Abstract
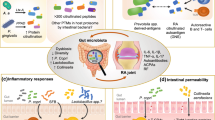
A growing amount of evidence suggests that gut microbiota plays an important role in human health, including a possible role in the pathogenesis of rheumatic and musculoskeletal diseases (RMD). We analysed the current evidence about the role of microbiota in rheumatoid arthritis (RA), spondyloarthritis (SpA), systemic lupus erythematosus (SLE) and systemic sclerosis (SSc). In RA, we found a general consensus regarding a reduction of diversity and a specific bacterial signature, with consistent changes according to the different ethnic and geographical areas. The major pathogenetic role in RA is recognised for P. copri, L. salivarius and Collinsella, even if findings become more heterogeneous when considering established disease. In SpA, we found a relative gut abundance of Akkermansia, Coprococcus, Ruminoccocus and a relative reduction in Bacterioides and Firmicutes spp. Human and preclinical data suggest loss of mucosal barrier, increased permeability and Th1- and Th17-mediated inflammation. Additionally, HLA-B27 seems to play a role in shaping the intestinal microbiota and the consequent inflammation. In SLE, the typical gut microbiota signature was characterised by a reduction in the Firmicutes/Bacteroidetes ratio and by enrichment of Rhodococcus, Eggerthella, Klebsiella, Prevotella, Eubacterium and Flavonifractor, even if their real pathogenic impact remains unclear. In SSc, gastrointestinal dysbiosis is well documented with an increase of pro-inflammatory species (Fusobacterium, Prevotella, Ruminococcus, Akkermansia, γ-Proteobacteria, Erwinia, Trabsulsiella, Bifidobacterium, Lactobacillus, Firmicutes and Actinobacteria) and a reduction of species as Faecalibacterium, Clostridium, Bacteroidetes and Rikenella. In conclusion, seems possible to recognise a distinct gut microbiota profile for each RMD, even if significant differences in bacterial species do exist between different studies and there is a high risk of bias due to the cross-sectional nature of such studies. Therefore longitudinal studies are needed, especially on patients with preclinical and early disease, to investigate the real role of gut microbiota in the pathogenesis of RMD.
Similar content being viewed by others
References
Thursby E, Juge N (2017) Introduction to the human gut microbiota. Biochem J 474:1823–1836. https://doi.org/10.1042/BCJ20160510
Rooks MG, Garrett WS (2016) Gut microbiota, metabolites and host immunity. Nat Rev Immunol 16:341–352. https://doi.org/10.1038/nri.2016.42
Dong Y, Yao J, Deng Q et al (2023) Relationship between gut microbiota and rheumatoid arthritis: a bibliometric analysis. Front Immunol 14:1131933
Levy M, Kolodziejczyk AA, Thaiss CA, Elinav E (2017) Dysbiosis and the immune system. Nat Rev Immunol 17:219–232. https://doi.org/10.1038/nri.2017.7
Bashir H, Singh S, Singh RP et al (2023) Age-mediated gut microbiota dysbiosis promotes the loss of dendritic cells tolerance. Aging Cell 22:e13838. https://doi.org/10.1111/acel.13838
Ursell LK, Metcalf JL, Parfrey LW, Knight R (2012) Defining the human microbiome. Nutr Rev 70:S38–S44. https://doi.org/10.1111/j.1753-4887.2012.00493.x
Smolen JS, Aletaha D, Barton A et al (2018) Rheumatoid arthritis. Nat Rev Dis Primer 4:18001. https://doi.org/10.1038/nrdp.2018.1
Alpizar-Rodriguez D, Lesker TR, Gronow A et al (2019) Prevotella copri in individuals at risk for rheumatoid arthritis. Ann Rheum Dis 78:590–593. https://doi.org/10.1136/annrheumdis-2018-214514
Scher JU, Sczesnak A, Longman RS et al (2013) Expansion of intestinal Prevotella copri correlates with enhanced susceptibility to arthritis. Elife 2:e01202. https://doi.org/10.7554/eLife.01202
Vaahtovuo J, Munukka E, Korkeamäki M et al (2008) Fecal microbiota in early rheumatoid arthritis. J Rheumatol 35:1500–1505
Chen J, Wright K, Davis JM et al (2016) An expansion of rare lineage intestinal microbes characterizes rheumatoid arthritis. Genome Med 8:43. https://doi.org/10.1186/s13073-016-0299-7
