Abstract
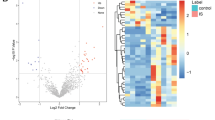
New data are accumulating on the involvement of interaction among circular RNAs (circRNAs), microRNAs (miRNAs/miRs), and messenger RNAs (mRNAs) in cerebral infarction (CI). This study aims to illustrate the GEO database-based identification of a circRNA-miRNA-mRNA crosstalk network underlying immune cell infiltration in CI. The differential analysis suggested that 1696 circRNAs, 1989 miRNAs, and 5550 mRNAs that were differentially expressed in CI samples were retrieved from GEO database. GO and KEGG functional enrichment analyses showed that the differentially expressed mRNAs were mainly associated with common risk factors of CI, such as immune and inflammatory response. Next, the circRNA-miRNA pairs and miRNA-mRNA pairs were predicted, and the circRNA-miRNA-mRNA network was constructed by Cytoscape software. Totally, 436 circRNA-miRNA pairs were obtained through the online database, and 2033 miRNA-mRNA pairs were used to construct the circRNA-miRNA-mRNA crosstalk network. A protein–protein interaction (PPI) network was constructed on the basis of the ceRNA network, followed by key gene identification in the GSE9877 dataset. FAS was identified as the key gene in CI. The constructed FAS-mediated circRNA-miRNA-mRNA crosstalk network included five upregulated circRNAs (hsa_circ_0075341, hsa_circ_0049637, hsa_circ_0001085, hsa_circ_0004808 and hsa_circ_0092337) and five downregulated miRNAs (hsa-miR-92a-2-5p, hsa-miR-1245b-3p, hsa-miR-592, hsa-miR-224-5p, and hsa-miR-30e-3p). Furthermore, the CIBERSORT algorithm indicated that FAS was associated with immune cell infiltration in CI. In conclusion, this study revealed a role for FAS-centered circRNA-miRNA-mRNA crosstalk network in regulating immune cell infiltration of CI, which may be a viable target for CI prevention.








Similar content being viewed by others
Data Availability
The data underlying this article will be shared on reasonable request to the corresponding author.
References
Agarwal V, Bell GW, Nam JW, Bartel DP (2015) Predicting effective microRNA target sites in mammalian mRNAs. Elife 4:e05005. https://doi.org/10.7554/eLife.05005
Brochard V, Combadiere B, Prigent A, Laouar Y, Perrin A, Beray-Berthat V, Bonduelle O, Alvarez-Fischer D, Callebert J, Launay JM, Duyckaerts C, Flavell RA, Hirsch EC, Hunot S (2009) Infiltration of CD4+ lymphocytes into the brain contributes to neurodegeneration in a mouse model of Parkinson disease. J Clin Invest 119(1):182–192. https://doi.org/10.1172/JCI36470
Chalbatani GM, Momeni SA, MohammadiHadloo MH, Karimi Z, Hadizadeh M, Jalali SA, Miri SR, Memari F, Hamblin MR (2022) Comprehensive analysis of ceRNA networks to determine genes related to prognosis, overall survival, and immune infiltration in clear cell renal carcinoma. Comput Biol Med 141:105043. https://doi.org/10.1016/j.compbiomed.2021.105043
Chen B, Yi J, Xu Y, Zheng P, Tang R, Liu B (2022) Construction of a circRNA-miRNA-mRNA network revealed the potential mechanism of Buyang Huanwu Decoction in the treatment of cerebral ischemia. Biomed Pharmacother 145:112445. https://doi.org/10.1016/j.biopha.2021.112445
Cheon SY, Kim EJ, Kim JM, Kam EH, Ko BW, Koo BN (2017) Regulation of microglia and macrophage polarization via apoptosis signal-regulating kinase 1 silencing after ischemic/hypoxic injury. Front Mol Neurosci 10:261. https://doi.org/10.3389/fnmol.2017.00261
Chu HX, Kim HA, Lee S, Moore JP, Chan CT, Vinh A, Gelderblom M, Arumugam TV, Broughton BR, Drummond GR, Sobey CG (2014) Immune cell infiltration in malignant middle cerebral artery infarction: comparison with transient cerebral ischemia. J Cereb Blood Flow Metab 34(3):450–459. https://doi.org/10.1038/jcbfm.2013.217
Clough E, Barrett T (2016) The gene expression omnibus database. Methods Mol Biol 1418:93–110. https://doi.org/10.1007/978-1-4939-3578-9_5
Glazar P, Papavasileiou P, Rajewsky N (2014) circBase: a database for circular RNAs. RNA 20(11):1666–1670. https://doi.org/10.1261/rna.043687.113
