Proteins: Structure, Function, and Bioinformatics, 87(8), 646–657. https://doi.org/10.1002/PROT.25689
Jaishankar, A., Tang, N. L. S., Jelani, M., Mabb, A. M., George, A. J., Hoffiz, Y. C., & Zhu, Y. (2018). A comprehensive atlas of E3 ubiquitin ligase mutations in neurological disorders. Frontiers in Genetics, 9, 29. https://doi.org/10.3389/fgene.2018.00029
Upadhyay, A., Joshi, V., Amanullah, A., Mishra, R., Arora, N., Prasad, A., & Mishra, A. (2017). E3 ubiquitin ligases neurobiological mechanisms: Development to degeneration. Frontiers in Molecular Neuroscience, 10, 151. https://doi.org/10.3389/FNMOL.2017.00151/BIBTEX
LeBlanc, N., Mallette, E., & Zhang, W. (2021). Targeted modulation of E3 ligases using engineered ubiquitin variants. The FEBS Journal, 288(7), 2143–2165. https://doi.org/10.1111/FEBS.15536
Yang, Q., Zhao, J., Chen, D., & Wang, Y. (2021). E3 ubiquitin ligases: Styles, structures and functions. Molecular Biomedicine. https://doi.org/10.1186/S43556-021-00043-2
Schmidt, M. F., Gan, Z. Y., Komander, D., & Dewson, G. (2021). Ubiquitin signalling in neurodegeneration: Mechanisms and therapeutic opportunities. Cell Death & Differentiation, 28(2), 570–590. https://doi.org/10.1038/s41418-020-00706-7
Roverato, N. D., Sailer, C., Catone, N., Aichem, A., Stengel, F., & Groettrup, M. (2021). Parkin is an E3 ligase for the ubiquitin-like modifier FAT10, which inhibits Parkin activation and mitophagy. Cell reports. https://doi.org/10.1016/J.CELREP.2021.108857
Wang, T., Wang, W., Wang, Q., Xie, R., Landay, A., & Chen, D. (2020). The E3 ubiquitin ligase CHIP in normal cell function and in disease conditions. Annals of the New York Academy of Sciences, 1460(1), 3–10. https://doi.org/10.1111/NYAS.14206
Metzger, M. B., Pruneda, J. N., Klevit, R. E., & Weissman, A. M. (2014). RING-type E3 ligases: Master manipulators of E2 ubiquitin-conjugating enzymes and ubiquitination. Biophysica Acta (BBA) - Molecular Cell Research, 1843(1), 47–60. https://doi.org/10.1016/J.BBAMCR.2013.05.026
Leung, E., & Hazrati, L. N. (2021). Breast cancer type 1 and neurodegeneration: Consequences of deficient DNA repair. Brain Communications. https://doi.org/10.1093/BRAINCOMMS/FCAB117
Gough, C. A., Gojobori, T., & Imanishi, T. (2007). Cancer-related mutations in BRCA1-BRCT cause long-range structural changes in protein–protein binding sites: A molecular dynamics study. Proteins: Structure, Function, and Bioinformatics, 66(1), 69–86. https://doi.org/10.1002/PROT.21188
Drikos, I., Nounesis, G., & Vorgias, C. E. (2009). Characterization of cancer-linked BRCA1-BRCT missense variants and their interaction with phosphoprotein targets. Proteins: Structure, Function, and Bioinformatics, 77(2), 464–476. https://doi.org/10.1002/PROT.22460
Christensen, D. E., Brzovic, P. S., & Klevit, R. E. (2007). E2–BRCA1 RING interactions dictate synthesis of mono- or specific polyubiquitin chain linkages. Nature Structural & Molecular Biology, 14(10), 941–948. https://doi.org/10.1038/nsmb1295
Song, S., Kim, S. Y., Hong, Y. M., Jo, D. G., Lee, J. Y., Shim, S. M., Chung, C. W., Seo, S. J., Yoo, Y. J., Koh, J. Y., Lee, M. C., Yates, A. J., Ichijo, H., & Jung, Y. K. (2003). Essential role of E2–25K/Hip-2 in mediating amyloid-beta neurotoxicity. Molecular Cell, 12(3), 553–563. https://doi.org/10.1016/J.MOLCEL.2003.08.005
