Abstract
Introduction
Mitochondria represent an important milieu for studying the pathogenesis of several major diseases. The need for organelle-level metabolic resolution exists, as mitochondrial/cytosolic metabolites are often diluted beyond detection limits in complex samples. Compartment-specific studies are still hindered by the lack of efficient, cost-effective fractioning methods—applicable to laboratories of all financial/analytical standing.
Objectives
We established a novel mitochondrial/cytosolic purification pipeline for complimentary GC-TOF–MS and 1H-NMR metabolomics using robust, commercially available fractionation strategies.
Methods
Magnetic based mitochondria isolation kits (MACS) were adapted for this purpose, accompanied by cytosolic filtering. Yield was assessed through the percentage recovery of citrate synthase (CS; a mitochondrial marker), purity by immunoblotting against compartment-specific proteins and integrity interrogated through the respiratory coupling ratio (RCR). The effects of the kit-based buffers on MS/NMR analyses of pure metabolite standards were evaluated. Finally, biological applicability to mammalian disease models was shown using Ndufs4 mouse brain tissue.
Results
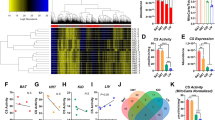
With minor modifications, MACS produced around 60% more mitochondria compared to a differential centrifugation method. Less than 15% of lysosomal LAMP-2 protein was found in the MACS isolates, confirming relative purity—while RCR’s above 6 indicate sufficient mitochondrial integrity. The filtering approach effectively depleted mitochondria from the cytosolic fraction, as indicated by negligible Hsp60 and CS levels. Our GC–MS pilot yielded 60–70 features per fraction, while NMR analyses could quantify 6–10 of the most abundant compounds in each fraction.
Conclusion
This study provides a simple and flexible solution for mitochondrial and cytosolic metabolomics in animal model tissues, towards large-scale application of such methodologies in disease research.



Similar content being viewed by others
Data availability
The metabolomics and metadata reported in this paper are available via Metabolomics Workbench, https://www.metabolomicsworkbench.org with unique file identifier Gunter007dekok_20191218_151540 and Data Track ID 1877.
References
Afanasyeva, M., Ustiugova, A., Golyshev, S., Kopylov, A., Bogolyubova, A., Demin, D., et al. (2018). Isolation of large amounts of highly pure mitochondria for “omics” studies. Biochemistry (Moscow),83, 76–85.
Aucamp, J., Bronkhorst, A. J., Peters, D. L., Van Dyk, H. C., Van der Westhuizen, F. H., & Pretorius, P. J. (2017). Kinetic analysis, size profiling, and bioenergetic association of DNA released by selected cell lines in vitro. Cellular and Molecular Life Sciences,74, 2689–2707.
Bayraktar, E. C., Baudrier, L., Özerdem, C., Lewis, C. A., Chan, S. H., Kunchok, T., et al. (2019). MITO-Tag Mice enable rapid isolation and multimodal profiling of mitochondria from specific cell types in vivo. Proceedings of the National Academy of Sciences,116, 303–312.
Beckonert, O., Keun, H. C., Ebbels, T. M., Bundy, J., Holmes, E., Lindon, J. C., et al. (2007). Metabolic profiling, metabolomic and metabonomic procedures for NMR spectroscopy of urine, plasma, serum and tissue extracts. Nature Protocols,2, 2692.
Birsoy, K., Wang, T., Chen, W. W., Freinkman, E., Abu-Remaileh, M., & Sabatini, D. M. (2015). An essential role of the mitochondrial electron transport chain in cell proliferation is to enable aspartate synthesis. Cell,162, 540–551.
Bissantz, C., Kuhn, B., & Stahl, M. (2010). A medicinal chemist's guide to molecular interactions. Journal of Medicinal Chemistry,53, 5061–5084.
Brand, M. D., & Nicholls, D. G. (2011). Assessing mitochondrial dysfunction in cells. Biochemical Journal,435, 297–312.
Brown, M.R., Sullivan, P.G., Dorenbos, K.A., Modafferi, E.A., Geddes, J.W., & Steward, O. (2004). Nitrogen disruption of synaptoneurosomes: An alternative method to isolate brain mitochondria. Journal of Neuroscience Methods,137, 299–303.
