Abstract
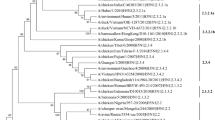
The highly pathogenic avian influenza (HPAI) H5N1 virus has received considerable attention during the past 2 decades due to its zoonotic and mutative features. This Virus is of special importance due to to the possibility of causing infection in human populations. According to it’s geographical location, Iran hosts a large number of aquatic migratory birds every year, and since these birds can be considered as the host of the H5 HPAI, the country is significantly at risk of this virus. the In this study, the molecular characteristics of hemagglutinin (HA) and neuraminidase (NA) genes of the H5N8 strain were identified in Malard county of Tehran province and Meighan wetland of Arak city, Markazi province were investigated. Based on the analysis of the amino acid sequence of the HA genes, the cleavage site of the gene includes the PLREKRRKR/GLF polybasic amino acid motif, which is a characteristic of highly pathogenic influenza viruses. The HA gene of two viruses had T156A, S123P, S133A mutations associated with the increased mammalian sialic acid-binding, and the NA gene of two viruses had H253Y mutations associated with the resistance to antiviral drugs. Phylogenetic analysis of the HA genes indicated the classification of these viruses in the 2.3.4.4 b subclade. Although the A/Goose/Iran/180/2016 virus was also an H5N8 2.3.4.4 b virus, its cluster was separated from the A/Chicken/Iran/162/2016 virus. This means that the entry of these viruses in to the country happened through more than one window. Furthermore, it seems that the introduction of these H5N8 HPAI strains in Iran probably occurred through the West Asia–East African flyway by wild migratory aquatic birds.





Similar content being viewed by others
References
Xu X, Subbarao K, Cox NJ, Guo Y (1999) Genetic characterization of the pathogenic influenza A/Goose/Guangdong/1/96 (H5N1) virus: similarity of its hemagglutinin gene to those of H5N1 viruses from the 1997 outbreaks in Hong Kong. Virology 261(1):15–19
Cattoli G, Fusaro A, Monne I, Capua I (2009) H5N1 virus evolution in Europe—an updated overview. Viruses 1(3):1351–1363
WHO, OIE, FAO, (2012) Continuedevolution of highly pathogenic avian influenza A (H5N1): updated nomenclature. Influenza Other Respir Viruses 6:1–5. https://doi.org/10.1111/j.1750-2659.2011.00298.x
Sun H, Pu J, Hu J, Liu L, Xu G, Gao GF et al (2016) Characterization of clade 2.3.4.4 highly pathogenic H5 avian influenza viruses in ducks and chickens. Vet Microbiol 182:116–22
Zhao K, Gu M, Zhong L, Duan Z, Zhang Y, Zhu Y et al (2013) Characterization of three H5N5 and one H5N8 highly pathogenic avian influenza viruses in China. Vet Microbiol 163(3–4):351–357
Li J, Gu M, Liu D, Liu B, Jiang K, Zhong L et al (2016) Phylogenetic and biological characterization of three K1203 (H5N8)-like avian influenza A virus reassortants in China in 2014. Adv Virol 161(2):289–302
Wu H, Peng X, Xu L, Jin C, Cheng L, Lu X et al (2014) Novel reassortant influenza A (H5N8) viruses in domestic ducks, eastern China. Emerg Infect Dis 20(8):1315
Lee EK, Lee YN, Song BM, Heo GB, Lee HS, Lee YJ. Pathogenesis of multiple subgroups of clade 2.3. 4.4. influenza A (H5N8) virus in mice and ferrets. 대한수의학회 학술대회발표집. 2016:274–5.
