Abstract
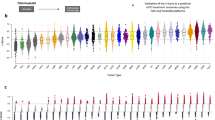
Advances in the understanding of checkpoint blockade immunotherapy have suggested that boosting the cancer-immunity cycle (CIC) can help induce regression of tumors. However, good efficacy only occurs in a subset of patients. Predictive biomarkers that can reflect the tumor microenvironment (TME) and CIC may have great potential. More recently, the presence of intratumoral tertiary lymphoid structures (TLSs) has also been correlated with clinical benefit in patients. In this study, we comprehensively measured the immunogram scores (IGSs) for the CIC and explored the associations between immunological and mutational features and a 12-chemokine metagene TLS signature in data from The Cancer Genome Atlas (TCGA). Three immunotherapy datasets were further applied for validation. In the TCGA dataset, we observed that the 12-chemokine TLS signature score was positively associated with the enhanced IGSs as represented by increased tumor mutational burden (TMB) and neoantigen burden (TNB), enriched immune cell (IC) infiltration, and elevated cytolytic activity and checkpoint expression. Specifically, in bladder cancer and melanoma, the high 12-chemokine TLS signature score was found to potentially reflect an expanded cancer-immunity status characterized by high TNB and an immune-inflamed feature. The predictive and prognostic value of the 12-chemokine TLS signature was further validated in several immunotherapy datasets. The score of the 12-chemokine TLS signature may serve as a pancancer marker of the immune-active phenotype. The 12-chemokine TLS signature showed promise as a predictive and prognostic biomarker for ICB efficacy, especially in melanoma and bladder cancer.





Similar content being viewed by others
Data availability
All data analyzed in our study were obtained from publicly available, online datasets. The raw data of this study are derived from the TCGA database and GEO data portal, as mentioned in the Materials and Methods.
Consent for publication
Not applicable.
References
Sharma P, Hu-Lieskovan S, Wargo JA, Ribas A. Primary, adaptive, and acquired resistance to cancer immunotherapy. Cell. 2017;168(4):707–23.
Havel JJ, Chowell D, Chan TA. The evolving landscape of biomarkers for checkpoint inhibitor immunotherapy. Nat Rev Cancer. 2019;19(3):133–50.
Carbone DP, Reck M, Paz-Ares L, Creelan B, Horn L, Steins M, Felip E, van den Heuvel MM, Ciuleanu TE, Badin F, et al. First-line nivolumab in Stage IV or recurrent non-small-cell lung cancer. N Engl J Med. 2017;376(25):2415–26.
Brahmer J, Reckamp KL, Baas P, Crino L, Eberhardt WE, Poddubskaya E, Antonia S, Pluzanski A, Vokes EE, Holgado E, et al. Nivolumab versus docetaxel in advanced squamous-cell non-small-cell lung cancer. N Engl J Med. 2015;373(2):123–35.
Kim SY, Halmos B. Choosing the best first-line therapy: NSCLC with no actionable oncogenic driver. Lung Cancer Manag. 2020;9(3):LMT36.
Gettinger SN, Choi J, Mani N, Sanmamed MF, Datar I, Sowell R, Du VY, Kaftan E, Goldberg S, Dong W, et al. A dormant TIL phenotype defines non-small cell lung carcinomas sensitive to immune checkpoint blockers. Nat Commun. 2018;9(1):3196.
Ott PA, Bang YJ, Piha-Paul SA, Razak ARA, Bennouna J, Soria JC, Rugo HS, Cohen RB, O’Neil BH, Mehnert JM, et al. T-cell-inflamed gene-expression profile, programmed death ligand 1 expression, and tumor mutational burden predict efficacy in patients treated with pembrolizumab across 20 cancers: KEYNOTE-028. J Clin Oncol. 2019;37(4):318–27.
Riaz N, Havel JJ, Makarov V, Desrichard A, Urba WJ, Sims JS, Hodi FS, Martin-Algarra S, Mandal R, Sharfman WH, et al. Tumor and microenvironment evolution during immunotherapy with nivolumab. Cell. 2017;171(4):934-949.e916.
Sautes-Fridman C, Petitprez F, Calderaro J, Fridman WH. Tertiary lymphoid structures in the era of cancer immunotherapy. Nat Rev Cancer. 2019;19(6):307–25.
Cabrita R, Lauss M, Sanna A, Donia M, Skaarup Larsen M, Mitra S, Johansson I, Phung B, Harbst K, Vallon-Christersson J, et al. Tertiary lymphoid structures improve immunotherapy and survival in melanoma. Nature. 2020;577(7791):561–5.
Coppola D, Nebozhyn M, Khalil F, Dai H, Yeatman T, Loboda A, Mule JJ. Unique ectopic lymph node-like structures present in human primary colorectal carcinoma are identified by immune gene array profiling. Am J Pathol. 2011;179(1):37–45.
Prabhakaran S, Rizk VT, Ma Z, Cheng CH, Berglund AE, Coppola D, Khalil F, Mule JJ, Soliman HH. Evaluation of invasive breast cancer samples using a 12-chemokine gene expression score: correlation with clinical outcomes. Breast Cancer Res. 2017;19(1):71.