de Pril, R., Fischer, D. F., Roos, R. A. C., & van Leeuwen, F. W. (2007). Ubiquitin-conjugating enzyme E2–25K increases aggregate formation and cell death in polyglutamine diseases. Molecular and cellular neurosciences, 34(1), 10–19. https://doi.org/10.1016/J.MCN.2006.09.006
Su, J., Huang, P., Qin, M., Lu, Q., Sang, X., Cai, Y., Wang, Y., Liu, F., Wu, R., Wang, X., Jiang, X., Wang, J., Sun, Q., Chen, S., & Xu, J. (2018). Reduction of HIP2 expression causes motor function impairment and increased vulnerability to dopaminergic degeneration in Parkinson’s disease models. Cell Death and Disease. https://doi.org/10.1038/s41419-018-1066-z
Tak, Y. J., & Kang, S. (2022). The E2 ubiquitin-conjugating enzyme HIP2 is a crucial regulator of quality control against mutant SOD1 proteotoxicity. Biochimica et Biophysica Acta (BBA) - Molecular Basis of Disease, 1868(2), 166316. https://doi.org/10.1016/J.BBADIS.2021.166316
Mezencev, R., & Chernoff, Y. O. (2020). Risk of Alzheimer’s disease in cancer patients: Analysis of mortality data from the US SEER population-based registries. Cancers, 12(4), 796. https://doi.org/10.3390/CANCERS12040796
Mano, T., Iwata, A., Nonaka, T., Tarutani, A., Hashimoto, T., Hasegawa, M., Iwatsubo, T., & Toda, T. (2018). Tau-related dysfunction of BRCA1 leads to reduced neuronal plasticity in Alzheimer’s disease. Alzheimer’s & Dementia. https://doi.org/10.1016/J.JALZ.2018.06.2771
Evans, T. A., Raina, A. K., Delacourte, A., Aprelikova, O., Lee, H. G., Zhu, X., Perry, G., & Smith, M. A. (2007). BRCA1 may modulate neuronal cell cycle re-entry in alzheimer disease. International Journal of Medical Sciences, 4(3), 140. https://doi.org/10.7150/IJMS.4.140
Goodsell, D. S. (2022). JCVI-syn3A minimal cell. RCSB Protein Data Bank. https://doi.org/10.2210/RCSB_PDB/GOODSELL-GALLERY-042
Barrett, T., Wilhite, S. E., Ledoux, P., Evangelista, C., Kim, I. F., Tomashevsky, M., Marshall, K. A., Phillippy, K. H., Sherman, P. M., Holko, M., Yefanov, A., Lee, H., Zhang, N., Robertson, C. L., Serova, N., Davis, S., & Soboleva, A. (2013). NCBI GEO: Archive for functional genomics data sets–update. Nucleic Acids Research. https://doi.org/10.1093/NAR/GKS1193
Edgar, R., Domrachev, M., & Lash, A. E. (2002). Gene expression omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Research, 30(1), 207–210. https://doi.org/10.1093/NAR/30.1.207
Hunt, G. P., Grassi, L., Henkin, R., Smeraldi, F., Spargo, T. P., Kabiljo, R., Koks, S., Ibrahim, Z., Dobson, R. J. B., Al-Chalabi, A., Barnes, M. R., & Iacoangeli, A. (2022). GEOexplorer: a webserver for gene expression analysis and visualisation. Nucleic Acids Research, 50(W1), W367–W374. https://doi.org/10.1093/NAR/GKAC364
Udhaya Kumar, S., Thirumal Kumar, D., Bithia, R., Sankar, S., Magesh, R., Sidenna, M., George Priya Doss, C., & Zayed, H. (2020). Analysis of differentially expressed genes and molecular pathways in familial hypercholesterolemia involved in atherosclerosis: A systematic and bioinformatics approach. Frontiers in Genetics, 11, 1–16. https://doi.org/10.3389/FGENE.2020.00734/BIBTEX