Vascellari M, Hablolvarid M, Shoushtari A, Hedayati A (2007) Mortality of wild swans associated with naturally infection with highly pathogenic H5N1 avian influenza virus in Iran. Arch Razi Inst 62(4):207–213
OIE. World Organisation for Animal Health, 2018. OIE situation report for avian influenza. http://www.oie.int/animal-healthin-the-world/update-on-avian-influenza/2018. Accessed 20 April 2022
OIE. World Organisation for Animal Health.Animal disease events. 2020. https://wahis.woah.org/#/events?viewAll=true. Accessed 23 April 2022
Fereidouni SR, Werner O, Starick E, Beer M, Harder TC, Aghakhan M et al (2010) Avian influenza virus monitoring in wintering waterbirds in Iran, 2003–2007. Virol J 7(1):1–14
Yegani S, Shoushtari A-H, Eshratabadi F, Molouki A (2019) Full sequence analysis of hemagglutinin and neuraminidase genes and proteins of highly pathogenic avian influenza H5N1 virus detected in Iran, 2015. Trop Anim Health Prod 51(3):605–612
Kord E, Kaffashi A, Ghadakchi H, Eshratabadi F, Bameri Z, Shoushtari A (2014) Molecular characterization of the surface glycoprotein genes of highly pathogenic H5N1 avian influenza viruses detected in Iran in 2011. Trop Anim Health Prod 46(3):549–554
Qi X, Li X, Rider P, Fan W, Gu H, Xu L et al (2009) Molecular characterization of highly pathogenic H5N1 avian influenza A viruses isolated from raccoon dogs in China. PLoS ONE 4(3):e4682
WHO. World Health Organization, 2018. Avian Influenza Weekly Update Number 631 (6 April 2018). Available at http://www.Wpro.who.int/emerging_diseases/AvianInfluenza/en/2018
OIE. World Organisation for Animal Health. Manual of Diagnostic Tests and Vaccines for Terrestrial Animals 2022. https://www.woah.org/en/what-we-do/standards/codes-and-manuals/terrestrial-manual-online-access/
Stear MJ (2005) OIE manual of diagnostic tests and vaccines for terrestrial animals (mammals, birds and bees) 5th edn volumes 1 & 2 World Organization for Animal Health 2004. Parasitology 130(6):727
Hoffimann E, Stech J, Guan Y, Webster R, Perez D (2001) Universal primer set for the full-length amplification of all influenza A viruses. Arch Virol 146(2275):89
Sanger F, Nicklen S, Coulson AR (1977) DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci 74(12):5463–5467
Slomka M, Pavlidis T, Banks J, Shell W, McNally A, Essen S et al (2007) Validated H5 Eurasian real-time reverse transcriptase–polymerase chain reaction and its application in H5N1 outbreaks in 2005–2006. Avian Dis 51(s1):373–377
Spackman E, Senne DA, Myers T, Bulaga LL, Garber LP, Perdue ML et al (2002) Development of a real-time reverse transcriptase PCR assay for type A influenza virus and the avian H5 and H7 hemagglutinin subtypes. J Clin Microbiol 40(9):3256–3260
Zhang Y, Aevermann BD, Anderson TK, Burke DF, Dauphin G, Gu Z et al (2017) Influenza research database: an integrated bioinformatics resource for influenza virus research. Nucleic Acids Res 45(D1):D466–D474
Gupta R, Brunak S (eds.) (2001) Prediction of glycosylation across the human proteome and the correlation to protein function. Pac Symp Biocomput.
de Graaf M, Fouchier RA (2014) Role of receptor binding specificity in influenza A virus transmission and pathogenesis. EMBO J 33(8):823–841
Skehel JJ, Wiley DC (2000) Receptor binding and membrane fusion in virus entry: the influenza hemagglutinin. Annu Rev Biochem 69:531–569
Ha Y, Stevens DJ, Skehel JJ, Wiley DC (2001) X-ray structures of H5 avian and H9 swine influenza virus hemagglutinins bound to avian and human receptor analogs. Proc Natl Acad Sci 98(20):11181–11186
Ohkawara A, Okamatsu M, Ozawa M, Chu DH, Nguyen LT, Hiono T et al (2017) Antigenic diversity of H5 highly pathogenic avian influenza viruses of clade 2.3.4.4 isolated in Asia. Microbiol Immunol 61(5):149–58