Chan, C. C., Lee, Y., Lam, H., & Zhang, X.-M. (2004). Analytical method validation and instrument performance verification. New York: Wiley.
Chen, W. W., Freinkman, E., Wang, T., Birsoy, K., & Sabatini, D. M. (2016). Absolute quantification of matrix metabolites reveals the dynamics of mitochondrial metabolism. Cell,166(1324–1337), e11.
Cheng, M. Y., Hartl, F.-U., Martin, J., Pollock, R. A., Kalousek, F., Neuper, W., et al. (1989). Mitochondrial heat-shock protein hsp60 is essential for assembly of proteins imported into yeast mitochondria. Nature Communications,337, 620.
Corcelli, A., Saponetti, M. S., Zaccagnino, P., Lopalco, P., Mastrodonato, M., Liquori, G. E., et al. (2010). Mitochondria isolated in nearly isotonic KCl buffer: Focus on cardiolipin and organelle morphology. Biochimica et Biophysica Acta,1798, 681–687.
de Bari, L., & Atlante, A. (2018). Including the mitochondrial metabolism of l-lactate in cancer metabolic reprogramming. Cellular and Molecular Life Sciences,75, 2763–2776.
Dona, A. C., Jiménez, B., Schäfer, H., Humpfer, E., Spraul, M., Lewis, M. R., et al. (2014). Precision high-throughput proton NMR spectroscopy of human urine, serum, and plasma for large-scale metabolic phenotyping. Analytical Chemistry,86, 9887–9894.
Dunn, W. B., Broadhurst, D. I., Atherton, H. J., Goodacre, R., & Griffin, J. L. (2011). Systems level studies of mammalian metabolomes: The roles of mass spectrometry and nuclear magnetic resonance spectroscopy. Chemical Society Reviews,40, 387–426.
Dunn, W. B., Wilson, I. D., Nicholls, A. W., & Broadhurst, D. (2012). The importance of experimental design and QC samples in large-scale and MS-driven untargeted metabolomic studies of humans. Bioanalysis,4, 2249–2264.
Esterhuizen, K., van der Westhuizen, F. H., & Louw, R. (2017). Metabolomics of mitochondrial disease. Mitochondrion,35, 97–110.
FDA, U.S. (2001) Guidance for industry, bioanalytical method validation. Federal Register 66.
Fiehn, O. (2002). Metabolomics—The link between genotypes and phenotypes. In C. Town (Ed.), Functional genomics (pp. 155–171). Dordrecht: Springer.
Franko, A., Baris, O. R., Bergschneider, E., Von Toerne, C., Hauck, S. M., Aichler, M., et al. (2013). Efficient isolation of pure and functional mitochondria from mouse tissues using automated tissue disruption and enrichment with anti-TOM22 magnetic beads. PLoS ONE,8, e82392.
Fratila, R. M., & Velders, A. H. (2011). Small-volume nuclear magnetic resonance spectroscopy. Annual Review of Analytical Chemistry,4, 227–249.
Frezza, C. (2017). Mitochondrial metabolites: Undercover signalling molecules. Interface Focus,7, 20160100.
García-Sevillano, M., García-Barrera, T., Navarro, F., Montero-Lobato, Z., & Gómez-Ariza, J. (2015). Shotgun metabolomic approach based on mass spectrometry for hepatic mitochondria of mice under arsenic exposure. BioMetals,28, 341–351.
Gielisch, I., & Meierhofer, D. (2015). Metabolome and proteome profiling of complex I deficiency induced by rotenone. Journal of Proteome Research,14, 224–235.
Glaves, J., Li, M., Mercier, P., Fahlman, R., & Sykes, B. (2014). High-throughput, multi-platform metabolomics on very small volumes: 1 H NMR metabolite identification in an unadulterated tube-in-tube system. Metabolomics,10, 1145–1151.
Go, Y.-M., Roede, J. R., Orr, M., Liang, Y., & Jones, D. P. (2014a). Integrated redox proteomics and metabolomics of mitochondria to identify mechanisms of cd toxicity. Toxicological Sciences,139, 59–73.
Go, Y.-M., Uppal, K., Walker, D.I., Tran, V., Dury, L., Strobel, F.H., Baubichon-Cortay, H., Pennell, K.D., Roede, J.R. and Jones, D.P. (2014b) Mitochondrial metabolomics using high-resolution Fourier-transform mass spectrometry. In Mass spectrometry in metabolomics (pp. 43–73). New York: Springer.