Messina JL, Fenstermacher DA, Eschrich S, Qu X, Berglund AE, Lloyd MC, Schell MJ, Sondak VK, Weber JS, Mule JJ. 12-Chemokine gene signature identifies lymph node-like structures in melanoma: potential for patient selection for immunotherapy? Sci Rep. 2012;2:765.
Danaher P, Warren S, Lu R, Samayoa J, Sullivan A, Pekker I, Wallden B, Marincola FM, Cesano A. Pan-cancer adaptive immune resistance as defined by the Tumor Inflammation Signature (TIS): results from The Cancer Genome Atlas (TCGA). J Immunother Cancer. 2018;6(1):63.
Chen DS, Mellman I. Oncology meets immunology: the cancer-immunity cycle. Immunity. 2013;39(1):1–10.
Smyth MJ, Ngiow SF, Ribas A, Teng MW. Combination cancer immunotherapies tailored to the tumour microenvironment. Nat Rev Clin Oncol. 2016;13(3):143–58.
Uno T, Takeda K, Kojima Y, Yoshizawa H, Akiba H, Mittler RS, Gejyo F, Okumura K, Yagita H, Smyth MJ. Eradication of established tumors in mice by a combination antibody-based therapy. Nat Med. 2006;12(6):693–8.
Karasaki T, Nagayama K, Kuwano H, Nitadori JI, Sato M, Anraku M, Hosoi A, Matsushita H, Morishita Y, Kashiwabara K, et al. An immunogram for the cancer-immunity cycle: towards personalized immunotherapy of lung cancer. J Thorac Oncol. 2017;12(5):791–803.
Thorsson V, Gibbs DL, Brown SD, Wolf D, Bortone DS, Ou Yang TH, Porta-Pardo E, Gao GF, Plaisier CL, Eddy JA, et al. The immune landscape of cancer. Immunity. 2018;48(4):812-830.e814.
Mariathasan S, Turley SJ, Nickles D, Castiglioni A, Yuen K, Wang Y, Kadel EE III, Koeppen H, Astarita JL, Cubas R, et al. TGFbeta attenuates tumour response to PD-L1 blockade by contributing to exclusion of T cells. Nature. 2018;554(7693):544–8.
Ulloa-Montoya F, Louahed J, Dizier B, Gruselle O, Spiessens B, Lehmann FF, Suciu S, Kruit WH, Eggermont AM, Vansteenkiste J, Brichard VG. Predictive gene signature in MAGE-A3 antigen-specific cancer immunotherapy. J Clin Oncol. 2013;31(19):2388–95.
Charoentong P, Finotello F, Angelova M, Mayer C, Efremova M, Rieder D, Hackl H, Trajanoski Z. Pan-cancer immunogenomic analyses reveal genotype–immunophenotype relationships and predictors of response to checkpoint blockade. Cell Rep. 2017;18(1):248–62.
Barbie DA, Tamayo P, Boehm JS, Kim SY, Moody SE, Dunn IF, Schinzel AC, Sandy P, Meylan E, Scholl C, et al. Systematic RNA interference reveals that oncogenic KRAS-driven cancers require TBK1. Nature. 2009;462(7269):108–12.
Xu L, Deng C, Pang B, Zhang X, Liu W, Liao G, Yuan H, Cheng P, Li F, Long Z, et al. TIP: a web server for resolving tumor immunophenotype profiling. Cancer Res. 2018;78(23):6575–80.
Zhu J, Armstrong AJ, Friedlander TW, Kim W, Pal SK, George DJ, Zhang T. Biomarkers of immunotherapy in urothelial and renal cell carcinoma: PD-L1, tumor mutational burden, and beyond. J Immunother Cancer. 2018;6(1):4.
Ye Y, Jing Y, Li L, Mills GB, Diao L, Liu H, Han L. Sex-associated molecular differences for cancer immunotherapy. Nat Commun. 2020;11(1):1779.
Huang Z, Chen B, Liu X, Li H, Xie L, Gao Y, Duan R, Li Z, Zhang J, Zheng Y, Su W. Effects of sex and aging on the immune cell landscape as assessed by single-cell transcriptomic analysis. Proc Natl Acad Sci USA. 2021;118(33):e2023216118.
Conforti F, Pala L, Bagnardi V, De Pas T, Martinetti M, Viale G, Gelber RD, Goldhirsch A. Cancer immunotherapy efficacy and patients’ sex: a systematic review and meta-analysis. Lancet Oncol. 2018;19(6):737–46.
Acknowledgements
We thank the professional English-speaking editors at American Journal Experts for assistance with improving the quality of the language.
Funding
Not applicable.
Author information
Authors and Affiliations
Contributions
XL, LY: Conceptualization. ZW: Acquisition of the data. XL, XL: Formal analysis and investigation. XL: Drafting of the manuscript. KO, ZW: Statistical analysis. LY: Supervision. All authors: Critical revision of the manuscript for important intellectual content.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no competing interests to declare.
Ethical approval
Not necessary.
Informed consent
Not necessary.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Li, X., Wan, Z., Liu, X. et al. A 12-chemokine gene signature is associated with the enhanced immunogram scores and is relevant for precision immunotherapy. Med Oncol 39, 43 (2022). https://doi.org/10.1007/s12032-021-01635-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12032-021-01635-2