Ha Y, Stevens DJ, Skehel JJ, Wiley DC (2002) H5 avian and H9 swine influenza virus haemagglutinin structures: possible origin of influenza subtypes. Embo J 21(5):865–875
Kaverin NV, Rudneva IA, Govorkova EA, Timofeeva TA, Shilov AA, Kochergin-Nikitsky KS et al (2007) Epitope mapping of the hemagglutinin molecule of a highly pathogenic H5N1 influenza virus by using monoclonal antibodies. J Virol 81(23):12911–12917
Kaverin NV, Rudneva IA, Ilyushina NA, Varich NL, Lipatov AS, Smirnov YA et al (2002) Structure of antigenic sites on the haemagglutinin molecule of H5 avian influenza virus and phenotypic variation of escape mutants. J Gen Virol 83(10):2497–2505
Reynolds CR, Islam SA, Sternberg MJ (2018) EzMol: a web server wizard for the rapid visualization and image production of protein and nucleic acid structures. J Mol Biol 430(15):2244–2248
Yen H-L, Hoffmann E, Taylor G, Scholtissek C, Monto AS, Webster RG et al (2006) Importance of neuraminidase active-site residues to the neuraminidase inhibitor resistance of influenza viruses. J Virol 80(17):8787–8795
Xu X, Zhu X, Dwek RA, Stevens J, Wilson IA (2008) Structural characterization of the 1918 influenza virus H1N1 neuraminidase. J Virol 82(21):10493–10501
Colman PM, Hoyne PA, Lawrence MC (1993) Sequence and structure alignment of paramyxovirus hemagglutinin-neuraminidase with influenza virus neuraminidase. J Virol 67(6):2972–2980
Colman PM (1994) Influenza virus neuraminidase: structure, antibodies, and inhibitors. Protein Sci 3(10):1687–1696
Burmeister W, Ruigrok R, Cusack S (1992) The 2.2 a resolution crystal structure of influenza B neuraminidase and its complex with sialic acid. EMBO J 11(1):49–56
Saito T, Taylor G, Laver W, Kawaoka Y, Webster R (1994) Antigenicity of the N8 influenza A virus neuraminidase: existence of an epitope at the subunit interface of the neuraminidase. J Virol 68(3):1790–1796
Wan H, Gao J, Xu K, Chen H, Couzens LK, Rivers KH et al (2013) Molecular basis for broad neuraminidase immunity: conserved epitopes in seasonal and pandemic H1N1 as well as H5N1 influenza viruses. J Virol 87(16):9290–9300
Roubidoux EK, McMahon M, Carreño JM, Capuano C, Jiang K, Simon V, et al. Novel epitopes of human monoclonal antibodies targeting the influenza virus N1 neuraminidase. bioRxiv. 2021.
Coker T, Meseko C, Odaibo G, Olaleye D (2014) Circulation of the low pathogenic avian influenza subtype H5N2 virus in ducks at a live bird market in Ibadan. Nigeria Infect Dis Poverty 3(1):1–6
Coyle PV, Al Molawi NH, Kacem MABH, El Kahlout RA, Al Kuwari E, Al Khal A et al (2022) Reporting of RT-PCR cycle threshold (Ct) values during the first wave of COVID-19 in Qatar improved result interpretation in clinical and public health settings. J Med Microbiol 71(5):001499
Lu Y, Yan J, Feng Y, Xu C, Shi W, Mao H (2008) Rapid detection of H5 avian influenza virus by TaqMan-MGB real-time RT-PCR. Lett Appl Microbiol 46(1):20–25
Yang F, Xu L, Liu F, Yao H, Wu N, Wu H (2020) Development and evaluation of a TaqMan MGB RT-PCR assay for detection of H5 and N8 subtype influenza virus. BMC Infect Dis 20(1):1–7
Das SR, Hensley SE, David A, Schmidt L, Gibbs JS, Puigbò P et al (2011) Fitness costs limit influenza A virus hemagglutinin glycosylation as an immune evasion strategy. Proc Natl Acad Sci 108(51):E1417–E1422
Linster M, van Boheemen S, de Graaf M, Schrauwen EJ, Lexmond P, Mänz B et al (2014) Identification, characterization, and natural selection of mutations driving airborne transmission of A/H5N1 virus. Cell 157(2):329–339