Gullberg, J., Jonsson, P., Nordström, A., Sjöström, M., & Moritz, T. (2004). Design of experiments: An efficient strategy to identify factors influencing extraction and derivatization of Arabidopsis thaliana samples in metabolomic studies with gas chromatography/mass spectrometry. Analytical Biochemistry,331, 283–295.
Hall, A. M., & Roberson, E. D. (2012). Mouse models of Alzheimer's disease. Brain Research Bulletin,88, 3–12.
Hohmann, M., Felbinger, C., Christoph, N., Wachter, H., Wiest, J., & Holzgrabe, U. (2014). Quantification of taurine in energy drinks using 1H NMR. Journal of Pharmaceutical and Biomedical Analysis,93, 156–160.
Hornig-Do, H.-T., Günther, G., Bust, M., Lehnartz, P., Bosio, A., & Wiesner, R. J. (2009). Isolation of functional pure mitochondria by superparamagnetic microbeads. Analytical Biochemistry,389, 1–5.
Hubbard, W. B., Harwood, C. L., Prajapati, P., Springer, J. E., Saatman, K. E., & Sullivan, P. G. (2019). Fractionated mitochondrial magnetic separation for isolation of synaptic mitochondria from brain tissue. Science Report,9, 9656.
Jin, F., Wu, Q., Lu, Y. F., Gong, Q. H., & Shi, J. S. (2008). Neuroprotective effect of resveratrol on 6-OHDA-induced Parkinson's disease in rats. European Journal of Pharmacology,600, 78–82.
Junghans, L., Teleki, A., Wijaya, A. W., Becker, M., Schweikert, M., & Takors, R. (2019). From nutritional wealth to autophagy: In vivo metabolic dynamics in the cytosol, mitochondrion and shuttles of IgG producing CHO cells. Metabolic Engineering,54, 145–159.
Kappler, L., Li, J., Häring, H.-U., Weigert, C., Lehmann, R., Xu, G., et al. (2016). Purity matters: A workflow for the valid high-resolution lipid profiling of mitochondria from cell culture samples. Scientific Reports,6, 21107.
Kayser, E. B., Sedensky, M. M., & Morgan, P. G. (2016). Region-specific defects of respiratory capacities in the Ndufs4(KO) mouse brain. PLoS ONE,11, e0148219.
Kruse, S. E., Watt, W. C., Marcinek, D. J., Kapur, R. P., Schenkman, K. A., & Palmiter, R. D. (2008). Mice with mitochondrial complex I deficiency develop a fatal encephalomyopathy. Cell Metabolism,7, 312–320.
Lake, N. J., Bird, M. J., Isohanni, P., & Paetau, A. (2015). Leigh syndrome: Neuropathology and pathogenesis. Journal of Neuropathology and Experimental Neurology,74, 482–492.
Lee, W. D., Mukha, D., Aizenshtein, E., & Shlomi, T. (2019). Spatial-fluxomics provides a subcellular-compartmentalized view of reductive glutamine metabolism in cancer cells. Nature Communications,10, 1351.
Lindon, J. C., Nicholson, J. K., & Everett, J. R. (1999). NMR spectroscopy of biofluids (pp. 1–88). Annual reports on NMR spectroscopy: Elsevier.
Lindeque, J.Z., Hidalgo, J, Louw, R., & van der Westhuizen, F.H. (2013). Systemic and organ specific metabolic differences in metallothionein knockout mice when challenged with exercise. Metabolomics,9, 418–432.
Liu, X., & Xu, G. (2018). Recent advances in using mass spectrometry for mitochondrial metabolomics and lipidomics—A review. Analytica Chimica Acta,1037, 3–12.
Luo, R., David, L., Hung, H., Devaney, J., & Gilson, M. K. (1999). Strength of solvent-exposed salt-bridges. The Journal of Physical Chemistry B,103, 727–736.
Mamer, O., Gravel, S.-P., Choinière, L., Chénard, V., St-Pierre, J., & Avizonis, D. (2013). The complete targeted profile of the organic acid intermediates of the citric acid cycle using a single stable isotope dilution analysis, sodium borodeuteride reduction and selected ion monitoring GC/MS. Metabolomics,9, 1019–1030.