Chiang H-I, Li J-R, Liu C-C et al (2019) An association of gut microbiota with different phenotypes in Chinese patients with rheumatoid arthritis. J Clin Med 8:1770. https://doi.org/10.3390/jcm8111770
Zhang X, Zhang D, Jia H et al (2015) The oral and gut microbiomes are perturbed in rheumatoid arthritis and partly normalized after treatment. Nat Med 21:895–905. https://doi.org/10.1038/nm.3914
Kishikawa T, Maeda Y, Nii T et al (2020) Metagenome-wide association study of gut microbiome revealed novel aetiology of rheumatoid arthritis in the Japanese population. Ann Rheum Dis 79:103–111. https://doi.org/10.1136/annrheumdis-2019-215743
Berthelot J-M, Lioté F, Sibilia J (2023) Methotrexate also improves rheumatoid arthritis through correction of microbiota dysbiosis. Joint Bone Spine 90:105602. https://doi.org/10.1016/j.jbspin.2023.105602
Vallier M, Segurens B, Larsonneur E et al (2023) Characterisation of gut microbiota composition in patients with axial spondyloarthritis and its modulation by TNF inhibitor treatment. RMD Open 9:e002794. https://doi.org/10.1136/rmdopen-2022-002794
Pianta A, Arvikar S, Strle K et al (2017) Evidence of the immune relevance of Prevotella copri, a gut microbe, in patients with rheumatoid arthritis. Arthritis Rheumatol Hoboken NJ 69:964–975. https://doi.org/10.1002/art.40003
Rashid T, Ebringer A (2012) Autoimmunity in rheumatic diseases is induced by microbial infections via crossreactivity or molecular mimicry. Autoimmune Dis 2012:539282. https://doi.org/10.1155/2012/539282
van Heemst J, Jansen DTSL, Polydorides S et al (2015) Crossreactivity to vinculin and microbes provides a molecular basis for HLA-based protection against rheumatoid arthritis. Nat Commun 6:6681. https://doi.org/10.1038/ncomms7681
Parsaei M, Sarafraz N, Moaddab SY, Ebrahimzadeh Leylabadlo H (2021) The importance of Faecalibacterium prausnitzii in human health and diseases. New Microb New Infect 43:100928. https://doi.org/10.1016/j.nmni.2021.100928
van Delft MAM, van der Woude D, Toes REM, Trouw LA (2019) Secretory form of rheumatoid arthritis-associated autoantibodies in serum are mainly of the IgM isotype, suggesting a continuous reactivation of autoantibody responses at mucosal surfaces. Ann Rheum Dis 78:146–148. https://doi.org/10.1136/annrheumdis-2018-213724
Kokkonen H, Mullazehi M, Berglin E et al (2011) Antibodies of IgG, IgA and IgM isotypes against cyclic citrullinated peptide precede the development of rheumatoid arthritis. Arthritis Res Ther 13:R13. https://doi.org/10.1186/ar3237
Rantapää-Dahlqvist S, de Jong BAW, Berglin E et al (2003) Antibodies against cyclic citrullinated peptide and IgA rheumatoid factor predict the development of rheumatoid arthritis. Arthritis Rheum 48:2741–2749. https://doi.org/10.1002/art.11223
Maeda Y, Takeda K (2019) Host–microbiota interactions in rheumatoid arthritis. Exp Mol Med 51:1–6. https://doi.org/10.1038/s12276-019-0283-6
Sieper J, Poddubnyy D (2017) Axial spondyloarthritis. Lancet Lond Engl 390:73–84. https://doi.org/10.1016/S0140-6736(16)31591-4
FitzGerald O, Ogdie A, Chandran V et al (2021) Psoriatic arthritis. Nat Rev Dis Primer 7:1–17. https://doi.org/10.1038/s41572-021-00293-y
Ciccia F, Rizzo A, Triolo G (2016) Subclinical gut inflammation in ankylosing spondylitis. Curr Opin Rheumatol 28:89–96. https://doi.org/10.1097/BOR.0000000000000239
Costello M-E, Elewaut D, Kenna TJ, Brown MA (2013) Microbes, the gut and ankylosing spondylitis. Arthritis Res Ther 15:214. https://doi.org/10.1186/ar4228
Gilis E, Mortier C, Venken K et al (2018) The role of the microbiome in gut and joint inflammation in psoriatic arthritis and spondyloarthritis. J Rheumatol Suppl 94:36–39. https://doi.org/10.3899/jrheum.180135
Zhang L, Han R, Zhang X et al (2019) Fecal microbiota in patients with ankylosing spondylitis: correlation with dietary factors and disease activity. Clin Chim Acta Int J Clin Chem 497:189–196. https://doi.org/10.1016/j.cca.2019.07.038