Wu, A. T. H., Lawal, B., Wei, L., Wen, Y. T., Tzeng, D. T. W., & Lo, W. C. (2021). Multiomics identification of potential targets for alzheimer disease and antrocin as a therapeutic candidate. Pharmaceutics, 13(10), 1555. https://doi.org/10.3390/PHARMACEUTICS13101555
Li, W., Bengtson, M. H., Ulbrich, A., Matsuda, A., Reddy, V. A., Orth, A., Chanda, S. K., Batalov, S., & Joazeiro, C. A. (2008). Genome-wide and functional annotation of human E3 ubiquitin ligases identifies MULAN, a mitochondrial e3 that regulates the organelle’s dynamics and signaling. PLOS ONE, 3(1), e1487. https://doi.org/10.1371/JOURNAL.PONE.0001487
Heberle, H., Meirelles, V. G., da Silva, F. R., Telles, G. P., & Minghim, R. (2015). InteractiVenn: A web-based tool for the analysis of sets through Venn diagrams. BMC Bioinformatics, 16(1), 1–7. https://doi.org/10.1186/S12859-015-0611-3/FIGURES/4
Szklarczyk, D., Franceschini, A., Wyder, S., Forslund, K., Heller, D., Huerta-Cepas, J., Simonovic, M., Roth, A., Santos, A., Tsafou, K. P., Kuhn, M., Bork, P., Jensen, L. J., & von Mering, C. (2015). STRING v10: Protein-protein interaction networks, integrated over the tree of life. Nucleic Acids Research, 43(D1), D447–D452. https://doi.org/10.1093/nar/gku1003
Oughtred, R., Chatr-aryamontri, A., Breitkreutz, B. J., Chang, C. S., Rust, J. M., Theesfeld, C. L., Heinicke, S., Breitkreutz, A., Chen, D., Hirschman, J., Kolas, N., Livstone, M. S., Nixon, J., O’Donnell, L., Ramage, L., Winter, A., Reguly, T., Sellam, A., Stark, C., … Tyers, M. (2016). BioGRID: A tool for studying biological interactions in yeast. Cold Spring Harbor protocols, 2016(1), pdb.top080754. https://doi.org/10.1101/PDB.TOP080754
Szklarczyk, D., Gable, A. L., Nastou, K. C., Lyon, D., Kirsch, R., Pyysalo, S., Doncheva, N. T., Legeay, M., Fang, T., Bork, P., Jensen, L. J., & von Mering, C. (2021). The STRING database in 2021: customizable protein–protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Research, 49(D1), D605–D612. https://doi.org/10.1093/NAR/GKAA1074
Cao, M., Pietras, C. M., Feng, X., Doroschak, K. J., Schaffner, T., Park, J., Zhang, H., Cowen, L. J., & Hescott, B. J. (2014). New directions for diffusion-based network prediction of protein function incorporating pathways with confidenc. Bioinformatics (Oxford, England). https://doi.org/10.1093/BIOINFORMATICS/BTU263
Cao, M., Zhang, H., Park, J., Daniels, N. M., Crovella, M. E., Cowen, L. J., & Hescott, B. (2013). Going the distance for protein function prediction: A new distance metric for protein interaction networks. PLoS ONE. https://doi.org/10.1371/JOURNAL.PONE.0076339
Shannon, P., Markiel, A., Ozier, O., Baliga, N. S., Wang, J. T., Ramage, D., Amin, N., Schwikowski, B., & Ideker, T. (2003). Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Research, 13(11), 2498. https://doi.org/10.1101/GR.1239303
Elbashir, M. K., Mohammed, M., Mwambi, H., & Omolo, B. (2023). Identification of hub genes associated with breast cancer using integrated gene expression data with protein-protein interaction network. Applied Sciences (Switzerland), 13(4), 2403. https://doi.org/10.3390/APP13042403/S1
