Abstract
Purpose
To investigate the potential mechanisms of miR-211-3p on induction chemotherapy (IC) sensitivity in hypopharyngeal squamous cell carcinoma (HSCC).
Methods
qRT-PCR was assessed to compare the miR-211-3p expression between IC sensitive and insensitive tumor tissues. The MTT assay was performed to analyze cell proliferation and viability to paclitaxel after alteration of miR-211-3p. Flow cytometry assay was conducted to explore cell apoptosis. Transwell assay was used to explore the effect of miR-211-3p on cell migration. Transcriptome sequencing was then performed to select differentially expressed genes (DEGs) after over-expression of miR-211-3p. GO and KEGG enrichment analyses were conducted to annotate DEGs. PPI analysis was conducted to screen candidate genes. The differential expression and survival status of candidate genes were further validated in TCGA-HNSCC data. The single sample GSEA method was used to investigate the association between downstream genes and immune cell infiltration.
Results
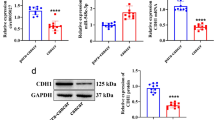
miR-211-3p was up-regulated in IC insensitive larynx–hypopharyngeal tumor tissues. Over-expression of miR-211-3p promoted cell proliferation and migration, and inhibited apoptosis.
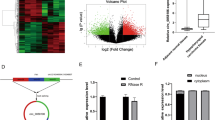
The IC50 value of miR-211-3p overexpression (OE) group was significantly higher than negative control (NC) group treated with paclitaxel, suggesting miR-211-3p enhanced IC insensitivity in HSCC. We found 778 DEG after over-expression of miR-211-3p and 11 significant genes were then identified. Finally, colony stimulating factor 2 (CSF2) and C-C motif chemokine ligand 20 (CCL20) were validated to be significantly high expressed and associated with poorer overall survival in head and neck squamous cell carcinoma, which were involved in TNF signaling pathway and then regulated immune cell infiltration.
Conclusion
The miR-211-3p could promote HSCC progression and upregulate CSF2/CCL20/TNF signaling to promote IC insensitivity in HSCC, which may provide new ideas for HSCC therapy.





Similar content being viewed by others
Data availability
The datasets generated during and/or analysed during the current study are not publicly available due to the raw data also forms part of an ongoing study, but are available from the corresponding author on reasonable request.
References
Bray F, Ferlay J, Soerjomataram I et al (2018) Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 68(6):394–424. https://doi.org/10.3322/caac.21492
Newman JR, Connolly TM, Illing EA et al (2015) Survival trends in hypopharyngeal cancer: a population-based review. Laryngoscope 125(3):624–629. https://doi.org/10.1002/lary.24915
Siegel RL, Miller KD, Fuchs HE et al (2021) (2021) Cancer statistics. CA Cancer J Clin 71(1):7–33. https://doi.org/10.3322/caac.21654
Gordin E (2018) Neoadjuvant chemotherapy for hypopharyngeal squamous cell carcinoma and personalized medicine in head and neck cancer. Ann Surg Oncol 25(4):848–849. https://doi.org/10.1245/s10434-017-6258-8
Lu TX, Rothenberg ME (2018) MicroRNA. J Allergy Clin Immunol 141(4):1202–1207. https://doi.org/10.1016/j.jaci.2017.08.034
Van Roosbroeck K, Fanini F, Setoyama T et al (2017) Combining anti-miR-155 with chemotherapy for the treatment of lung cancers. Clin Cancer Res 23(11):2891–2904. https://doi.org/10.1158/1078-0432.CCR-16-1025
Wu K, Hu Y, Yan K et al (2020) microRNA-10b confers cisplatin resistance by activating AKT/mTOR/P70S6K signaling via targeting PPARγ in esophageal cancer. J Cell Physiol 235(2):1247–1258. https://doi.org/10.1002/jcp.29040
Xue J, Chi Y, Chen Y et al (2016) MiRNA-621 sensitizes breast cancer to chemotherapy by suppressing FBXO11 and enhancing p53 activity. Oncogene 35(4):448–458. https://doi.org/10.1038/onc.2015.96
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-Seq data with DESeq2. Genome Biol 15(12):550. https://doi.org/10.1186/s13059-014-0550-8
Yu G, Wang LG, Han Y et al (2012) clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16(5):284–287. https://doi.org/10.1089/omi.2011.0118
Wu T, Hu E, Xu S et al (2021) clusterProfiler 4.0: a universal enrichment tool for interpreting omics data. Innovation (NY) 2(3):100141. https://doi.org/10.1016/j.xinn.2021.100141
Szklarczyk D, Gable AL, Lyon D et al (2019) STRING v11: protein–protein association networks with increased coverage, supporting functional discovery in genomewide experimental datasets. Nucleic Acids Res 47(D1):D607–D613. https://doi.org/10.1093/nar/gky1131
Kohl M, Wiese S, Warscheid B (2011) Cytoscape: software for visualization and analysis of biological networks. Methods Mol Biol 696:291–303. https://doi.org/10.1007/978-1-60761-987-1_18
Hänzelmann S, Castelo R, Guinney J (2013) GSVA: gene set variation analysis for microarray and RNA-Seq data. BMC Bioinform 14(1):1–15. https://doi.org/10.1186/1471-2105-14-7