Wen C, Zheng Z, Shao T et al (2017) Quantitative metagenomics reveals unique gut microbiome biomarkers in ankylosing spondylitis. Genome Biol 18:142. https://doi.org/10.1186/s13059-017-1271-6
Tito RY, Cypers H, Joossens M et al (2017) Brief report: dialister as a microbial marker of disease activity in spondyloarthritis. Arthritis Rheumatol Hoboken NJ 69:114–121. https://doi.org/10.1002/art.39802
Breban M, Tap J, Leboime A et al (2017) Faecal microbiota study reveals specific dysbiosis in spondyloarthritis. Ann Rheum Dis 76:1614–1622. https://doi.org/10.1136/annrheumdis-2016-211064
Scher JU, Ubeda C, Artacho A et al (2015) Decreased bacterial diversity characterizes the altered gut microbiota in patients with psoriatic arthritis, resembling dysbiosis in inflammatory bowel disease. Arthritis Rheumatol Hoboken NJ 67:128–139. https://doi.org/10.1002/art.38892
Mielants H, De Keyser F, Baeten D, Van den Bosch F (2005) Gut inflammation in the spondyloarthropathies. Curr Rheumatol Rep 7:188–194. https://doi.org/10.1007/s11926-996-0038-y
Gill T, Asquith M, Rosenbaum JT, Colbert RA (2015) The intestinal microbiome in spondyloarthritis. Curr Opin Rheumatol 27:319–325. https://doi.org/10.1097/BOR.0000000000000187
Turpin W, Espin-Garcia O, Xu W et al (2016) Association of host genome with intestinal microbial composition in a large healthy cohort. Nat Genet 48:1413–1417. https://doi.org/10.1038/ng.3693
Asquith M, Sternes PR, Costello M-E et al (2019) HLA alleles associated with risk of ankylosing spondylitis and rheumatoid arthritis influence the gut microbiome. Arthritis Rheumatol Hoboken NJ 71:1642–1650. https://doi.org/10.1002/art.40917
Berland M, Meslier V, Berreira Ibraim S et al (2023) Both disease activity and HLA-B27 status are associated with gut microbiome dysbiosis in spondyloarthritis patients. Arthritis Rheumatol Hoboken NJ 75:41–52. https://doi.org/10.1002/art.42289
Ciccia F, Guggino G, Rizzo A et al (2017) Dysbiosis and zonulin upregulation alter gut epithelial and vascular barriers in patients with ankylosing spondylitis. Ann Rheum Dis 76:1123–1132. https://doi.org/10.1136/annrheumdis-2016-210000
Veys EM, van Leare M (1973) Serum IgG, IgM, and IgA levels in ankylosing spondylitis. Ann Rheum Dis 32:493–496. https://doi.org/10.1136/ard.32.6.493
Laurent MR, Panayi GS (1983) Acute-phase proteins and serum immunoglobulins in ankylosing spondylitis. Ann Rheum Dis 42:524–528. https://doi.org/10.1136/ard.42.5.524
Demetter P, Van Huysse JA, De Keyser F et al (2002) Increase in lymphoid follicles and leukocyte adhesion molecules emphasizes a role for the gut in spondyloarthropathy pathogenesis. J Pathol 198:517–522. https://doi.org/10.1002/path.1235
Romero-Sánchez C, Bautista-Molano W, Parra V et al (2017) Gastrointestinal symptoms and elevated levels of anti-saccharomyces cerevisiae antibodies are associated with higher disease activity in Colombian patients with spondyloarthritis. Int J Rheumatol 2017:4029584. https://doi.org/10.1155/2017/4029584
Wallis D, Asaduzzaman A, Weisman M et al (2013) Elevated serum anti-flagellin antibodies implicate subclinical bowel inflammation in ankylosing spondylitis: an observational study. Arthritis Res Ther 15:R166. https://doi.org/10.1186/ar4350
Henke MT, Kenny DJ, Cassilly CD et al (2019) Ruminococcus gnavus, a member of the human gut microbiome associated with Crohn’s disease, produces an inflammatory polysaccharide. Proc Natl Acad Sci USA 116:12672–12677. https://doi.org/10.1073/pnas.1904099116
Smith PM, Howitt MR, Panikov N et al (2013) The microbial metabolites, short-chain fatty acids, regulate colonic Treg cell homeostasis. Science 341:569–573. https://doi.org/10.1126/science.1241165
Fanouriakis A, Tziolos N, Bertsias G, Boumpas DT (2021) Update οn the diagnosis and management of systemic lupus erythematosus. Ann Rheum Dis 80:14–25. https://doi.org/10.1136/annrheumdis-2020-218272