Reimand, J., Arak, T., & Vilo, J. (2011). g:Profiler—a web server for functional interpretation of gene lists (2011 update). Nucleic Acids Research, 39(2), W307–W315. https://doi.org/10.1093/NAR/GKR378
Xie, Z., Bailey, A., Kuleshov, M. V., Clarke, D. J. B., Evangelista, J. E., Jenkins, S. L., Lachmann, A., Wojciechowicz, M. L., Kropiwnicki, E., Jagodnik, K. M., Jeon, M., & Ma’ayan, A. (2021). Gene set knowledge discovery with enrichr. Current Protocols, 1(3), e90. https://doi.org/10.1002/CPZ1.90
Ge, S. X., Jung, D., Jung, D., & Yao, R. (2020). ShinyGO: A graphical gene-set enrichment tool for animals and plants. Bioinformatics, 36(8), 2628–2629. https://doi.org/10.1093/BIOINFORMATICS/BTZ931
Letunic, I., & Bork, P. (2018). 20 years of the SMART protein domain annotation resource. Nucleic Acids Research, 46(D1), D493–D496. https://doi.org/10.1093/NAR/GKX922
Bailey, T. L., Johnson, J., Grant, C. E., & Noble, W. S. (2015). The MEME suite. Nucleic Acids Research, 43(W1), W39–W49. https://doi.org/10.1093/NAR/GKV416
Obenauer, J. C., Cantley, L. C., & Yaffe, M. B. (2003). Scansite 20: Proteome-wide prediction of cell signaling interactions using short sequence motifs. Nucleic Acids Research, 31(13), 3635–3641. https://doi.org/10.1093/NAR/GKG584
Ashkenazy, H., Abadi, S., Martz, E., Chay, O., Mayrose, I., Pupko, T., & Ben-Tal, N. (2016). ConSurf 2016: An improved methodology to estimate and visualize evolutionary conservation in macromolecules. Nucleic Acids Research. https://doi.org/10.1093/nar/gkw408
Xu, H., Zhou, J., Lin, S., Deng, W., Zhang, Y., & Xue, Y. (2017). PLMD: An updated data resource of protein lysine modifications. Journal of Genetics and Genomics = Yi chuan xue bao, 44(5), 243–250. https://doi.org/10.1016/J.JGG.2017.03.007
Bendl, J., Stourac, J., Salanda, O., Pavelka, A., Wieben, E. D., Zendulka, J., Brezovsky, J., & Damborsky, J. (2014). PredictSNP: Robust and accurate consensus classifier for prediction of disease-related mutations. PLoS Computational Biology. https://doi.org/10.1371/JOURNAL.PCBI.1003440
Pejaver, V., Urresti, J., Lugo-Martinez, J., Pagel, K. A., Lin, G. N., Nam, H. J., Mort, M., Cooper, D. N., Sebat, J., Iakoucheva, L. M., Mooney, S. D., & Radivojac, P. (2020). Inferring the molecular and phenotypic impact of amino acid variants with MutPred2. Nature Communications. https://doi.org/10.1038/S41467-020-19669-X
Venselaar, H., te Beek, T. A. H., Kuipers, R. K. P., Hekkelman, M. L., & Vriend, G. (2010). Protein structure analysis of mutations causing inheritable diseases. An e-Science approach with life scientist friendly interfaces. BMC Bioinformatics, 11(1), 1–10. https://doi.org/10.1186/1471-2105-11-548/FIGURES/5
Brzovic, P. S., Rajagopal, P., Hoyt, D. W., King, M. C., & Klevit, R. E. (2001). Structure of a BRCA1-BARD1 heterodimeric RING-RING complex. Nature Structural Biology, 8(10), 833–837. https://doi.org/10.1038/nsb1001-833
Burley, S. K., Bhikadiya, C., Bi, C., Bittrich, S., Chao, H., Chen, L., Craig, P. A., Crichlow, G. V., Dalenberg, K., Duarte, J. M., Dutta, S., Fayazi, M., Feng, Z., Flatt, J. W., Ganesan, S., Ghosh, S., Goodsell, D. S., Green, R. K., Guranovic, V., … Zardecki, C. (2023). RCSB Protein Data Bank (RCSB.org): delivery of experimentally-determined PDB structures alongside one million computed structure models of proteins from artificial intelligence/machine learning. Nucleic Acids Research, 51(D1), D488–D508. https://doi.org/10.1093/NAR/GKAC1077
