Abstract
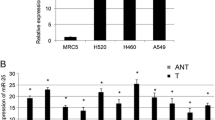
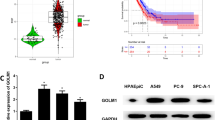
Lung adenocarcinoma (LUAD) is the most frequent histological subtype of non-small cell lung cancer. Cyclin B1 (CCNB1) is the vital initiator and controller of mitosis. Studies have indicated that CCNB1 overexpression is closely associated with cell proliferation and tumorigenesis in many cancers. Thus, discovery of molecular mechanism of CCNB1 in LUAD is conducive to developing new diagnostic or therapeutic targets for LUAD. We acquired mature miRNA and mRNA expression information of LUAD from TCGA database, as well as related clinical data. CCNB1 expression in normal and LUAD tissue was analyzed. Relationship between CCNB1 and patient’s survival and clinical stage was analyzed. Upstream regulatory gene miRNA of CCNB1 was predicted. qRT-PCR and western blot examined expression levels of CCNB1 and miR-139-5p in cells. CCK-8 tested cell proliferation. Scratch healing and Transwell determined cell migration and invasion. Flow cytometry analyzed the cell cycle. Dual-luciferase verified targeting relationship between the two genes. Compared to controls, CCNB1 expression was prominently high in LUAD patient samples, and associated with advanced tumor stages and shorter overall survival. MiR-139-5p expressed an evidently negative correlation with CCNB1 and was predicted to target CCNB1. MiR-139-5p mimics reduced CCNB1 mRNA and protein expression, and suppressed luciferase activity in a target-specific manner, as confirmed by a control construct with a mutated miR-139-5p binding site. CCNB1 overexpression fostered progression of LUAD cells. Mechanistically, miR-139-5p might negatively regulate CCNB1 in LUAD, thereby suppressing cell proliferation, migration, invasion and cell cycle.





Similar content being viewed by others
Data Availability
The data used to support the findings of this study are included within the article. The data and materials in the current study are available from the corresponding author on reasonable request.
References
Siegel, R. L., Miller, K. D., & Jemal, A. (2019). Cancer statistics, 2019. CA: A Cancer Journal for Clinicians, 69, 7–34. https://doi.org/10.3322/caac.21551
Thomson, C. S., & Forman, D. (2009). Cancer survival in England and the influence of early diagnosis: What can we learn from recent EUROCARE results? British Journal of Cancer, 101(Suppl 2), S102-109. https://doi.org/10.1038/sj.bjc.6605399
Jemal, A., Siegel, R., Xu, J., & Ward, E. (2010). Cancer statistics, 2010. CA: A Cancer Journal for Clinicians, 60, 277–300. https://doi.org/10.3322/caac.20073
Zhou, C., & Yao, L. D. (2016). Strategies to improve outcomes of patients with EGRF-mutant non-small cell lung cancer: Review of the literature. Journal of Thoracic Oncology, 11, 174–186. https://doi.org/10.1016/j.jtho.2015.10.002
Villanueva, N., & Bazhenova, L. (2018). New strategies in immunotherapy for lung cancer: Beyond PD-1/PD-L1. Therapeutic Advances in Respiratory Disease, 12, 1753466618794133. https://doi.org/10.1177/1753466618794133
Riaz, S. P., et al. (2012). Trends in incidence of small cell lung cancer and all lung cancer. Lung Cancer, 75, 280–284. https://doi.org/10.1016/j.lungcan.2011.08.004
Shin, J. U., et al. (2012). Prognostic significance of ATM and cyclin B1 in pancreatic neuroendocrine tumor. Tumour Biology, 33, 1645–1651. https://doi.org/10.1007/s13277-012-0420-5
Krek, W., & Nigg, E. A. (1991). Differential phosphorylation of vertebrate p34cdc2 kinase at the G1/S and G2/M transitions of the cell cycle: Identification of major phosphorylation sites. EMBO Journal, 10, 305–316.
