Abstract
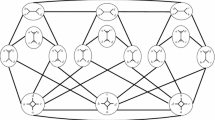
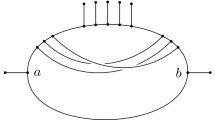
Suppose G is a phylogenetic network given as a rooted acyclic directed graph. Let X be a subset of the vertex set containing the root, all leaves, and all vertices of outdegree 1. A vertex is “regular” if it has a unique parent, and “hybrid” if it has two parents. Consider the case where each gene is binary. Assume an idealized system of inheritance in which no homoplasies occur at regular vertices, but homoplasies can occur at hybrid vertices. Under our model, the distances between taxa are shown to be described using a system of numbers called “originating weights” and “homoplasy weights.” Assume that the distances are known between all members of X. Sufficient conditions are given such that the graph G and all the originating and homoplasy weights can be reconstructed from the given distances.
Similar content being viewed by others
References
Bandelt, H.-J., Dress, A., 1992. Split decomposition: a new and useful approach to phylogenetic analysis of distance data. Mol. Phylogenet. Evol. 1, 242–252.
Baroni, M., Semple, C., Steel, M., 2004. A framework for representing reticulate evolution. Ann. Comb. 8, 391–408.
Bordewich, M., Semple, C., 2007. Computing the minimum number of hybridization events for a consistent evolutionary history. Discret. Appl. Math. 155, 914–928.
Felsenstein, J., 2004. Inferring Phylogenies. Sinauer, Sunderland.
Fitch, W.M., 1981. A non-sequential method for constructing trees and hierarchical classifications. J. Mol. Evol. 18, 30–37.
Gusfield, D., Eddhu, S., Langley, C., 2004a. Optimal, efficient reconstruction of phylogenetic networks with constrained recombination. J. Bioinform. Comput. Biol. 2, 173–213.
Gusfield, D., Eddhu, S., Langley, C., 2004b. The fine structure of galls in phylogenetic networks. INFORMS J. Comput. 16(4), 459–469.
Hein, J., 1990. Reconstructing evolution of sequences subject to recombination using parsimony. Math. Biosci. 98, 185–200.
Hein, J., 1993. A heuristic method to reconstruct the history of sequences subject to recombination. J. Mol. Evol. 36, 396–405.
Li, W.-H., 1981. A simple method for constructing phylogenetic trees from distance matrices. Proc. Natl. Acad. Sci. USA 78, 1085–1089.
Moret, B.M.E., Nakhleh, L., Warnow, T., Linder, C.R., Tholse, A., Padolina, A., Sun, J., Timme, R., 2004. Phylogenetic networks: modeling, reconstructibility, and accuracy. IEEE/ACM Trans. Comput. Biol. Bioinform. 1, 13–23.
Nakhleh, L., Warnow, T., Linder, C.R., 2004. Reconstructing reticulate evolution in species-theory and practice. In: Bourne, P.E., Gusfield, D. (Eds.), Proceedings of the Eighth Annual International Conference on Computational Molecular Biology, pp. 337–346. RECOMB’04, San Diego, California, 27–31 March 2004. ACM, New York.
Saitou, N., Nei, M., 1987. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 4, 406–425.
Sattath, S., Tversky, A., 1977. Additive similarity trees. Psychometrika 42, 319–345.
Sokal, R.R., Sneath, P.H.A., 1963. Numerical Taxonomy. Freeman, San Francisco.
Wang, L., Zhang, K., Zhang, L., 2001. Perfect phylogenetic networks with recombination. J. Comput. Biol. 8, 69–78.
Willson, S.J., 2006. Unique reconstruction of tree-like phylogenetic networks from distances between leaves. Bull. Math. Biol. 68, 919–944.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Willson, S.J. Reconstruction of Some Hybrid Phylogenetic Networks with Homoplasies from Distances. Bull. Math. Biol. 69, 2561–2590 (2007). https://doi.org/10.1007/s11538-007-9232-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11538-007-9232-y