Han B, Zhang Y, Zhang Y, Bai Y, Chen X, Huang R, Wu F, Leng S, Chao J, Zhang JH, Hu G, Yao H (2018) Novel insight into circular RNA HECTD1 in astrocyte activation via autophagy by targeting MIR142-TIPARP: implications for cerebral ischemic stroke. Autophagy 14(7):1164–1184. https://doi.org/10.1080/15548627.2018.1458173
Hu XL, Su Q, Meng DL, Ren YS, Su ZQ (2021) Circular RNA expression alteration and bioinformatics analysis in patients with acute cerebral infarction injury. Bioengineered 12(2):11490–11505. https://doi.org/10.1080/21655979.2021.2009960
Irmady K, Jackman KA, Padow VA, Shahani N, Martin LA, Cerchietti L, Unsicker K, Iadecola C, Hempstead BL (2014) Mir-592 regulates the induction and cell death-promoting activity of p75NTR in neuronal ischemic injury. J Neurosci 34(9):3419–3428. https://doi.org/10.1523/JNEUROSCI.1982-13.2014
Krzyzowska M, Kowalczyk A, Skulska K, Thorn K, Eriksson K (2021) Fas/FasL contributes to HSV-1 brain infection and neuroinflammation. Front Immunol 12:714821. https://doi.org/10.3389/fimmu.2021.714821
Li T, Shao Y, Fu L, Xie Y, Zhu L, Sun W, Yu R, Xiao B, Guo J (2018) Plasma circular RNA profiling of patients with gastric cancer and their droplet digital RT-PCR detection. J Mol Med (Berl) 96(1):85–96. https://doi.org/10.1007/s00109-017-1600-y
Liang Z, Chi YJ, Lin GQ, Luo SH, Jiang QY, Chen YK (2018) MiRNA-26a promotes angiogenesis in a rat model of cerebral infarction via PI3K/AKT and MAPK/ERK pathway. Eur Rev Med Pharmacol Sci 22(11):3485–3492. https://doi.org/10.26355/eurrev_201806_15175
Liu M, Wang Q, Shen J, Yang BB, Ding X (2019) Circbank: a comprehensive database for circRNA with standard nomenclature. RNA Biol 16(7):899–905. https://doi.org/10.1080/15476286.2019.1600395
Liu X, Zhong G, Li W, Zeng Y, Wu M (2021a) The construction and comprehensive analysis of a ceRNA immunoregulatory network and tissue-infiltrating immune cells in atrial fibrillation. Int J Gen Med 14:9051–9066. https://doi.org/10.2147/IJGM.S338797
Liu Z, Wu X, Yu Z, Tang X (2021b) Reconstruction of circRNA-miRNA-mRNA associated ceRNA networks reveal functional circRNAs in intracerebral hemorrhage. Sci Rep 11(1):11584. https://doi.org/10.1038/s41598-021-91059-9
Lu G, Wong MS, Xiong MZQ, Leung CK, Su XW, Zhou JY, Poon WS, Zheng VZY, Chan WY, Wong GKC (2017) Circulating microRNAs in delayed cerebral infarction after aneurysmal subarachnoid hemorrhage. J Am Heart Assoc 6(4):e005363. https://doi.org/10.1161/JAHA.116.005363
Mahovic D, Zurak N, Lakusic N, Sporis D, Zarkovic N, Stancin N, Bosnar-Puretic M (2013) The dynamics of soluble Fas/APO 1 apoptotic biochemical marker in acute ischemic stroke patients. Adv Med Sci 58(2):298–303. https://doi.org/10.2478/ams-2013-0014
Meng HL, Li XX, Chen YT, Yu LJ, Zhang H, Lao JM, Zhang X, Xu Y (2016) Neuronal soluble Fas ligand drives M1-microglia polarization after cerebral ischemia. CNS Neurosci Ther 22(9):771–781. https://doi.org/10.1111/cns.12575
Newman AM, Liu CL, Green MR, Gentles AJ, Feng W, Xu Y, Hoang CD, Diehn M, Alizadeh AA (2015) Robust enumeration of cell subsets from tissue expression profiles. Nat Methods 12(5):453–457. https://doi.org/10.1038/nmeth.3337
Paules C, Youssef L, Miranda J, Crovetto F, Estanyol JM, Fernandez G, Crispi F, Gratacos E (2020) Maternal proteomic profiling reveals alterations in lipid metabolism in late-onset fetal growth restriction. Sci Rep 10(1):21033. https://doi.org/10.1038/s41598-020-78207-3
Qi X, Zhang DH, Wu N, Xiao JH, Wang X, Ma W (2015) ceRNA in cancer: possible functions and clinical implications. J Med Genet 52(10):710–718. https://doi.org/10.1136/jmedgenet-2015-103334
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13(11):2498–2504. https://doi.org/10.1101/gr.1239303
Shao S, Wang C, Wang S, Zhang H, Zhang Y (2020) Hsa_circ_0075341 is up-regulated and exerts oncogenic properties by sponging miR-149–5p in cervical cancer. Biomed Pharmacother 121:109582. https://doi.org/10.1016/j.biopha.2019.109582
Song A, Yang Y, He H, Sun J, Chang Q, Xue Q (2021) Inhibition of long non-coding RNA KCNQ1OT1 attenuates neuroinflammation and neuronal apoptosis through regulating NLRP3 expression via sponging miR-30e-3p. J Inflamm Res 14:1731–1742. https://doi.org/10.2147/JIR.S291274