Mason, S., Terburgh, K., & Louw, R. (2018). Miniaturized 1H-NMR method for analyzing limited-quantity samples applied to a mouse model of Leigh disease. Metabolomics,14, 74.
Matuszczyk, J.-C., Teleki, A., Pfizenmaier, J., & Takors, R. (2015). Compartment-specific metabolomics for CHO reveals that ATP pools in mitochondria are much lower than in cytosol. Biotechnology Journal,10, 1639–1650.
Moffett, J. R., Ross, B., Arun, P., Madhavarao, C. N., & Namboodiri, A. M. (2007). N-Acetylaspartate in the CNS: From neurodiagnostics to neurobiology. Progress in neurobiology,81, 89–131.
Nagrath, D., Caneba, C., Karedath, T., & Bellance, N. (2011). Metabolomics for mitochondrial and cancer studies. Biochimica et Biophysica Acta Bioenergetics,1807, 650–663.
Nunnari, J., & Suomalainen, A. (2012). Mitochondria: in sickness and in health. Cell,148, 1145–1159.
Palmieri, F., & Monne, M. (2016). Discoveries, metabolic roles and diseases of mitochondrial carriers: A review. Biochimica et Biophysica Acta,1863, 2362–2378.
Pan, D., Lindau, C., Lagies, S., Wiedemann, N., & Kammerer, B. (2018). Metabolic profiling of isolated mitochondria and cytoplasm reveals compartment-specific metabolic responses. Metabolomics,14, 59.
Passarella, S., & Schurr, A. (2018). l-Lactate transport and metabolism in mitochondria of Hep G2 cells—The cori cycle revisited. Frontiers in Oncology,8, 120.
Patti, M.-E., & Corvera, S. (2010). The role of mitochondria in the pathogenesis of type 2 diabetes. Endocrine Reviews,31, 364–395.
Paul, M. H., & Sperling, E. (1952). Cyclophorase system. XXIII. Correlation of cyclophorase activity and mitochondrial density in striated muscle. Proceedings of the Society for Experimental Biology and Medicine,79, 352–354.
Picard, M., Wallace, D. C., & Burelle, Y. (2016). The rise of mitochondria in medicine. Mitochondrion,30, 105–116.
Picard, M., White, K., & Turnbull, D. M. (2012). Mitochondrial morphology, topology, and membrane interactions in skeletal muscle: A quantitative three-dimensional electron microscopy study. Journal of Applied Physiology,114, 161–171.
Pustylnikov, S., Costabile, F., Beghi, S., & Facciabene, A. (2018). Targeting mitochondria in cancer: Current concepts and immunotherapy approaches. Transl Res,202, 35–51.
Roede, J. R., Park, Y., Li, S., Strobel, F. H., & Jones, D. P. (2012). Detailed mitochondrial phenotyping by high resolution metabolomics. PLoS ONE,7, e33020.
Ruggeri, B. A., Camp, F., & Miknyoczki, S. (2014). Animal models of disease: Pre-clinical animal models of cancer and their applications and utility in drug discovery. Biochemical Pharmacology,87, 150–161.
Schymanski, E. L., Jeon, J., Gulde, R., Fenner, K., Ruff, M., Singer, H. P., et al. (2014). Identifying small molecules via high resolution mass spectrometry: Communicating confidence. Environmental Science and Technology,48, 2097–2098.
Shepherd, D., & Garland, P. (1969). The kinetic properties of citrate synthase from rat liver mitochondria. Biochemical Journal,114, 597–610.
Steinhauser, M. L., & Lechene, C. P. (2013). Quantitative imaging of subcellular metabolism with stable isotopes and multi-isotope imaging mass spectrometry. Seminars in Cell & Developmental Biology,24, 661–667.
Tahrir, F. G., Shanmughapriya, S., Ahooyi, T. M., Knezevic, T., Gupta, M. K., Kontos, C. D., et al. (2018). Dysregulation of mitochondrial bioenergetics and quality control by HIV-1 Tat in cardiomyocytes. Journal of Cellular Physiology,233, 748–758.