Lee C-W, Suarez DL, Tumpey TM, Sung H-W, Kwon Y-K, Lee Y-J et al (2005) Characterization of highly pathogenic H5N1 avian influenza A viruses isolated from South Korea. J Virol 79(6):3692–3702
Burke DF, Smith DJ (2014) A recommended numbering scheme for influenza A HA subtypes. PLoS ONE 9(11):e112302
Ibrahim E, Sirawan A, El-Bazzal B, El Hage J, Abi Said M, Zaraket H et al (2016) Complete genome sequence of the first H5N1 avian influenza virus isolated from chickens in Lebanon in 2016. Genome Announc 4(5):e01062-e1116
Gohrbandt S, Veits J, Hundt J, Bogs J, Breithaupt A, Teifke JP et al (2011) Amino acids adjacent to the haemagglutinin cleavage site are relevant for virulence of avian influenza viruses of subtype H5. J Gen Virol 92(1):51–59
Nguyen HT, Nguyen T, Mishin VP, Sleeman K, Balish A, Jones J et al (2013) Antiviral susceptibility of highly pathogenic avian influenza A (H5N1) viruses isolated from poultry, Vietnam, 2009–2011. Emerg Infect Dis 19(12):1963
Yang H, Carney PJ, Mishin VP, Guo Z, Chang JC, Wentworth DE et al (2016) Molecular characterizations of surface proteins hemagglutinin and neuraminidase from recent H5Nx avian influenza viruses. J Virol 90(12):5770–5784
Takashita E, Ejima M, Ra Itoh, Miura M, Ohnishi A, Nishimura H et al (2014) A community cluster of influenza A (H1N1) pdm09 virus exhibiting cross-resistance to oseltamivir and peramivir in Japan, November to December 2013. Eurosurveillance 19(1):20666
Sonnberg S, Webby RJ, Webster RG (2013) Natural history of highly pathogenic avian influenza H5N1. Virus Res 178(1):63–77
Ghafouri SA, Langeroudi AG, Maghsoudloo H, Tehrani F, Khaltabadifarahani R, Abdollahi H et al (2017) Phylogenetic study-based hemagglutinin (HA) gene of highly pathogenic avian influenza virus (H5N1) detected from backyard chickens in Iran, 2015. Virus Genes 53(1):117–120
Smith GJ, Donis RO (2015) Nomenclature updates resulting from the evolution of avian influenza A (H5) virus clades 2.1.3.2a, 2.2.1, and 2.3.4 during 2013–2014. Influenza Other Respir Viruses 9(5):271–6
Song Y, Cui J, Song H, Ye J, Zhao Z, Wu S et al (2015) New reassortant H5N8 highly pathogenic avian influenza virus from waterfowl in Southern China. Front Microbiol 6:1170
Shi W, Gao GF (2021) Emerging H5N8 avian influenza viruses. Science 372(6544):784–786
Usui T, Soda K, Tomioka Y, Ito H, Yabuta T, Takakuwa H et al (2017) Characterization of clade 2.3.4.4 H5N8 highly pathogenic avian influenza viruses from wild birds possessing atypical hemagglutinin polybasic cleavage sites. Virus Genes 53(1):44–51
Wilks S, de Graaf M, Smith DJ, Burke DF (2012) A review of influenza haemagglutinin receptor binding as it relates to pandemic properties. Vaccine 30(29):4369–4376
Bogs J, Veits J, Gohrbandt S, Hundt J, Stech O, Breithaupt A et al (2010) Highly pathogenic H5N1 influenza viruses carry virulence determinants beyond the polybasic hemagglutinin cleavage site. PLoS ONE 5(7):e11826
Li K, Guan Y, Wang J, Smith G, Xu K, Duan L et al (2004) Genesis of a highly pathogenic and potentially pandemic H5N1 influenza virus in eastern Asia. Nature 430(6996):209–213
Matrosovich M, Zhou N, Kawaoka Y, Webster R (1999) The surface glycoproteins of H5 influenza viruses isolated from humans, chickens, and wild aquatic birds have distinguishable properties. J Virol 73(2):1146–1155
Wilson IA, Skehel JJ, Wiley D (1981) Structure of the haemagglutinin membrane glycoprotein of influenza virus at 3 Å resolution. Nature 289(5796):366–373
Stevens J, Blixt O, Tumpey TM, Taubenberger JK, Paulson JC, Wilson IA (2006) Structure and receptor specificity of the hemagglutinin from an H5N1 influenza virus. Science 312(5772):404–10