Lei Y, Liu Q, Li Q et al (2023) Exploring the complex relationship between microbiota and systemic lupus erythematosus. Curr Rheumatol Rep 25:107–116. https://doi.org/10.1007/s11926-023-01102-z
Hevia A, Milani C, López P et al (2014) Intestinal dysbiosis associated with systemic lupus erythematosus. MBio 5:e01548-e11514. https://doi.org/10.1128/mBio.01548-14
He Z, Shao T, Li H et al (2016) Alterations of the gut microbiome in Chinese patients with systemic lupus erythematosus. Gut Pathog 8:64. https://doi.org/10.1186/s13099-016-0146-9
Santiago-Rodriguez TM, Hollister EB (2019) Human virome and disease: high-throughput sequencing for virus discovery, identification of phage-bacteria dysbiosis and development of therapeutic approaches with emphasis on the human gut. Viruses 11:656. https://doi.org/10.3390/v11070656
Li B-Z, Wang H, Li X-B et al (2022) Altered gut fungi in systemic lupus erythematosus—a pilot study. Front Microbiol 13:1031079. https://doi.org/10.3389/fmicb.2022.1031079
Liu F, Ren T, Li X et al (2021) Distinct microbiomes of gut and saliva in patients with systemic lupus erythematous and clinical associations. Front Immunol 12:626217. https://doi.org/10.3389/fimmu.2021.626217
Wang X, Shu Q, Song L et al (2022) Gut microbiota in systemic lupus erythematosus and correlation with diet and clinical manifestations. Front Med 9:915179. https://doi.org/10.3389/fmed.2022.915179
Azzouz D, Omarbekova A, Heguy A et al (2019) Lupus nephritis is linked to disease-activity associated expansions and immunity to a gut commensal. Ann Rheum Dis 78:947–956. https://doi.org/10.1136/annrheumdis-2018-214856
Toumi E, Goutorbe B, Plauzolles A et al (2022) Gut microbiota in systemic lupus erythematosus patients and lupus mouse model: a cross species comparative analysis for biomarker discovery. Front Immunol 13:943241. https://doi.org/10.3389/fimmu.2022.943241
Gerges MA, Esmaeel NE, Makram WK et al (2021) Altered profile of fecal microbiota in newly diagnosed systemic lupus erythematosus Egyptian patients. Int J Microbiol 2021:9934533. https://doi.org/10.1155/2021/9934533
Li Y, Wang H-F, Li X et al (1979) (2019) Disordered intestinal microbes are associated with the activity of Systemic Lupus Erythematosus. Clin Sci Lond Engl 133:821–838. https://doi.org/10.1042/CS20180841
López P, de Paz B, Rodríguez-Carrio J et al (2016) Th17 responses and natural IgM antibodies are related to gut microbiota composition in systemic lupus erythematosus patients. Sci Rep 6:24072. https://doi.org/10.1038/srep24072
Guo M, Wang H, Xu S et al (2020) Alteration in gut microbiota is associated with dysregulation of cytokines and glucocorticoid therapy in systemic lupus erythematosus. Gut Microb 11:1758–1773. https://doi.org/10.1080/19490976.2020.1768644
Allanore Y, Simms R, Distler O et al (2015) Systemic sclerosis. Nat Rev Dis Primer 1:15002. https://doi.org/10.1038/nrdp.2015.2
McMahan ZH (2019) Gastrointestinal involvement in systemic sclerosis: an update. Curr Opin Rheumatol 31:561–568. https://doi.org/10.1097/BOR.0000000000000645
Feng X, Li X-Q, Jiang Z (2021) Prevalence and predictors of small intestinal bacterial overgrowth in systemic sclerosis: a systematic review and meta-analysis. Clin Rheumatol 40:3039–3051. https://doi.org/10.1007/s10067-020-05549-8
Caimmi C, Caramaschi P, Venturini A et al (2018) Malnutrition and sarcopenia in a large cohort of patients with systemic sclerosis. Clin Rheumatol 37:987–997. https://doi.org/10.1007/s10067-017-3932-y
Volkmann ER, Chang Y-L, Barroso N et al (2016) Association of systemic sclerosis with a unique colonic microbial consortium. Arthritis Rheumatol Hoboken NJ 68:1483–1492. https://doi.org/10.1002/art.39572
Andréasson K, Alrawi Z, Persson A et al (2016) Intestinal dysbiosis is common in systemic sclerosis and associated with gastrointestinal and extraintestinal features of disease. Arthritis Res Ther 18:278. https://doi.org/10.1186/s13075-016-1182-z