Terburgh, K., Lindeque, Z., Mason, S., Van der Westhuizen, F., & Louw, R. (2019). Metabolomics of Ndufs4−/− skeletal muscle: Adaptive mechanisms converge at the ubiquinone-cycle. Biochimica et Biophysica Acta,1865, 98–106.
Tsai, C. (1967). Spontaneous decarboxylation of oxalacetic acid. Canadian Journal of Chemistry,45, 873–880.
Tsai, G., & Coyle, J. T. (1995). N-acetylaspartate in neuropsychiatric disorders. Progress in Neurobiology,46, 531–540.
Tzelepis, F., Blagih, J., Khan, N., Gillard, J., Mendonca, L., Roy, D.G., Ma, E.H., Joubert, P., Jones, R.G., and Divangahi, M. (2018) Mitochondrial cyclophilin D regulates T cell metabolic responses and disease tolerance to tuberculosis. Science Immunology3.
Van Aardt, W. J., Le Roux, J. M., Lindeque, J. Z., Mason, S., & Louw, R. (2016). The effect of temperature on the respiration and metabolism of the African burrowing scorpion (Opistophthalmus latimanus). Comparative Biochemistry and Physiology Part D: Genomics and Proteomics,20, 50–56.
Van Vranken, J. G., & Rutter, J. (2016). The whole (cell) is less than the sum of its parts. Cell,166, 1078–1079.
Vendelin, M., Béraud, N., Guerrero, K., Andrienko, T., Kuznetsov, A. V., Olivares, J., et al. (2005). Mitochondrial regular arrangement in muscle cells: A “crystal-like” pattern. American Journal of Physiology-Cell Physiology,288, C757–C767.
Wang, A. C., & Bax, A. (1993). Minimizing the effects of radio-frequency heating in multidimensional NMR experiments. Journal of biomolecular NMR,3, 715–720.
Warrack, B. M., Hnatyshyn, S., Ott, K. H., Reily, M. D., Sanders, M., Zhang, H., et al. (2007). Normalization strategies for metabonomic analysis of urine samples. Journal of Chromatography B. Analytical Technologies in the Biomedical and Life Sciences,877, 547–552.
Wu, M., Neilson, A., Swift, A. L., Moran, R., Tamagnine, J., Parslow, D., et al. (2007). Multiparameter metabolic analysis reveals a close link between attenuated mitochondrial bioenergetic function and enhanced glycolysis dependency in human tumor cells. American Journal of Physiology: Cell Physiology,292, C125–C136.
Yao, J., Irwin, R. W., Zhao, L., Nilsen, J., Hamilton, R. T., & Brinton, R. D. (2009). Mitochondrial bioenergetic deficit precedes Alzheimer's pathology in female mouse model of Alzheimer's disease. Proceedings of the National Academy of Sciences of the USA,106, 14670–14675.
Acknowledgements
This work is based on the research supported by the National Research Foundation of South Africa (NRF; Grant Numbers 92736 and 117424) and the Technological Innovation Agency of the Department of Science and Technology of South Africa (TIA; Grant Number Metabol.01). Opinions expressed and conclusions arrived at, are those of the authors and are not necessarily to be attributed to the NRF or the TIA. We acknowledge and thank Dr’s Shayne Mason and J Zander Lindeque of the NWU, Human Metabolomics, for their invaluable assistance during metabolomics and data analyses, as well as Mr’s Luciano WIllemse and Charl du Plessis for helping in acquiring chemometrics data for the chemical standard mixtures investigated in this study.
Author information
Authors and Affiliations
Contributions
The study was conceived by RL, but planned and executed in equal parts by both authors. Data were acquired predominantly by GV and analysed concomitantly by both authors. The manuscript was compiled by GV under supervision of RL.
Corresponding author
Ethics declarations
Conflict of interest
Both authors declare to have no conflicts of interest.
Ethical approval
The AnimCare animal research ethics committee of North-West University approved (NWU-00568-19-S5) the animal protocols used here. All animals were maintained and all procedures performed in accordance with the code of ethics in research, training and testing of drugs in South Africa and complied with national legislation.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
van der Walt, G., Louw, R. Novel mitochondrial and cytosolic purification pipeline for compartment-specific metabolomics in mammalian disease model tissues. Metabolomics 16, 78 (2020). https://doi.org/10.1007/s11306-020-01697-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11306-020-01697-9