Kozakov, D., Grove, L. E., Hall, D. R., Bohnuud, T., Mottarella, S. E., Luo, L., Xia, B., Beglov, D., & Vajda, S. (2015). The FTMap family of web servers for determining and characterizing ligand-binding hot spots of proteins. Nature Protocols, 10(5), 733–755. https://doi.org/10.1038/nprot.2015.043
de Vries, S. J., & Bonvin, A. M. J. J. (2011). CPORT: A consensus interface predictor and its performance in prediction-driven docking with HADDOCK. PLoS ONE. https://doi.org/10.1371/JOURNAL.PONE.0017695
Desta, I. T., Porter, K. A., Xia, B., Kozakov, D., & Vajda, S. (2020). Performance and its limits in rigid body protein-protein docking. Structure, 28(9), 1071-1081.e3. https://doi.org/10.1016/j.str.2020.06.006
Vajda, S., Yueh, C., Beglov, D., Bohnuud, T., Mottarella, S. E., Xia, B., Hall, D. R., & Kozakov, D. (2017). New additions to the ClusPro server motivated by CAPRI. Proteins: Structure Function, and Bioinformatics. https://doi.org/10.1002/prot.25219
Kozakov, D., Beglov, D., Bohnuud, T., Mottarella, S. E., Xia, B., Hall, D. R., & Vajda, S. (2013). How good is automated protein docking? Proteins: Structure Function, and Bioinformatics, 81(12), 2159–2166. https://doi.org/10.1002/prot.24403
Kozakov, D., Hall, D. R., Xia, B., Porter, K. A., Padhorny, D., Yueh, C., Beglov, D., & Vajda, S. (2017). The ClusPro web server for protein–protein docking. Nature Protocols, 12(2), 255–278. https://doi.org/10.1038/nprot.2016.169
Porter, K. A., Xia, B., Beglov, D., Bohnuud, T., Alam, N., Schueler-Furman, O., & Kozakov, D. (2017). ClusPro peptidock: Efficient global docking of peptide recognition motifs using FFT. Bioinformatics, 33(20), 3299–3301. https://doi.org/10.1093/bioinformatics/btx216
van Zundert, G. C. P., Rodrigues, J. P. G. L. M., Trellet, M., Schmitz, C., Kastritis, P. L., Karaca, E., Melquiond, A. S. J., van Dijk, M., de Vries, S. J., & Bonvin, A. M. J. J. (2016). The HADDOCK2.2 web server: User-friendly integrative modeling of biomolecular complexes. Journal of Molecular Biology, 428(4), 720–725. https://doi.org/10.1016/J.JMB.2015.09.014
Dominguez, C., Boelens, R., & Bonvin, A. M. J. J. (2003). HADDOCK: A protein-protein docking approach based on biochemical or biophysical information. Journal of the American Chemical Society, 125(7), 1731–1737. https://doi.org/10.1021/JA026939X/SUPPL_FILE/JA026939XSI20021128_085857.TXT
Venkatraman, V., Yang, Y. D., Sael, L., & Kihara, D. (2009). Protein-protein docking using region-based 3D Zernike descriptors. BMC Bioinformatics. https://doi.org/10.1186/1471-2105-10-407
Porter, K. A., Desta, I., Kozakov, D., & Vajda, S. (2019). What method to use for protein–protein docking? Current Opinion in Structural Biology, 55, 1–7. https://doi.org/10.1016/j.sbi.2018.12.010
Van Der Spoel, D., Lindahl, E., Hess, B., Groenhof, G., Mark, A. E., & Berendsen, H. J. C. (2005). GROMACS: Fast, flexible, and free. Journal of Computational Chemistry, 26(16), 1701–1718. https://doi.org/10.1002/JCC.20291