Morgan, D. O. (1995). Principles of CDK regulation. Nature, 374, 131–134. https://doi.org/10.1038/374131a0
Chai, N., et al. (2018). FOXM1 promotes proliferation in human hepatocellular carcinoma cells by transcriptional activation of CCNB1. Biochemical and Biophysical Research Communications, 500, 924–929. https://doi.org/10.1016/j.bbrc.2018.04.201
Fang, Y., Yu, H., Liang, X., Xu, J., & Cai, X. (2014). Chk1-induced CCNB1 overexpression promotes cell proliferation and tumor growth in human colorectal cancer. Cancer Biology & Therapy, 15, 1268–1279. https://doi.org/10.4161/cbt.29691
Li, S., et al. (2018). Identification of an eight-gene prognostic signature for lung adenocarcinoma. Cancer Manag Res, 10, 3383–3392. https://doi.org/10.2147/CMAR.S173941
Bartel, D. P. (2004). MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell, 116, 281–297. https://doi.org/10.1016/s0092-8674(04)00045-5
Bartel, D. P. (2009). MicroRNAs: Target recognition and regulatory functions. Cell, 136, 215–233. https://doi.org/10.1016/j.cell.2009.01.002
Huang, L. L., et al. (2017). Potential role of miR-139-5p in cancer diagnosis, prognosis and therapy. Oncology Letters, 14, 1215–1222. https://doi.org/10.3892/ol.2017.6351
Krishnan, K., et al. (2013). miR-139-5p is a regulator of metastatic pathways in breast cancer. RNA, 19, 1767–1780. https://doi.org/10.1261/rna.042143.113
Bushati, N., & Cohen, S. M. (2007). microRNA functions. Annual Review of Cell and Developmental Biology, 23, 175–205. https://doi.org/10.1146/annurev.cellbio.23.090506.123406
Adam, L., et al. (2009). miR-200 expression regulates epithelial-to-mesenchymal transition in bladder cancer cells and reverses resistance to epidermal growth factor receptor therapy. Clinical Cancer Research, 15, 5060–5072. https://doi.org/10.1158/1078-0432.CCR-08-2245
Qiu, G., Lin, Y., Zhang, H., & Wu, D. (2015). miR-139-5p inhibits epithelial-mesenchymal transition, migration and invasion of hepatocellular carcinoma cells by targeting ZEB1 and ZEB2. Biochemical and Biophysical Research Communications, 463, 315–321. https://doi.org/10.1016/j.bbrc.2015.05.062
Asangani, I. A., et al. (2008). MicroRNA-21 (miR-21) post-transcriptionally downregulates tumor suppressor Pdcd4 and stimulates invasion, intravasation and metastasis in colorectal cancer. Oncogene, 27, 2128–2136. https://doi.org/10.1038/sj.onc.1210856
Shao, Y., Liang, B., Long, F., & Jiang, S. J. (2017). Diagnostic MicroRNA biomarker discovery for non-small-cell lung cancer adenocarcinoma by integrative bioinformatics analysis. BioMed Research International, 2017, 2563085. https://doi.org/10.1155/2017/2563085
Survival Analysis v. 2.37–7 (2014).
Robinson, M. D., McCarthy, D. J., & Smyth, G. K. (2010). edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics (Oxford, England), 26, 139–140. https://doi.org/10.1093/bioinformatics/btp616
Zhuang, L., Yang, Z., & Meng, Z. (2018). Upregulation of BUB1B, CCNB1, CDC7, CDC20, and MCM3 in tumor tissues predicted worse overall survival and disease-free survival in hepatocellular carcinoma patients. BioMed research international, 2018, 7897346. https://doi.org/10.1155/2018/7897346
Wang, S., Sun, H., Zhan, X., & Wang, Q. (2020). MicroRNA718 serves a tumorsuppressive role in nonsmall cell lung cancer by directly targeting CCNB1. International Journal of Molecular Medicine, 45, 33–44. https://doi.org/10.3892/ijmm.2019.4396
Choudhury, T. R. Is Hepatoglobin therapy justified? J Assoc Physicians India 39, 724; author reply 724–725 (1991).
Greenawalt, E. J., et al. (2019). Targeting of SGK1 by miR-576-3p inhibits lung adenocarcinoma migration and invasion. Molecular Cancer Research, 17, 289–298. https://doi.org/10.1158/1541-7786.MCR-18-0364
Pan, Z. H., Guo, X. Q., Shan, J., & Luo, S. X. (2018). LINC00324 exerts tumor-promoting functions in lung adenocarcinoma via targeting miR-615-5p/AKT1 axis. European Review for Medical and Pharmacological Sciences, 22, 8333–8342. https://doi.org/10.26355/eurrev_201812_16531
Zhu, D., et al. (2019). MiR-138-5p suppresses lung adenocarcinoma cell epithelial-mesenchymal transition, proliferation and metastasis by targeting ZEB2. Pathology, Research and Practice, 215, 861–872. https://doi.org/10.1016/j.prp.2019.01.029
Zhou, L. Y., Zhang, F. W., Tong, J., & Liu, F. (2020). MiR-191-5p inhibits lung adenocarcinoma by repressing SATB1 to inhibit Wnt pathway. Molecular Genetics & Genomic Medicine, 8, e1043. https://doi.org/10.1002/mgg3.1043
Wang, F., Chen, X., Yu, X., & Lin, Q. (2019). Degradation of CCNB1 mediated by APC11 through UBA52 ubiquitination promotes cell cycle progression and proliferation of non-small cell lung cancer cells. Am J Transl Res, 11, 7166–7185.