Lin AH, Cannon PM (2002) Use of pseudotyped retroviral vectors to analyze the receptor-binding pocket of hemagglutinin from a pathogenic avian influenza A virus (H7 subtype). Virus Res 83(1–2):43–56
Xu D, Newhouse EI, Amaro RE, Pao HC, Cheng LS, Markwick PR et al (2009) Distinct glycan topology for avian and human sialopentasaccharide receptor analogues upon binding different hemagglutinins: a molecular dynamics perspective. J Mol Biol 387(2):465–491
Su Y, Yang H-Y, Zhang B-J, Jia H-L, Tien P (2008) Analysis of a point mutation in H5N1 avian influenza virus hemagglutinin in relation to virus entry into live mammalian cells. Adv Virol 153(12):2253–2261
Hiono T, Ohkawara A, Ogasawara K, Okamatsu M, Tamura T, Chu D-H et al (2015) Genetic and antigenic characterization of H5 and H7 influenza viruses isolated from migratory water birds in Hokkaido, Japan and Mongolia from 2010 to 2014. Virus Genes 51(1):57–68
Kanehira K, Uchida Y, Takemae N, Hikono H, Tsunekuni R, Saito T (2015) Characterization of an H5N8 influenza A virus isolated from chickens during an outbreak of severe avian influenza in Japan in April 2014. Adv Virol 160(7):1629–1643
Vigerust DJ, Shepherd VL (2007) Virus glycosylation: role in virulence and immune interactions. Trends Microbiol 15(5):211–218
Bao D, Xue R, Zhang M, Lu C, Ma T, Ren C et al (2021) N-linked glycosylation plays an important role in budding of neuraminidase protein and virulence of influenza viruses. J Virol 95(3):e02042-e2120
Wu J, Zhang F, Wang M, Xu C, Song J, Zhou J et al (2010) Characterization of neuraminidases from the highly pathogenic avian H5N1 and 2009 pandemic H1N1 influenza A viruses. PLoS ONE 5(12):e15825
Yee KS, Carpenter TE, Cardona CJ (2009) Epidemiology of H5N1 avian influenza. Comp Immunol Microbiol Infect Dis 32(4):325–340
Eichelberger MC, Monto AS (2019) Neuraminidase, the forgotten surface antigen, emerges as an influenza vaccine target for broadened protection. J Infect Dis 219(1):75–80
De Jong MD, Thanh TT, Khanh TH, Hien VM, Smith GJ, Chau NV et al (2005) Oseltamivir resistance during treatment of influenza A (H5N1) infection. N Engl J Med 353(25):2667–2672
Le QM, Kiso M, Someya K, Sakai YT, Nguyen TH, Nguyen KH et al (2005) Avian flu: isolation of drug-resistant H5N1 virus. Nature 437(7062):1108
Tisoncik-Go J, Cordero KS, Rong L (2013) Analysis of oseltamivir resistance substitutions in influenza virus glycoprotein neuraminidase using a lentivirus-based surrogate assay system. Virol Sinica 28(2):81–91
Colman PM, Varghese J, Laver W (1983) Structure of the catalytic and antigenic sites in influenza virus neuraminidase. Nature 303(5912):41–44
Gubareva LV, Webster RG, Hayden FG (2001) Comparison of the activities of zanamivir, oseltamivir, and RWJ-270201 against clinical isolates of influenza virus and neuraminidase inhibitor-resistant variants. Antimicrob Agents Chemother 45(12):3403–3408
von Itzstein M, Wu WY, Kok GB, Pegg MS, Dyason JC, Jin B et al (1993) Rational design of potent sialidase-based inhibitors of influenza virus replication. Nature 363(6428):418–423
Author information
Authors and Affiliations
Contributions
MM: Researcher, Wrote the main Manuscript text, prepared figures and Tables. HK: Supervisor. AS: Supervisor, Producer. MHFM: Advisor, Phylogenetic Analyser. GNB: Advisor, Editor. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Additional information
Edited by Zhen Fu.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Motahhar, M., Keyvanfar, H., Shoushtari, A. et al. The arrival of highly pathogenic avian influenza viruses H5N8 in Iran through two windows, 2016. Virus Genes 58, 527–539 (2022). https://doi.org/10.1007/s11262-022-01930-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-022-01930-8