Abstract
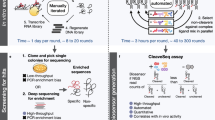
Small molecule aptamers discovered by traditional selection methods usually lack conformational changes upon target binding. This limits the use of aptamers as molecular probes for small molecule detection and regulatory elements of genetic circuits. Here, we report a new method called capture and in vitro transcription-systematic evolution of ligands by exponential enrichment (CIVT-SELEX) to select DNA aptamers that can not only bind to small molecule ligands but also undergo significant conformational changes. Through this method, we select a structure-switching aptamer of uridine-5′-diphosphate (UDP). Taking advantage of its conformational changes, we first construct a UDP-responsive transcriptional switch by inserting the aptamer in a genetic circuit and demonstrate that it can respond to the addition of UDP and regulate the transcription of downstream genes. We also build a UDP aptamer-based biosensor that can be used for active glycosyltransferase screening. We believe this method can provide a universal platform for selecting small molecule aptamers with conformational changes and expand the use of aptamers in small molecule detection and genetic regulation.

Similar content being viewed by others
References
Wang X, Zhang XJ, Li Y, Zhang GR, Li J, Wang XQ, Tan W. CCS Chem, 2022, 4: 2581–2587
Li L, Xu S, Yan H, Li X, Yazd HS, Li X, Huang T, Cui C, Jiang J, Tan W. Angew Chem Int Ed, 2021, 60: 2221–2231
Huang R, Xi Z, He N. Sci China Chem, 2015, 58: 1122–1130
Tian L, Shao M, Gong Y, Chao Y, Wei T, Yang K, Chen Q, Liu Z. Sci China Chem, 2022, 65: 574–583
Wang Z, Huang C, Sun N, Deng C. Sci China Chem, 2021, 64: 932–947
Liu M, Zhang M, Chen J, Yang R, Huang Z, Liu Z, Li N, Shui L. Sci China Chem, 2022, 65: 2023–2030
Tuerk C, Gold L. Science, 1990, 249: 505–510
Yu H, Alkhamis O, Canoura J, Liu Y, Xiao Y. Angew Chem Int Ed, 2021, 60: 16800–16823
Huang PJJ, Liu J. ACS Chem Biol, 2022, 17: 2121–2129
Nutiu R, Li Y. J Am Chem Soc, 2003, 125: 4771–4778
Stojanovic MN, de Prada P, Landry DW. J Am Chem Soc, 2001, 123: 4928–4931
Wang J, Yang L, Cui X, Zhang Z, Dong L, Guan N. ACS Synth Biol, 2017, 6: 758–765
Hoetzel J, Suess B. J Mol Biol, 2022, 434: 167631
Ma Q, Zhang C, Zhang M, Han D, Tan W. Small Struct, 2021, 2: 2100051
Guo S, Xu Z, Lin L, Guo Y, Li J, Lu C, Shi X, Yang H. Int J Mol Sci, 2023, 24: 2833
Mendonsa SD, Bowser MT. J Am Chem Soc, 2004, 126: 20–21
Nguyen VT, Kwon YS, Kim JH, Gu MB. Chem Commun, 2014, 50: 10513–10516
Spiga FM, Maietta P, Guiducci C. ACS Comb Sci, 2015, 17: 326–333
Boussebayle A, Torka D, Ollivaud S, Braun J, Bofill-Bosch C, Dombrowski M, Groher F, Hamacher K, Suess B. Nucleic Acids Res, 2019, 47: 4883–4895
Huang M, Song J, Huang P, Chen X, Wang W, Zhu Z, Song Y, Yang C. Anal Chem, 2019, 91: 10879–10886
Lyu C, Khan IM, Wang Z. Talanta, 2021, 229: 122274
Bowles D, Isayenkova J, Lim EK, Poppenberger B. Curr Opin Plant Biol, 2005, 8: 254–263
Guo S, Wang M, Xu W, Zou F, Lin J, Peng Q, Xu W, Xu S, Shi X. Phytochemistry, 2022, 193: 113007
Boussebayle A, Groher F, Suess B. Methods, 2019, 161: 10–15
Alam KK, Tawiah KD, Lichte MF, Porciani D, Burke DH. ACS Synth Biol, 2017, 6: 1710–1721
McKeague M, Velu R, Hill K, Bardóczy V, Mészáros T, DeRosa M. Toxins, 2014, 6: 2435–2452
Liu X, Lu Q, Chen S, Wang F, Hou J, Xu Z, Meng C, Hu T, Hou Y. Molecules, 2018, 23: 2337
Alkhamis O, Yang W, Farhana R, Yu H, Xiao Y. Nucleic Acids Res, 2020, 48: e120
Liu J, Lu Y. Angew Chem Int Ed, 2005, 45: 90–94
Zhang Z, Oni O, Liu J. Nucleic Acids Res, 2017, 45: 7593–7601
Kypr J, Kejnovska I, Renciuk D, Vorlickova M. Nucleic Acids Res, 2009, 37: 1713–1725
Sengupta A, Gavvala K, Koninti RK, Hazra P. J Photochem Photobiol B-Biol, 2014, 140: 240–248
Canoura J, Wang Z, Yu H, Alkhamis O, Fu F, Xiao Y. J Am Chem Soc, 2018, 140: 9961–9971
Canoura J, Yu H, Alkhamis O, Roncancio D, Farhana R, Xiao Y. J Am Chem Soc, 2021, 143: 805–816
Acknowledgements
We would like to thank Dr. J. Yang, Dr. Y. Zheng, Dr. F. Li, Dr. M. Chen, and Dr. L. Chen for their helpful discussion and suggestions. This work was supported by the National Natural Science Foundation of China (32001037, 22176035), the National Key R&D Program of China (2020YFA0210800, 2018YFA0902600), the Natural Science Foundation of Fujian Province (2020J01491, 2020J05120), and Fuzhou University Research Fund (GXRC-20033).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Conflict of interest
The authors declare no conflict of interest.
Supporting information
The supporting information is available online at http://chem.scichina.com and http://link.springer.com/journal/11426. The supporting materials are published as submitted, without typesetting or editing. The responsibility for scientific accuracy and content remains entirely with the authors.
Supporting information
11426_2022_1540_MOESM1_ESM.pdf
Selecting Small Molecule DNA Aptamers with Significant Conformational Changes for Constructing Transcriptional Switches and Biosensors
Rights and permissions
About this article
Cite this article
Guo, S., Lin, J., Lin, L. et al. Selecting small molecule DNA aptamers with significant conformational changes for constructing transcriptional switches and biosensors. Sci. China Chem. 66, 1529–1536 (2023). https://doi.org/10.1007/s11426-022-1540-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11426-022-1540-y