Wang, J., et al. (2020). PKMYT1 is associated with prostate cancer malignancy and may serve as a therapeutic target. Gene, 744, 144608. https://doi.org/10.1016/j.gene.2020.144608
Li, P., Xiao, Z., Luo, J., Zhang, Y., & Lin, L. (2019). MiR-139-5p, miR-940 and miR-193a-5p inhibit the growth of hepatocellular carcinoma by targeting SPOCK1. Journal of Cellular and Molecular Medicine, 23, 2475–2488. https://doi.org/10.1111/jcmm.14121
Liu, J., et al. (2018). Tumor-suppressor role of miR-139-5p in endometrial cancer. Cancer Cell International, 18, 51. https://doi.org/10.1186/s12935-018-0545-8
Yong-Hao, Y., Xian-Guo, W., Ming, X., & Jin-Ping, Z. (2019). Expression and clinical significance of miR-139-5p in non-small cell lung cancer. Journal of International Medical Research, 47, 867–874. https://doi.org/10.1177/0300060518815379
Wang, K., Jin, J., Ma, T., & Zhai, H. (2017). MiR-139-5p inhibits the tumorigenesis and progression of oral squamous carcinoma cells by targeting HOXA9. Journal of Cellular and Molecular Medicine, 21, 3730–3740. https://doi.org/10.1111/jcmm.13282
Pajic, M., et al. (2018). miR-139-5p modulates radiotherapy resistance in breast cancer by repressing multiple gene networks of DNA repair and ROS defense. Cancer Research, 78, 501–515. https://doi.org/10.1158/0008-5472.Can-16-3105
Pang, C., et al. (2016). MiR-139-5p is increased in the peripheral blood of patients with prostate cancer. Cellular physiology and biochemistry: International journal of experimental cellular physiology, biochemistry, and pharmacology, 39, 1111–1117. https://doi.org/10.1159/000447819
Yang, B., et al. (2019). Downregulation of miR-139-5p promotes prostate cancer progression through regulation of SOX5. Biomedicine & Pharmacotherapy, 109, 2128–2135. https://doi.org/10.1016/j.biopha.2018.09.029
Wu, J., Zhang, T., Chen, Y., & Ha, S. (2020). MiR-139-5p influences hepatocellular carcinoma cell invasion and proliferation capacities via decreasing SLITRK4 expression. Bioscience Reports. https://doi.org/10.1042/BSR20193295
Wu, J., et al. (2019). LncSNHG3/miR-139-5p/BMI1 axis regulates proliferation, migration, and invasion in hepatocellular carcinoma. OncoTargets and Therapy, 12, 6623–6638. https://doi.org/10.2147/ott.S196630
Acknowledgements
Not applicable.
Funding
Not applicable.
Author information
Authors and Affiliations
Contributions
All authors contributed to data analysis, drafting and revising the article, gave final approval of the version to be published, and agreed to be accountable for all aspects of the work.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no potential conflicts of interest.
Ethical Approval
Not applicable.
Consent for Publication
All authors consent to submit the manuscript for publication.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
12033_2022_465_MOESM1_ESM.tif
Supplementary Fig. 1 LUAD malignant progression was suppressed by silencing CCNB1. A: CCNB1 level in H1299 cells; B: The proliferation ability of H1299 cells; C: The invasion of H1299 cells (100×); D: The migration ability of H1299 cells (40×); E: The cell cycle of H1299. The cell grouping settings were as follows: H1299 cells with si-NC and si-CCNB1. * denotes p<0.05 (TIF 3302 kb)
Rights and permissions
About this article
Cite this article
Bao, B., Yu, X. & Zheng, W. MiR-139-5p Targeting CCNB1 Modulates Proliferation, Migration, Invasion and Cell Cycle in Lung Adenocarcinoma. Mol Biotechnol 64, 852–860 (2022). https://doi.org/10.1007/s12033-022-00465-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-022-00465-5