Bindea G, Mlecnik B, Tosolini M et al (2013) Spatiotemporal dynamics of intratumoral immune cells reveal the immune landscape in human cancer. Immunity 39(4):782–795. https://doi.org/10.1016/j.immuni.2013.10.003
Qin X, Guo H, Wang X et al (2019) Exosomal miR-196a derived from cancer-associated fibroblasts confers cisplatin resistance in head and neck cancer through targeting CDKN1B and ING5. Genome Biol 20(1):12. https://doi.org/10.1186/s13059-018-1604-0
Sun X, Xiao D, Xu T et al (2016) miRNA-24-3p promotes cell proliferation and regulates chemosensitivity in head and neck squamous cell carcinoma by targeting CHD5. Future Oncol 12(23):2701–2712. https://doi.org/10.2217/fon-2016-0179
Peng F, Zhang H, Du Y et al (2015) miR-23a promotes cisplatin chemoresistance and protects against cisplatin-induced apoptosis in tongue squamous cell carcinoma cells through Twist. Oncol Rep 33(2):942–950. https://doi.org/10.3892/or.2014.3664
Chang KW, Liu CJ, Chu TH et al (2008) Association between high miR-211 microRNA expression and the poor prognosis of oral carcinoma. J Dent Res 87(11):1063–1068. https://doi.org/10.1177/154405910808701116
Chen YF, Yang CC, Kao SY et al (2016) MicroRNA-211 enhances the oncogenicity of carcinogen-induced oral carcinoma by repressing TCF12 and increasing antioxidant activity. Cancer Res 76(16):4872–4886. https://doi.org/10.1158/0008-5472.CAN-15-1664
Chu TH, Yang CC, Liu CJ et al (2013) miR-211 promotes the progression of head and neck carcinomas by targeting TGFβRII. Cancer Lett 337(1):115–124. https://doi.org/10.1016/j.canlet.2013.05.032
Lian M, Wang H, Fang J et al (2017) Microarray gene expression analysis of chemosensitivity for docetaxel, cisplatin and 5-fluorouracil (TPF) combined chemotherapeutic regimen in hypopharyngeal squamous cell carcinoma. Chin J Cancer Res 29(3):204–212. https://doi.org/10.21147/j.issn.1000-9604.2017.03.06
Hong IS (2016) Stimulatory versus suppressive effects of GM-CSF on tumor progression in multiple cancer types. Exp Mol Med 48(7):e242. https://doi.org/10.1038/emm.2016.64
Sielska M, Przanowski P, Wylot B et al (2013) Distinct roles of CSF family cytokines in macrophage infiltration and activation in glioma progression and injury response. J Pathol 230(3):310–321. https://doi.org/10.1002/path.4192
Xu Z, Zhang Y, Xu M et al (2019) Demethylation and overexpression of CSF2 are involved in immune response, chemotherapy resistance, and poor prognosis in colorectal cancer. OncoTargets Ther 12:11255–11269. https://doi.org/10.2147/OTT.S216829
Wang TT, Zhao YL, Peng LS et al (2017) Tumour-activated neutrophils in gastric cancer foster immune suppression and disease progression through GM-CSF-PD-L1 pathway. Gut 66(11):1900–1911. https://doi.org/10.1136/gutjnl-2016-313075
Wang D, Yang L, Yu W et al (2019) Colorectal cancer cell-derived CCL20 recruits regulatory T cells to promote chemoresistance via FOXO1/CEBPB/NF-κB signaling. J Immunother Cancer 7(1):215. https://doi.org/10.1186/s40425-019-0701-2
Chen W, Qin Y, Wang D et al (2018) CCL20 triggered by chemotherapy hinders the therapeutic efficacy of breast cancer. PLoS Biol 16(7):e2005869. https://doi.org/10.1371/journal.pbio.2005869
He H, Wu J, Zang M et al (2017) CCR6+ B lymphocytes responding to tumor cell-derived CCL20 support hepatocellular carcinoma progression via enhancing angiogenesis. Am J Cancer Res 7(5):1151–1163
Song Q, Shang J, Zhang C et al (2020) Transcription factor RUNX3 promotes CD8+ T cell recruitment by CCL3 and CCL20 in lung adenocarcinoma immune microenvironment. J Cell Biochem 121(5–6):3208–3220. https://doi.org/10.1002/jcb.29587
Acknowledgements
We thank all individuals who take part in this research.
Funding
This project was supported and funded by Beijing Natural Science Foundation Program and Scientific Research Key Program of Beijing Municipal Commission of Education (KZ201910025034); Beijing Municipal Administration of Hospitals’ Ascent Plan (DFL20180202); National Key R&D Program of China (2020YFB1312805); National Natural Science Foundation of China (82002880); Beijing Municipal Administration of Hospitals Incubating Program (PX2021008); and Beijing Hospitals Authority Youth Programme (QML20200205).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by LW, RW and TH. The first draft of the manuscript was written by LW, RW and JF, and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
Financial interests: all authors declare they have no financial interests. The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical approval
No ethical approval is required.
Informed consent
Informed consent is not required.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Wang, L., Wang, R., Huang, T. et al. miR-211-3p enhances induction chemotherapy insensitivity by upregulating CSF2/CCL20/TNF signaling in hypopharyngeal squamous cell carcinoma. Mol Biol Rep 49, 6103–6112 (2022). https://doi.org/10.1007/s11033-022-07401-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-022-07401-5