Robertson, M. J., Tirado-Rives, J., & Jorgensen, W. L. (2015). Improved peptide and protein torsional energetics with the OPLS-AA force field. Journal of Chemical Theory and Computation, 11(7), 3499–3509. https://doi.org/10.1021/acs.jctc.5b00356
Scardoni, G., Tosadori, G., Faizan, M., Spoto, F., Fabbri, F., & Laudanna, C. (2014). Biological network analysis with CentiScaPe: Centralities and experimental dataset integration. F1000Research. https://doi.org/10.12688/F1000RESEARCH.4477.2
Chin, C. H., Chen, S. H., Wu, H. H., Ho, C. W., Ko, M. T., & Lin, C. Y. (2014). cytoHubba: Identifying hub objects and sub-networks from complex interactome. BMC Systems Biology, 8(4), 1–7. https://doi.org/10.1186/1752-0509-8-S4-S11/TABLES/4
Lorick, K. L., Jensen, J. P., Fang, S., Ong, A. M., Hatakeyama, S., & Weissman, A. M. (1999). RING fingers mediate ubiquitin-conjugating enzyme (E2)-dependent ubiquitination. Proceedings of the National Academy of Sciences of the United States of America, 96(20), 11364. https://doi.org/10.1073/PNAS.96.20.11364
Ying, M., Huang, X., Zhao, H., Wu, Y., Wan, F., Huang, C., & Jie, K. (2011). Comprehensively surveying structure and function of RING domains from Drosophila melanogaster. PLoS ONE, 6(9), e23863. https://doi.org/10.1371/JOURNAL.PONE.0023863
Ashkenazy, H., Abadi, S., Martz, E., Chay, O., Mayrose, I., Pupko, T., & Ben-Tal, N. (2016). ConSurf 2016: An improved methodology to estimate and visualize evolutionary conservation in macromolecules. Nucleic Acids Research, 44, W344. https://doi.org/10.1093/NAR/GKW408
Le Guilloux, V., Schmidtke, P., & Tuffery, P. (2009). Fpocket: An open source platform for ligand pocket detection. BMC Bioinformatics, 10(1), 1–11. https://doi.org/10.1186/1471-2105-10-168/TABLES/1
Hsu, K. S., & Kao, H. Y. (2018). PML: Regulation and multifaceted function beyond tumor suppression. Cell & Bioscience, 8(1), 1–21. https://doi.org/10.1186/S13578-018-0204-8
Tessier, S., Ferhi, O., Geoffroy, M. C., González-Prieto, R., Canat, A., Quentin, S., Pla, M., Niwa-Kawakita, M., Bercier, P., Rérolle, D., Tirard, M., Therizols, P., Fabre, E., Vertegaal, A. C. O., de Thé, H., & Lallemand-Breitenbach, V. (2022). Exploration of nuclear body-enhanced sumoylation reveals that PML represses 2-cell features of embryonic stem cells. Nature Communications, 13(1), 1–15. https://doi.org/10.1038/s41467-022-33147-6
Ivanschitz, L., De Thé, H., & Le Bras, M. (2013). PML, SUMOylation, and senescence. Frontiers in Oncology. https://doi.org/10.3389/FONC.2013.00171
Rabellino, A., & Scaglioni, P. P. (2013). PML degradation: Multiple ways to eliminate PML. Frontiers in Oncology. https://doi.org/10.3389/FONC.2013.00060/BIBTEX
Clark, S. L., Rodriguez, A. M., Snyder, R. R., Hankins, G. D. V., & Boehning, D. (2012). Structure-function of the tumor suppressor BRCA1. Computational and Structural Biotechnology Journal, 1(1), e201204005. https://doi.org/10.5936/CSBJ.201204005
Chatterjee, P., Karn, R., Emerson, I. A., & Banerjee, S. (2023). Docking and molecular dynamics simulation revealed the potential inhibitory activity of amygdalin in triple-negative breast cancer therapeutics targeting the brct domain of BARD1 receptor. Molecular Biotechnology, 2023, 1–19. https://doi.org/10.1007/S12033-023-00680-8