Volkmann ER, Hoffmann-Vold A-M, Chang Y-L et al (2017) Systemic sclerosis is associated with specific alterations in gastrointestinal microbiota in two independent cohorts. BMJ Open Gastroenterol 4:e000134. https://doi.org/10.1136/bmjgast-2017-000134
Patrone V, Puglisi E, Cardinali M et al (2017) Gut microbiota profile in systemic sclerosis patients with and without clinical evidence of gastrointestinal involvement. Sci Rep 7:14874. https://doi.org/10.1038/s41598-017-14889-6
Wang Y, Wei J, Zhang W et al (2022) Gut dysbiosis in rheumatic diseases: a systematic review and meta-analysis of 92 observational studies. EBioMedicine 80:104055. https://doi.org/10.1016/j.ebiom.2022.104055
Tan TC, Chandrasekaran L, Leung YY et al (2023) Gut microbiome profiling in systemic sclerosis: a metagenomic approach. Clin Exp Rheumatol 41:1578–1588. https://doi.org/10.55563/clinexprheumatol/jof7nx
Yong WC, Upala S, Sanguankeo A (2018) Helicobacter pylori infection in systemic sclerosis: a systematic review and meta-analysis of observational studies. Clin Exp Rheumatol 36(Suppl 113):168–174
Efthymiou G, Liaskos C, Simopoulou T et al (2020) Antigen-specific humoral responses against Helicobacter pylori in patients with systemic sclerosis. Immunol Res 68:39–47. https://doi.org/10.1007/s12026-020-09124-w
Bellocchi C, Volkmann ER (2018) Update on the gastrointestinal microbiome in systemic sclerosis. Curr Rheumatol Rep 20:49. https://doi.org/10.1007/s11926-018-0758-9
Pittman N, Rawn SM, Wang M et al (2018) Treatment of small intestinal bacterial overgrowth in systemic sclerosis: a systematic review. Rheumatol Oxf Engl 57:1802–1811. https://doi.org/10.1093/rheumatology/key175
Frech TM, Khanna D, Maranian P et al (2011) Probiotics for the treatment of systemic sclerosis-associated gastrointestinal bloating/distention. Clin Exp Rheumatol 29:S22-25
Marighela TF, Arismendi MI, Marvulle V et al (2019) Effect of probiotics on gastrointestinal symptoms and immune parameters in systemic sclerosis: a randomized placebo-controlled trial. Rheumatol Oxf Engl 58:1985–1990. https://doi.org/10.1093/rheumatology/kez160
Mehta H, Goulet P-O, Mashiko S et al (2017) Early-life antibiotic exposure causes intestinal dysbiosis and exacerbates skin and lung pathology in experimental systemic sclerosis. J Invest Dermatol 137:2316–2325. https://doi.org/10.1016/j.jid.2017.06.019
Gómez-Hurtado I, Santacruz A, Peiró G et al (2011) Gut microbiota dysbiosis is associated with inflammation and bacterial translocation in mice with CCl4-induced fibrosis. PLoS ONE 6:e23037. https://doi.org/10.1371/journal.pone.0023037
Chioma OS, Mallott E, Shah-Gandhi B et al (2023) Low gut microbial diversity augments estrogen-driven pulmonary fibrosis in female-predominant interstitial lung disease. Cells 12:766. https://doi.org/10.3390/cells12050766
Acknowledgements
We thank Dr Marta Fraccaro for revising the English language in the text and Dr Francesca Pistillo for help in reviewing the literature for the spondylarthritis section.
Funding
No funding to declare.
Author information
Authors and Affiliations
Contributions
RB and MR were responsible for the conceptualisation of the study. RB and DB were responsible for writing the first draft. RB, DB, EB and AM were responsible for carrying out the review for the different sections. All authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
All authors have completed the COI disclosure form.
Statements on human and animal rights
This article does not contain any studies with human participants or animals performed by any of the authors.
Informed consent
None.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Bixio, R., Bertelle, D., Bertoldo, E. et al. The potential pathogenic role of gut microbiota in rheumatic diseases: a human-centred narrative review. Intern Emerg Med (2023). https://doi.org/10.1007/s11739-023-03496-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11739-023-03496-1