Su B, Wang X, Sun Y, Long M, Zheng J, Wu W, Li L (2020) miR-30e-3p promotes cardiomyocyte autophagy and inhibits apoptosis via regulating Egr-1 during ischemia/hypoxia. Biomed Res Int 2020:7231243. https://doi.org/10.1155/2020/7231243
Szklarczyk D, Gable AL, Lyon D, Junge A, Wyder S, Huerta-Cepas J, Simonovic M, Doncheva NT, Morris JH, Bork P, Jensen LJ, Mering CV (2019) STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res 47(D1):D607–D613. https://doi.org/10.1093/nar/gky1131
Takeda H, Yamaguchi T, Yano H, Tanaka J (2021) Microglial metabolic disturbances and neuroinflammation in cerebral infarction. J Pharmacol Sci 145(1):130–139. https://doi.org/10.1016/j.jphs.2020.11.007
Tan ZR, Zhang C, Tian FF (2019) Spectrum of clinical features and neuroimaging findings in acute cerebral infarction patients with unusual ipsilateral motor impairment- a series of 22 cases. BMC Neurol 19(1):279. https://doi.org/10.1186/s12883-019-1516-y
Tao Z, Rao G, Wu S, Lin Y, Wang J, Chen Z (2021) Rehabilitation evaluation of hemiplegic patients with anterior circulation cerebral infarction based on cranial magnetic stimulation. J Healthc Eng 2021:7868419. https://doi.org/10.1155/2021/7868419
Tay Y, Rinn J, Pandolfi PP (2014) The multilayered complexity of ceRNA crosstalk and competition. Nature 505(7483):344–352. https://doi.org/10.1038/nature12986
Tokar T, Pastrello C, Rossos AEM, Abovsky M, Hauschild AC, Tsay M, Lu R, Jurisica I (2018) mirDIP 4.1-integrative database of human microRNA target predictions. Nucleic Acids Res 46(D1):D360–D370. https://doi.org/10.1093/nar/gkx1144
Yamada A, Arakaki R, Saito M, Kudo Y, Ishimaru N (2017) Dual role of Fas/FasL-mediated signal in peripheral immune tolerance. Front Immunol 8:403. https://doi.org/10.3389/fimmu.2017.00403
Yamamura S, Imai-Sumida M, Tanaka Y, Dahiya R (2018) Interaction and cross-talk between non-coding RNAs. Cell Mol Life Sci 75(3):467–484. https://doi.org/10.1007/s00018-017-2626-6
Yan B, Ren F, Shang W, Gong X (2022) Transcriptomic analysis reveals genetic cross-talk between periodontitis and hypothyroidism. Dis Markers 2022:5736394. https://doi.org/10.1155/2022/5736394
Yan Z, Xiao Y, Chen Y, Luo G (2020) Screening and identification of epithelial-to-mesenchymal transition-related circRNA and miRNA in prostate cancer. Pathol Res Pract 216(2):152784. https://doi.org/10.1016/j.prp.2019.152784
Zhang W, Liu Y, Min Z, Liang G, Mo J, Ju Z, Zeng B, Guan W, Zhang Y, Chen J, Zhang Q, Li H, Zeng C, Wei Y, Chan GC (2022) circMine: a comprehensive database to integrate, analyze and visualize human disease-related circRNA transcriptome. Nucleic Acids Res 50(D1):D83–D92. https://doi.org/10.1093/nar/gkab809
Zhang X, Ha T, Lu C, Lam F, Liu L, Schweitzer J, Kalbfleisch J, Kao RL, Williams DL, Li C (2015) Poly (I:C) therapy decreases cerebral ischaemia/reperfusion injury via TLR3-mediated prevention of Fas/FADD interaction. J Cell Mol Med 19(3):555–565. https://doi.org/10.1111/jcmm.12456
Zhang Z, He J, Wang B (2021) Circular RNA circ_HECTD1 regulates cell injury after cerebral infarction by miR-27a-3p/FSTL1 axis. Cell Cycle 20(9):914–926. https://doi.org/10.1080/15384101.2021.1909885
Author information
Authors and Affiliations
Contributions
KY and JC designed the study. ZHF and BR collated the data, carried out data analyses, and produced the initial draft of the manuscript. KY and JC contributed to drafting the manuscript. All authors have read and approved the final submitted manuscript.
Corresponding author
Ethics declarations
Ethics Approval and Consent to Participate
This article does not contain any studies with human participants or animals performed by any of the authors.
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ying, K., Chen, J., Fu, Z. et al. FAS-mediated circRNA-miRNA-mRNA Crosstalk Network Regulates Immune Cell Infiltration in Cerebral Infarction. J Mol Neurosci 73, 117–128 (2023). https://doi.org/10.1007/s12031-023-02100-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12031-023-02100-7