Gouveia, C., Peña-Guerrero, J., Fernández-Rubio, C., García-Sosa, A. T., & Nguewa, P. A. (2023). BRCT domains: Structure, functions, and implications in disease—New therapeutic targets for innovative drug discovery against infections. Pharmaceutics, 15(7), 1839. https://doi.org/10.3390/PHARMACEUTICS15071839
Witus, S. R., Stewart, M. D., & Klevit, R. E. (2021). The BRCA1/BARD1 ubiquitin ligase and its substrates. The Biochemical Journal, 478(18), 3467. https://doi.org/10.1042/BCJ20200864
Pejaver, V., Urresti, J., Lugo-Martinez, J., Pagel, K. A., Lin, G. N., Nam, H. J., Mort, M., Cooper, D. N., Sebat, J., Iakoucheva, L. M., Mooney, S. D., & Radivojac, P. (2020). Inferring the molecular and phenotypic impact of amino acid variants with MutPred2. Nature Communications, 11(1), 1–13. https://doi.org/10.1038/s41467-020-19669-x
Wu, W., Koike, A., Takeshita, T., & Ohta, T. (2008). The ubiquitin E3 ligase activity of BRCA1 and its biological functions. Cell Division, 3(1), 1–10. https://doi.org/10.1186/1747-1028-3-1/FIGURES/2
(2007). New partners for BRCA1. Nature Structural & Molecular Biology, 14(10), 879–879. https://doi.org/10.1038/nsmb1007-879
Merkley, N., & Shaw, G. S. (2004). Solution structure of the flexible class II ubiquitin-conjugating enzyme Ubc1 provides insights for polyubiquitin chain assembly. The Journal of Biological Chemistry, 279(45), 47139–47147. https://doi.org/10.1074/JBC.M409576200
Middleton, A. J., Teyra, J., Zhu, J., Sidhu, S. S., & Day, C. L. (2021). Identification of ubiquitin variants that inhibit the E2 ubiquitin conjugating enzyme, Ube2k. ACS Chemical Biology, 16(9), 1745–1756. https://doi.org/10.1021/ACSCHEMBIO.1C00445/ASSET/IMAGES/LARGE/CB1C00445_0006.JPEG
Kalchman, M. A., Graham, R. K., Xia, G., Koide, H. B., Hodgson, J. G., Graham, K. C., Goldberg, Y. P., Gietz, R. D., Pickart, C. M., & Hayden, M. R. (1996). Huntingtin is ubiquitinated and interacts with a specific ubiquitin-conjugating enzyme. The Journal of Biological Chemistry, 271(32), 19385–19394. https://doi.org/10.1074/JBC.271.32.19385
Meiklejohn, H., Mostaid, M. S., Luza, S., Mancuso, S. G., Kang, D., Atherton, S., Rothmond, D. A., Weickert, C. S., Opazo, C. M., Pantelis, C., Bush, A. I., Everall, I. P., & Bousman, C. A. (2019). Blood and brain protein levels of ubiquitin-conjugating enzyme E2K (UBE2K) are elevated in individuals with schizophrenia. Journal of Psychiatric Research, 113, 51–57. https://doi.org/10.1016/J.JPSYCHIRES.2019.03.005
Su, J., Huang, P., Qin, M., Lu, Q., Sang, X., Cai, Y., Wang, Y., Liu, F., Wu, R., Wang, X., Jiang, X., Wang, J., Sun, Q., Chen, S., & Xu, J. (2018). Reduction of HIP2 expression causes motor function impairment and increased vulnerability to dopaminergic degeneration in Parkinson’s disease models. Cell Death & Disease. https://doi.org/10.1038/S41419-018-1066-Z
Gupta, R., Sahu, M., Srivastava, D., Tiwari, S., Ambasta, R. K., & Kumar, P. (2021). Post-translational modifications: Regulators of neurodegenerative proteinopathies. Ageing Research Reviews. https://doi.org/10.1016/j.arr.2021.101336
Liu, W., Tang, X., Qi, X., Fu, X., Ghimire, S., Ma, R., Li, S., Zhang, N., & Si, H. (2020). The ubiquitin conjugating enzyme: An important ubiquitin transfer platform in ubiquitin-proteasome system. International Journal of Molecular Sciences. https://doi.org/10.3390/IJMS21082894
Radivojac, P., Vacic, V., Haynes, C., Cocklin, R. R., Mohan, A., Heyen, J. W., Goebl, M. G., & Iakoucheva, L. M. (2010). Identification, analysis, and prediction of protein ubiquitination sites. Proteins: Structure, Function, and Bioinformatics, 78(2), 365–380. https://doi.org/10.1002/PROT.22555
Chen, R. H., Lee, Y. R., & Yuan, W. C. (2012). The role of PML ubiquitination in human malignancies. Journal of biomedical science, 19(1), 81. https://doi.org/10.1186/1423-0127-19-81/FIGURES/1
Miyahara, K., Takano, N., Yamada, Y., Kazama, H., Tokuhisa, M., Hino, H., Fujita, K., Barroga, E., Hiramoto, M., Handa, H., Kuroda, M., Ishikawa, T., & Miyazawa, K. (2021). BRCA1 degradation in response to mitochondrial damage in breast cancer cells. Scientific Reports, 11(1), 1–13. https://doi.org/10.1038/s41598-021-87698-7
Kurihara, M., Mano, T., Saito, Y., Murayama, S., Toda, T., & Iwata, A. (2020). Colocalization of BRCA1 with tau aggregates in human tauopathies. Brain Sciences. https://doi.org/10.3390/BRAINSCI10010007
Clark, K. A., Paquette, A., Tao, K., Bell, R., Boyle, J. L., Rosenthal, J., Snow, A. K., Stark, A. W., Thompson, B. A., Unger, J., Gertz, J., Varley, K. E., Boucher, K. M., Goldgar, D. E., Foulkes, W. D., Thomas, A., & Tavtigian, S. V. (2022). Comprehensive evaluation and efficient classification of BRCA1 RING domain missense substitutions. American Journal of Human Genetics, 109(6), 1153. https://doi.org/10.1016/J.AJHG.2022.05.004
Brzovic, P. S., Meza, J. E., King, M. C., & Klevit, R. E. (2001). BRCA1 RING domain cancer-predisposing mutations. Structural consequences and effects on protein-protein interactions. The Journal of Biological Chemistry, 276(44), 41399–41406. https://doi.org/10.1074/JBC.M106551200
Wezyk, M., & Zekanowski, C. (2018). Role of BRCA1 in neuronal death in Alzheimer’s disease. ACS Chemical Neuroscience. https://doi.org/10.1021/acschemneuro.8b00149
Acknowledgements
We would like to thank the senior management of Delhi Technological University for their constant support and guidance. The authors would like to thank the University Grant Commission, Government of India, for providing junior research fellowship to NR, NTA ref no 201610125084.
Funding
Not applicable.
Author information
Authors and Affiliations
Contributions
PK: conceptualized, designed, and supervised this work. MS and NR: drafted the manuscript, analyzed, and interpreted the data, and did the artwork. All authors have read and given consent for submission.
Corresponding author
Ethics declarations
Competing interest
The author declared no potential competing interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval and Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sahu, M., Rani, N. & Kumar, P. Simulation and Computational Study of RING Domain Mutants of BRCA1 and Ube2k in AD/PD Pathophysiology. Mol Biotechnol 66, 1095–1115 (2024). https://doi.org/10.1007/s12033-023-01006-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-023-01006-4